Automatic gene expression estimation from microarray images. Daniel O. Dantas Adviser: : Junior Barrera
|
|
|
- Cathleen Hoover
- 5 years ago
- Views:
Transcription
1 Automatic gene expression estimation from microarray images Daniel O. Dantas Adviser: : Junior Barrera IME-USP
2 Summary Introduction Problem definition Solution strategy Image segmentation Signal estimation Validation Conclusion
3 Introduction Microarray is a hybridization based technology used to measure the relative abundance of mrna from two samples (cancer and normal tissue, bacteries under normal and stressing conditions) Hybridization = matching of pairs of nucleic acid
4 Data acquisition
5 What is it for? Used to compare gene expression under different conditions. Gene expression is the entire process that takes the information contained in genes on DNA and turns that information into proteins. (edtech.clas.pdx.edu)
6 Knowledge evolution in Gene expression genetics
7 How does it work? Fix in a glass slide samples of cdna. Extract mrna from the two kinds of cells you want to analyze. Label copies of the mrna from each sample with different fluorescent dyes. Pour the two soups onto the glass slide and leave it there for some hours.
8 Cinética de hibridização cdnas Captura das Imagens Cy5 Misturas de cdnas marcados CY3 e CY5 Cy3
9 How does it work? If the mrna finds a matching cdna, they will hybridize. The more mrna in a sample, the more the respective color will lit. The scanner measures the light emitted by the fluorchrome when excited by a light at an appropriate wavelength.
10 A scanned image of a microarray slide
11 A microarray slide Is a small glass slide with about 1 x3 The resolution of a typical microarray image is about 10µm m (1000 pixels/cm). Each pixel of one channel has 16 bits = 2 bytes (ranges from 0 to 65535) 2 bytes x 2 channels x 2000 x 4000 = 32MB
12 A microarray slide The red channel represents the cy5 (wavelength = 635nm) The green channel represents the cy3 (wavelength = 532nm)
13 Problem definition Create a table with the estimated gene expression of each gene spoted in the slide automatically and reliably.
14 Aplication We can use the expression data to compare the behavior of many genes and classify them using clustering techniques, for example.
15 Available solutions Scanalyze: usually doesn t t find misaligned spots. SpotFinder(TIGR): subarrays must be placed manually. Arrayvision: very good on locating misaligned spots; many options. UCSF Spot: : does everything automatically if the image is perfect. Quantarray,, F-scanF scan, Dapple, Genepix, Imagene etc. All of them require user interaction to some level.
16 Our aim... Is to reduce the user interaction, doing the job automatically and measuring correctly the relative mrna concentrations. This will make the process cheaper and faster. User interaction makes the segmentation subjective. Eliminating that, the results may be more reproducible.
17 Solution strategy Manual steps Tilt correction (optional) Microarray geometry parameter setting Automatic steps Subarray gridding using image profiles Spots gridding using image profiles Spots detection Gene expression generation
18 Our software
19 Parameter setting In this window the user sets parameters for a whole family of arrays He can save in a file for reusing them
20 Microarray image segmentation process Hirata R, Barrera J, Hashimoto R, Dantas D, Esteves G. In press, 2002.
21 Microarray image segmentation process Delimited the spot, we must choose which pixels will be used in the signal estimation Or We we can can use select every some pixel of them based on the histogram information Example 15% of intensity of foreground The same is done in the background
22 A vertical image profile is the sum of the spots values of each image line
23 The subarray gridding Is done by filtering the horizontal and vertical profiles
24 And finally taking the local minima of the filtered profile
25 the same is done with the horizontal profile. Here the result
26 Spots gridding is done separately for each subarray
27 The profile filtering is simpler having just one step, and also uses local minima
28 The spots detection step is basically the application of the Watershed operator
29 To avoid oversegmentation the image must be filtered
30 The filtered image also gives markers that will be used in Watershed
31 We give as input to the Watershed the markers, grid and the filtered image gradient
32 Here the resulting grid in white and spots cortours in light blue
33 Segmentation example
34 Segmentation example
35 Raw data to the gene expression estimation step The raw data of a spot consists on: the pixels values of both channels inside its rectangular region of interest which pixels belong to foreground or backround Foreground is the region with spotted cdna Background is the region without it.
36 Raw data to the gene expression estimation step
37 Gene expression estimation Is to find a value that represents the relative quantity of mrna in the two samples.
38 Some techniques to estimate gene expression Linear regression or least-squares squares fit of the values of pixels in the two channels.
39 Some techniques to estimate gene expression (ch1i-ch1b) ch1b) / (ch2i-ch2b) ch2b) where chxi is the estimated foreground intensity and chxb is the estimated backround intensity of channel X.
40 Some techniques to estimate gene expression To estimate chxi and chxb we can do: mean or median of all pixels in the foreground and background. mean or median of some percentiles in the foreground and background (fixed region method) mean or median of higher percentiles of all the pixels in the rectangle to estimate chxi and of lower percentiles to estimate the chxb.. Foreground and background information is ignored (histogram method)
41 Some techniques to estimate gene expression In both, fixed region and histogram method, we look at parts of graphics like this, with the ordered values of the pixels of both channels.
42 Some techniques to estimate gene expression This graphic shows the quotient green/red, obtained by dividing the curves of the last graphic.
43 43A - IL1
44 43A - Trp1
45 43B - IL1
46 Validation We made controlled experiments to test the expression estimation techniques. The objective of the experiment was to test how expression was affected by: position in the slide dilution of cdna length of mrna fragments being marked with cy3 or cy5
47 Validation We spotted microarrays with 32 blocks, each block with 6 genes x 5 dilutions x 2 repetitions + 4 landmarks = 64 spots We made six slides like this and, onto them, we poured six different mrna soups: Dilution gene 43A 43B 44A 44B 45A 45B Irf Trp ST IL Q Lys
48 Validation Here each point is the value of a spot obtained by the fixed region method. Spots from different dilutions are grouped. The black ones are from the three bigger mrna fragments, and the red, from the three smaller. sqrt( ( (ch1i A -ch1b A ) x (ch2i B -ch2b B ) ) sqrt( ( (ch2i A -ch2b A ) x (ch1i B -ch1b B ) )
49 Validation And here is the best result, obtained with the histogram method. sqrt( ( (ch1i A -ch1b A ) x (ch2i B -ch2b B ) ) sqrt( ( (ch2i A -ch2b A ) x (ch1i B -ch1b B ) )
50 Validation Applying the least-squares squares fit to the data of each spot, we obtain results like this for the six genes. gene stddev mean Irf 0,2749 1,6176 1,00 Trp 0,4605 0,3999 0,25 ST0280 0,9945 1,4849 0,92 IL 1,8427 9,8712 6,10 Q 2,1836 6,9623 4,30 Lys 3,3600 2,1883 1,35
51 Validation Applying the histogram method to the data of each spot, we obtain results like this for the six genes. gene stddev mean Irf 0,2768 1,6411 1,00 Trp 0,6370 2,0420 1,24 ST0280 0,5019 2,4680 1,50 IL 1,5947 9,2869 5,66 Q 1,1552 7,3863 4,50 Lys 2,6532 6,5680 4,00
52 Normalization The expected expression of the gene IRF was 1.0 but the expression found was 1.6 This is due to the physical properties of the dyes.
53 Normalization When we have a single slide, we must eliminate the constant k assuming, when appropriate, that we can normalize all the spots using the expression of a housekeeping gene
54 Normalization When we have a single slide, we must eliminate the constant k assuming, when appropriate, that we can normalize all the spots using the expression of a housekeeping gene x = k (ch1i ch1b) (ch2i ch2b)
55 Normalization by swap Consists on eliminating the influence of the dyes properties by using two slides, and swapping the dye used to label the mrna sample. Use it if you find the single slide normalization hypotheses too strong.
56 Normalization by swap Better results can be achieved by doing swap experiments. x = k (ch1i A ch1b A ) = (ch2i B ch2b B ) 1 (ch2i A ch2b A ) (ch1i B ch1b B ) k
57 Normalization by swap Better results can be achieved by doing swap experiments. x = (ch1i (ch2i A A ch1b ch2b A A ) ) (ch2i (ch1i B B ch2b ch1b B B ) )
58 Normalization by swap Using the data obtained by least-sqares sqares fit from the two slides, the deviations decreases in all genes. gene stddev mean Irf 0,2224 0,9130 1,00 Trp 0,4039 0,7801 0,85 ST0280 0,5492 1,1251 1,23 IL 0,4567 3,4928 3,83 Q 0,9869 3,7146 4,07 Lys 1,3503 1,2297 1,35
59 Normalization by swap Using the data obtained by the histogram method from the two slides, the deviations decreases in all genes too. gene stddev mean Irf 0,0389 0,8959 1,00 Trp 0,1325 1,1096 1,24 ST0280 0,1482 1,3645 1,52 IL 0,2716 3,4475 3,85 Q 0,2964 3,8226 4,27 Lys 0,6194 2,7546 3,07
60 Normalization by swap Assuming that the best estimators are the ones with smaller standard deviation, we analyzed the resulting standard deviation of some different ways of choosing the pixels.
61 Normalization by swap Standard deviations using different values of percentiles for the foreground and bachground.. Histogram method. G43: dilution = 5 Higher background Lower background Higher foreground Lower foreground
62 Normalization by swap Standard deviations using different values of percentiles for the foreground and bachground.. Histogram method. G44: dilution = 2 G45: dilution = 10
63 Gene expression generation The program saves the expression data in a tab separated text file The file has the same format of the ones generated by ScanAlyze
64 Conclusion We created an automatic method for segmenting microarray images and estimating gene expression. The process was validated by controlled biochemical experiments. Some future steps: Automatic tilt correction Automatic identification of bad spots Statistically test if the controlled experiments represent properly real experiments. Automatic choice of the best estimation method Assign error bars to expression
Quality control of microarrays
 Quality control of microarrays Solveig Mjelstad Angelskår Intoduction to Microarray technology September 2009 Overview of the presentation 1. Image analysis 2. Quality Control (QC) general concepts 3.
Quality control of microarrays Solveig Mjelstad Angelskår Intoduction to Microarray technology September 2009 Overview of the presentation 1. Image analysis 2. Quality Control (QC) general concepts 3.
Automated cdna microarray image segmentation
 Automated cdna microarray image segmentation Author Liew, Alan Wee-Chung, Yan, Hong Published 2007 Conference Title Proceedings of the International Symposium on Computational Models for Life Sciences
Automated cdna microarray image segmentation Author Liew, Alan Wee-Chung, Yan, Hong Published 2007 Conference Title Proceedings of the International Symposium on Computational Models for Life Sciences
Preparation of Sample Hybridization Scanning and Image Analysis
 Preparation of Sample Hybridization Scanning and Image Analysis Sample preparation 1. Design experiment 2. Perform experiment Question? Replicates? Test? mutant wild type 3. Precipitate RNA 4. Label RNA
Preparation of Sample Hybridization Scanning and Image Analysis Sample preparation 1. Design experiment 2. Perform experiment Question? Replicates? Test? mutant wild type 3. Precipitate RNA 4. Label RNA
Microarray Data Pre-processing. Ana H. Barragan Lid
 Microarray Data Pre-processing Ana H. Barragan Lid Hybridized Microarray Imaged in a microarray scanner Scanner produces fluorescence intensity measurements Intensities correspond to levels of hybridization
Microarray Data Pre-processing Ana H. Barragan Lid Hybridized Microarray Imaged in a microarray scanner Scanner produces fluorescence intensity measurements Intensities correspond to levels of hybridization
Developed by BioDiscovery, Inc. DualChip evaluation software User Manual Version 1.1
 Developed by BioDiscovery, Inc. DualChip evaluation software User Manual Version 1.1 1 Table of contents 1. INTRODUCTION...3 2. SCOPE OF DELIVERY...4 3. INSTALLATION PROCEDURES...5 3.1. SYSTEM REQUIREMENTS...
Developed by BioDiscovery, Inc. DualChip evaluation software User Manual Version 1.1 1 Table of contents 1. INTRODUCTION...3 2. SCOPE OF DELIVERY...4 3. INSTALLATION PROCEDURES...5 3.1. SYSTEM REQUIREMENTS...
Low-level Analysis. cdna Microarrays. Lecture 2 Low Level Gene Expression Data Analysis. Stat 697K, CS 691K, Microbio 690K
 Lecture 2 Low Level Gene Expression Data nalysis Stat 697K, CS 691K, icrobio 690K Statistical Challenges odel variation of data not related to gene expression Compare expression for the same gene across
Lecture 2 Low Level Gene Expression Data nalysis Stat 697K, CS 691K, icrobio 690K Statistical Challenges odel variation of data not related to gene expression Compare expression for the same gene across
Computational Genomics. High-throughput experimental biology
 Computational Genomics 10-810/02 810/02-710, Spring 2009 Gene Expression Analysis Data pre-processing processing Eric Xing Lecture 15, March 4, 2009 Reading: class assignment Eric Xing @ CMU, 2005-2009
Computational Genomics 10-810/02 810/02-710, Spring 2009 Gene Expression Analysis Data pre-processing processing Eric Xing Lecture 15, March 4, 2009 Reading: class assignment Eric Xing @ CMU, 2005-2009
In our previous lecture, we understood the vital parameters to be taken into consideration before data acquisition and scanning.
 Interactomics: Protein Arrays & Label Free Biosensors Professor Sanjeeva Srivastava MOOC NPTEL Course Indian Institute of Technology Bombay Module 7 Lecture No 34 Software for Image scanning and data processing
Interactomics: Protein Arrays & Label Free Biosensors Professor Sanjeeva Srivastava MOOC NPTEL Course Indian Institute of Technology Bombay Module 7 Lecture No 34 Software for Image scanning and data processing
GenePix Application Note
 GenePix Application Note Determining the Signal-to-Noise Ratio and Optimal Photomultiplier gain setting in the GenePix 4000B Siobhan Pickett, M.S., Sean Carriedo, Ph.D. and Chang Wang, Ph.D. Axon Instruments,
GenePix Application Note Determining the Signal-to-Noise Ratio and Optimal Photomultiplier gain setting in the GenePix 4000B Siobhan Pickett, M.S., Sean Carriedo, Ph.D. and Chang Wang, Ph.D. Axon Instruments,
Scanning and Image Processing -by Steve Clough
 Scanning and Image Processing -by Steve Clough cdna microarrays use two dyes with well separated emission spectra such as Cy3 and Cy5 to allow direct comparisons on single slide GSI Lumonics Ratio of Expression
Scanning and Image Processing -by Steve Clough cdna microarrays use two dyes with well separated emission spectra such as Cy3 and Cy5 to allow direct comparisons on single slide GSI Lumonics Ratio of Expression
MICROARRAY IMAGE ANALYSIS PROGRAM
 Revision submitted for publication to Loyola Schools Review, 13 November 2002 MICROARRAY IMAGE ANALYSIS PROGRAM Paul Ignatius D. Echevarria Jerome C. Punzalan John Paul C. Vergara Department of Information
Revision submitted for publication to Loyola Schools Review, 13 November 2002 MICROARRAY IMAGE ANALYSIS PROGRAM Paul Ignatius D. Echevarria Jerome C. Punzalan John Paul C. Vergara Department of Information
GenePix Application Note
 GenePix Application Note Biological Relevance of GenePix Results Shawn Handran, Ph.D. and Jack Y. Zhai, Ph.D. Axon Instruments, Inc. 3280 Whipple Road, Union City, CA 94587 Last Updated: Aug 22, 2003.
GenePix Application Note Biological Relevance of GenePix Results Shawn Handran, Ph.D. and Jack Y. Zhai, Ph.D. Axon Instruments, Inc. 3280 Whipple Road, Union City, CA 94587 Last Updated: Aug 22, 2003.
Donuts, Scratches and Blanks: Robust Model-Based Segmentation of Microarray Images
 Donuts, Scratches and Blanks: Robust Model-Based Segmentation of Microarray Images Qunhua Li 1,a, Chris Fraley 1,a, Roger E. Bumgarner 2,b, Ka Yee Yeung 2,c, Adrian E. Raftery 1,a Technical Report no.
Donuts, Scratches and Blanks: Robust Model-Based Segmentation of Microarray Images Qunhua Li 1,a, Chris Fraley 1,a, Roger E. Bumgarner 2,b, Ka Yee Yeung 2,c, Adrian E. Raftery 1,a Technical Report no.
ScanArray Overview. Principle of Operation. Instrument Components
 ScanArray Overview The GSI Lumonics ScanArrayÒ Microarray Analysis System is a scanning laser confocal fluorescence microscope that is used to determine the fluorescence intensity of a two-dimensional
ScanArray Overview The GSI Lumonics ScanArrayÒ Microarray Analysis System is a scanning laser confocal fluorescence microscope that is used to determine the fluorescence intensity of a two-dimensional
Steps involved in microarray analysis after the experiments
 Steps involved in microarray analysis after the experiments Scanning slides to create images Conversion of images to numerical data Processing of raw numerical data Further analysis Clustering Integration
Steps involved in microarray analysis after the experiments Scanning slides to create images Conversion of images to numerical data Processing of raw numerical data Further analysis Clustering Integration
NPTEL VIDEO COURSE PROTEOMICS PROF. SANJEEVA SRIVASTAVA
 HANDOUT LECTURE-31 MICROARRAY WORK-FLOW: IMAGE SCANNING AND DATA PROCESSING Slide 1: This module contains the summary of the discussion with Mr. Pankaj Khanna, an application specialist from Spinco Biotech,
HANDOUT LECTURE-31 MICROARRAY WORK-FLOW: IMAGE SCANNING AND DATA PROCESSING Slide 1: This module contains the summary of the discussion with Mr. Pankaj Khanna, an application specialist from Spinco Biotech,
Scan slides (Axon Genepix 4200AL)
 Page 1 Scan slides (Axon Genepix 4200AL) We need to scan the slides on both channels (Cy3 and Cy5) to obtain a 16-bit grayscale TIFF file for each. Typically these files are about 20-26Mb per channel,
Page 1 Scan slides (Axon Genepix 4200AL) We need to scan the slides on both channels (Cy3 and Cy5) to obtain a 16-bit grayscale TIFF file for each. Typically these files are about 20-26Mb per channel,
NPTEL VIDEO COURSE PROTEOMICS PROF. SANJEEVA SRIVASTAVA
 LECTURE-31 MICROARRAY WORK-FLOW: IMAGE SACNNING AND DATA PROCESSING TRANSCRIPT Welcome to the proteomics course. In today s lecture we will talk about microarray work-flow the image scanning and processing.
LECTURE-31 MICROARRAY WORK-FLOW: IMAGE SACNNING AND DATA PROCESSING TRANSCRIPT Welcome to the proteomics course. In today s lecture we will talk about microarray work-flow the image scanning and processing.
EmbryoCellect. RHS Scanning and Analysis Instructions. for. Genepix Pro Software
 EmbryoCellect RHS Scanning and Analysis Instructions for Genepix Pro Software EmbryoCellect Genepix Pro Scanning and Analysis Technical Data Sheet Version 1.0 October 2015 1 Copyright Reproductive Health
EmbryoCellect RHS Scanning and Analysis Instructions for Genepix Pro Software EmbryoCellect Genepix Pro Scanning and Analysis Technical Data Sheet Version 1.0 October 2015 1 Copyright Reproductive Health
Point Calibration. July 3, 2012
 Point Calibration July 3, 2012 The purpose of the Point Calibration process is to generate a map of voltages (for galvos) or motor positions of the pointing device to the voltages or pixels of the reference
Point Calibration July 3, 2012 The purpose of the Point Calibration process is to generate a map of voltages (for galvos) or motor positions of the pointing device to the voltages or pixels of the reference
Multiplexing as Essential Tool for Modern Biology
 Multiplexing as Essential Tool for Modern Biology Bio-Plex Seminar, Debrecen, 2012. Gyula Csanádi, PhD. The "Age of "-omics" Studying interrelationships at different level of complexity Genes - Unveiling
Multiplexing as Essential Tool for Modern Biology Bio-Plex Seminar, Debrecen, 2012. Gyula Csanádi, PhD. The "Age of "-omics" Studying interrelationships at different level of complexity Genes - Unveiling
Instructions for Howto Scan µarrays
 Instructions for Howto Scan µarrays Introduction After probing the µarray slides with samples, one is now ready to scan them. To scan a µarrays slide is too convert the biological information trapped on
Instructions for Howto Scan µarrays Introduction After probing the µarray slides with samples, one is now ready to scan them. To scan a µarrays slide is too convert the biological information trapped on
Products - Microarray Scanners - Laser Scanners - InnoScan 900 Series and MAPIX Software
 Products - Microarray Scanners - Laser Scanners - InnoScan 900 Series and MAPIX Software Arrayit offers the world s only next generation microarray scanning technology, with proprietary rotary motion control,
Products - Microarray Scanners - Laser Scanners - InnoScan 900 Series and MAPIX Software Arrayit offers the world s only next generation microarray scanning technology, with proprietary rotary motion control,
Microarray Image Analysis: Background Estimation using Region and Filtering Techniques
 Microarray Image Analysis: Background Estimation using Region and Filtering Techniques Anders Bengtsson December 9, 2003 Abstract This report examines properties of two main methods used for local background
Microarray Image Analysis: Background Estimation using Region and Filtering Techniques Anders Bengtsson December 9, 2003 Abstract This report examines properties of two main methods used for local background
Product Information. Introduction
 Microarray Scanner Calibration Slide To quantitatively analyze scanners performance and output, adjust and fine-tune scanners, and perform comparative analysis for multiple scanner units. To verify scanners
Microarray Scanner Calibration Slide To quantitatively analyze scanners performance and output, adjust and fine-tune scanners, and perform comparative analysis for multiple scanner units. To verify scanners
Instructions for Mapping * µarray Images using GenePix 5.0
 Instructions for Mapping * µarray Images using GenePix 5.0 Preliminary Information Make sure that the GenePix 5.0 software has been installed on your computer and you have the USB hardware dongle that
Instructions for Mapping * µarray Images using GenePix 5.0 Preliminary Information Make sure that the GenePix 5.0 software has been installed on your computer and you have the USB hardware dongle that
LSM 780 Confocal Microscope Standard Operation Protocol
 LSM 780 Confocal Microscope Standard Operation Protocol Basic Operation Turning on the system 1. Sign on log sheet according to Actual start time 2. Check Compressed Air supply for the air table 3. Switch
LSM 780 Confocal Microscope Standard Operation Protocol Basic Operation Turning on the system 1. Sign on log sheet according to Actual start time 2. Check Compressed Air supply for the air table 3. Switch
Abstract. comment reviews reports deposited research refereed research interactions information
 http://genomebiology.com/21/2/11/research/47.1 Research Sources of nonlinearity in cdna microarray expression measurements Latha Ramdas*, Kevin R Coombes, Keith Baggerly, Lynne Abruzzo, W Edward Highsmith,
http://genomebiology.com/21/2/11/research/47.1 Research Sources of nonlinearity in cdna microarray expression measurements Latha Ramdas*, Kevin R Coombes, Keith Baggerly, Lynne Abruzzo, W Edward Highsmith,
Preprocessing and Segregating Offline Gujarati Handwritten Datasheet for Character Recognition
 Preprocessing and Segregating Offline Gujarati Handwritten Datasheet for Character Recognition Hetal R. Thaker Atmiya Institute of Technology & science, Kalawad Road, Rajkot Gujarat, India C. K. Kumbharana,
Preprocessing and Segregating Offline Gujarati Handwritten Datasheet for Character Recognition Hetal R. Thaker Atmiya Institute of Technology & science, Kalawad Road, Rajkot Gujarat, India C. K. Kumbharana,
Microarray BASICA: Background Adjustment, Segmentation, Image Compression and Analysis of Microarray Images
 EURASIP Journal on Applied Signal Processing 24:1, 92 17 c 24 Hindawi Publishing Corporation Microarray BASICA: Background Adjustment, Segmentation, Image Compression and Analysis of Microarray Images
EURASIP Journal on Applied Signal Processing 24:1, 92 17 c 24 Hindawi Publishing Corporation Microarray BASICA: Background Adjustment, Segmentation, Image Compression and Analysis of Microarray Images
Exercise questions for Machine vision
 Exercise questions for Machine vision This is a collection of exercise questions. These questions are all examination alike which means that similar questions may appear at the written exam. I ve divided
Exercise questions for Machine vision This is a collection of exercise questions. These questions are all examination alike which means that similar questions may appear at the written exam. I ve divided
880 Quantum Electronics Optional Lab Construct A Pulsed Dye Laser
 880 Quantum Electronics Optional Lab Construct A Pulsed Dye Laser The goal of this lab is to give you experience aligning a laser and getting it to lase more-or-less from scratch. There is no write-up
880 Quantum Electronics Optional Lab Construct A Pulsed Dye Laser The goal of this lab is to give you experience aligning a laser and getting it to lase more-or-less from scratch. There is no write-up
GALILEO TMA CK 4500 HTS Tissue Microarray Platform
 GALILEO TMA CK 4500 HTS Tissue Microarray Platform Tissue Microarray (TMA) A Block Of Samples From Hundreds Of Blocks (S. M. Hewitt, M.D., Ph.D., Tissue Array Research Program, LP, CCR, NCI, NIH) TMA technology
GALILEO TMA CK 4500 HTS Tissue Microarray Platform Tissue Microarray (TMA) A Block Of Samples From Hundreds Of Blocks (S. M. Hewitt, M.D., Ph.D., Tissue Array Research Program, LP, CCR, NCI, NIH) TMA technology
Graphics packages can be bit-mapped or vector. Both types of packages store graphics in a different way.
 Graphics packages can be bit-mapped or vector. Both types of packages store graphics in a different way. Bit mapped packages (paint packages) work by changing the colour of the pixels that make up the
Graphics packages can be bit-mapped or vector. Both types of packages store graphics in a different way. Bit mapped packages (paint packages) work by changing the colour of the pixels that make up the
Computer Vision. Howie Choset Introduction to Robotics
 Computer Vision Howie Choset http://www.cs.cmu.edu.edu/~choset Introduction to Robotics http://generalrobotics.org What is vision? What is computer vision? Edge Detection Edge Detection Interest points
Computer Vision Howie Choset http://www.cs.cmu.edu.edu/~choset Introduction to Robotics http://generalrobotics.org What is vision? What is computer vision? Edge Detection Edge Detection Interest points
Computer Graphics Fundamentals
 Computer Graphics Fundamentals Jacek Kęsik, PhD Simple converts Rotations Translations Flips Resizing Geometry Rotation n * 90 degrees other Geometry Rotation n * 90 degrees other Geometry Translations
Computer Graphics Fundamentals Jacek Kęsik, PhD Simple converts Rotations Translations Flips Resizing Geometry Rotation n * 90 degrees other Geometry Rotation n * 90 degrees other Geometry Translations
Multifluorescence The Crosstalk Problem and Its Solution
 Multifluorescence The Crosstalk Problem and Its Solution If a specimen is labeled with more than one fluorochrome, each image channel should only show the emission signal of one of them. If, in a specimen
Multifluorescence The Crosstalk Problem and Its Solution If a specimen is labeled with more than one fluorochrome, each image channel should only show the emission signal of one of them. If, in a specimen
Image Capture TOTALLAB
 1 Introduction In order for image analysis to be performed on a gel or Western blot, it must first be converted into digital data. Good image capture is critical to guarantee optimal performance of automated
1 Introduction In order for image analysis to be performed on a gel or Western blot, it must first be converted into digital data. Good image capture is critical to guarantee optimal performance of automated
Attenuation length in strip scintillators. Jonathan Button, William McGrew, Y.-W. Lui, D. H. Youngblood
 Attenuation length in strip scintillators Jonathan Button, William McGrew, Y.-W. Lui, D. H. Youngblood I. Introduction The ΔE-ΔE-E decay detector as described in [1] is composed of thin strip scintillators,
Attenuation length in strip scintillators Jonathan Button, William McGrew, Y.-W. Lui, D. H. Youngblood I. Introduction The ΔE-ΔE-E decay detector as described in [1] is composed of thin strip scintillators,
Locating Molecules Using GSD Technology Project Folders: Organization of Experiment Files...1
 .....................................1 1 Project Folders: Organization of Experiment Files.................................1 2 Steps........................................................................2
.....................................1 1 Project Folders: Organization of Experiment Files.................................1 2 Steps........................................................................2
Topics. - How to calibrate the LSM scanner. - How to clean the microscope. - How to adjust the pinhole alignment. - How to adjust the Collimator
 Topics - How to calibrate the LSM scanner - How to measure the PSF - How to clean the microscope - How to adjust the pinhole alignment - How to adjust the Collimator How to calibrate the LSM scanner The
Topics - How to calibrate the LSM scanner - How to measure the PSF - How to clean the microscope - How to adjust the pinhole alignment - How to adjust the Collimator How to calibrate the LSM scanner The
Image Analysis for Fluorescence
 Image Analysis for Fluorescence Terminology Table Image Analysis Macro Colocalization Intensity Dye AFI The extraction of meaningful information from digital images by means of digital image processing
Image Analysis for Fluorescence Terminology Table Image Analysis Macro Colocalization Intensity Dye AFI The extraction of meaningful information from digital images by means of digital image processing
Introduction to Image Analysis with
 Introduction to Image Analysis with PLEASE ENSURE FIJI IS INSTALLED CORRECTLY! WHAT DO WE HOPE TO ACHIEVE? Specifically, the workshop will cover the following topics: 1. Opening images with Bioformats
Introduction to Image Analysis with PLEASE ENSURE FIJI IS INSTALLED CORRECTLY! WHAT DO WE HOPE TO ACHIEVE? Specifically, the workshop will cover the following topics: 1. Opening images with Bioformats
CS 445 HW#2 Solutions
 1. Text problem 3.1 CS 445 HW#2 Solutions (a) General form: problem figure,. For the condition shown in the Solving for K yields Then, (b) General form: the problem figure, as in (a) so For the condition
1. Text problem 3.1 CS 445 HW#2 Solutions (a) General form: problem figure,. For the condition shown in the Solving for K yields Then, (b) General form: the problem figure, as in (a) so For the condition
Calculate Ratiometric FRET-Images with the FRET-Image-Script
 Tutorial Calculate Ratiometric FRET-Images with the FRET-Image-Script Summary This tutorial shows step-by-step, how the "FRET Image" script of SymPhoTime 64 can be used to calculate pixel-by-pixel the
Tutorial Calculate Ratiometric FRET-Images with the FRET-Image-Script Summary This tutorial shows step-by-step, how the "FRET Image" script of SymPhoTime 64 can be used to calculate pixel-by-pixel the
Multi-channel imaging cytometry with a single detector
 Multi-channel imaging cytometry with a single detector Sarah Locknar 1, John Barton 1, Mark Entwistle 2, Gary Carver 1 and Robert Johnson 1 1 Omega Optical, Brattleboro, VT 05301 2 Philadelphia Lightwave,
Multi-channel imaging cytometry with a single detector Sarah Locknar 1, John Barton 1, Mark Entwistle 2, Gary Carver 1 and Robert Johnson 1 1 Omega Optical, Brattleboro, VT 05301 2 Philadelphia Lightwave,
IncuCyte ZOOM Fluorescent Processing Overview
 IncuCyte ZOOM Fluorescent Processing Overview The IncuCyte ZOOM offers users the ability to acquire HD phase as well as dual wavelength fluorescent images of living cells producing multiplexed data that
IncuCyte ZOOM Fluorescent Processing Overview The IncuCyte ZOOM offers users the ability to acquire HD phase as well as dual wavelength fluorescent images of living cells producing multiplexed data that
Chapter 4. Displaying and Summarizing Quantitative Data. Copyright 2012, 2008, 2005 Pearson Education, Inc.
 Chapter 4 Displaying and Summarizing Quantitative Data Copyright 2012, 2008, 2005 Pearson Education, Inc. Dealing With a Lot of Numbers Summarizing the data will help us when we look at large sets of quantitative
Chapter 4 Displaying and Summarizing Quantitative Data Copyright 2012, 2008, 2005 Pearson Education, Inc. Dealing With a Lot of Numbers Summarizing the data will help us when we look at large sets of quantitative
RealSpot: software validating results from DNA microarray data analysis with spot images
 Physiol Genomics 21: 284 291, 2005. First published February 1, 2005; doi:10.1152/physiolgenomics.00236.2004. RealSpot: software validating results from DNA microarray data analysis with spot images Zhongming
Physiol Genomics 21: 284 291, 2005. First published February 1, 2005; doi:10.1152/physiolgenomics.00236.2004. RealSpot: software validating results from DNA microarray data analysis with spot images Zhongming
Influence of Dictionary Size on the Lossless Compression of Microarray Images
 Influence of Dictionary Size on the Lossless Compression of Microarray Images Robert Bierman 1, Rahul Singh 1 Department of Computer Science, San Francisco State University, San Francisco, CA bierman@sfsu.edu,
Influence of Dictionary Size on the Lossless Compression of Microarray Images Robert Bierman 1, Rahul Singh 1 Department of Computer Science, San Francisco State University, San Francisco, CA bierman@sfsu.edu,
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Purcell and beta factor without the diamond host for three wavelengths within the NV spectrum. Purcell factor for a dipole oriented along the a) x-axis, b)
Supplementary Figures Supplementary Figure 1. Purcell and beta factor without the diamond host for three wavelengths within the NV spectrum. Purcell factor for a dipole oriented along the a) x-axis, b)
Supplementary Information. Stochastic Optical Reconstruction Microscopy Imaging of Microtubule Arrays in Intact Arabidopsis thaliana Seedling Roots
 Supplementary Information Stochastic Optical Reconstruction Microscopy Imaging of Microtubule Arrays in Intact Arabidopsis thaliana Seedling Roots Bin Dong 1,, Xiaochen Yang 2,, Shaobin Zhu 1, Diane C.
Supplementary Information Stochastic Optical Reconstruction Microscopy Imaging of Microtubule Arrays in Intact Arabidopsis thaliana Seedling Roots Bin Dong 1,, Xiaochen Yang 2,, Shaobin Zhu 1, Diane C.
Standard Operating Procedure
 Standard Operating Procedure Nanosurf Atomic Force Microscopy Operation Facility NCCRD Nanotechnology Center for Collaborative Research and Development Department of Chemistry and Engineering Physics The
Standard Operating Procedure Nanosurf Atomic Force Microscopy Operation Facility NCCRD Nanotechnology Center for Collaborative Research and Development Department of Chemistry and Engineering Physics The
#P Quality Measures for Imaging-based Cellular Assays
 Abstract #P1224 - Quality Measures for Imaging-based Cellular Assays Ilya Ravkin, Vitra Bioscience, Inc. Z-factor and related measures are useful in estimating assay variability in HTS caused by assay
Abstract #P1224 - Quality Measures for Imaging-based Cellular Assays Ilya Ravkin, Vitra Bioscience, Inc. Z-factor and related measures are useful in estimating assay variability in HTS caused by assay
Imagers- Molecular, Cell Standard Operating Procedures
 Bio-Rad ChemiDoc XRS and Image Lab Software Jump to Export Images to other Apps Floid cell imaging station Life technologies Jump to Chemi-luminescence Protocol Imagers- Molecular, Cell Standard Operating
Bio-Rad ChemiDoc XRS and Image Lab Software Jump to Export Images to other Apps Floid cell imaging station Life technologies Jump to Chemi-luminescence Protocol Imagers- Molecular, Cell Standard Operating
Quick Guide. NucleoCounter NC-3000
 Quick Guide NucleoCounter NC-3000 Table of contents Setting up the FlexiCyte Protocol 2 Editing Image Capture and Analysis Parameters 3 Optimizing Exposure Time 4 Compensation for Spectral Overlap 6 Creating
Quick Guide NucleoCounter NC-3000 Table of contents Setting up the FlexiCyte Protocol 2 Editing Image Capture and Analysis Parameters 3 Optimizing Exposure Time 4 Compensation for Spectral Overlap 6 Creating
Renishaw InVia Raman microscope
 Laser Spectroscopy Labs Renishaw InVia Raman microscope Operation instructions 1. Turn On the power switch, system power switch is located towards the back of the system on the right hand side. Wait ~10
Laser Spectroscopy Labs Renishaw InVia Raman microscope Operation instructions 1. Turn On the power switch, system power switch is located towards the back of the system on the right hand side. Wait ~10
Cell Biology and Bioimaging Core
 Cell Biology and Bioimaging Core Leica TCS SP5 Operating Instructions Starting up the instrument 1. First, log in the log book located on the confocal desk. Include your name, your lab s PI, an account
Cell Biology and Bioimaging Core Leica TCS SP5 Operating Instructions Starting up the instrument 1. First, log in the log book located on the confocal desk. Include your name, your lab s PI, an account
Image analysis. CS/CME/BioE/Biophys/BMI 279 Oct. 31 and Nov. 2, 2017 Ron Dror
 Image analysis CS/CME/BioE/Biophys/BMI 279 Oct. 31 and Nov. 2, 2017 Ron Dror 1 Outline Images in molecular and cellular biology Reducing image noise Mean and Gaussian filters Frequency domain interpretation
Image analysis CS/CME/BioE/Biophys/BMI 279 Oct. 31 and Nov. 2, 2017 Ron Dror 1 Outline Images in molecular and cellular biology Reducing image noise Mean and Gaussian filters Frequency domain interpretation
(12) United States Patent (10) Patent No.: US 6,525,828 B1
 USOO6525828B1 (12) United States Patent (10) Patent No.: US 6,525,828 B1 Grosskopf (45) Date of Patent: *Feb. 25, 2003 (54) CONFOCAL COLOR 5,978,095 A 11/1999 Tanaami... 356/445 6,031,661. A 2/2000 Tanaami...
USOO6525828B1 (12) United States Patent (10) Patent No.: US 6,525,828 B1 Grosskopf (45) Date of Patent: *Feb. 25, 2003 (54) CONFOCAL COLOR 5,978,095 A 11/1999 Tanaami... 356/445 6,031,661. A 2/2000 Tanaami...
Spotxel 1.7 Microarray Image and Data Analysis Software User s Guide
 Spotxel 1.7 Microarray Image and Data Analysis Software User s Guide 27 April 2017 - Rev 7 Spotxel is only intended for research and not intended or approved for diagnosis of disease in humans or animals.
Spotxel 1.7 Microarray Image and Data Analysis Software User s Guide 27 April 2017 - Rev 7 Spotxel is only intended for research and not intended or approved for diagnosis of disease in humans or animals.
Comparing FCS and FRAP as methodologies for calculating diffusion
 Bi/BE 227 Winter 2018 Assignment #4 Comparing FCS and FRAP as methodologies for calculating diffusion Schedule: Jan 29: Assignment Jan 29-Feb 14: Work on assignment Feb 14: Student PowerPoint presentations.
Bi/BE 227 Winter 2018 Assignment #4 Comparing FCS and FRAP as methodologies for calculating diffusion Schedule: Jan 29: Assignment Jan 29-Feb 14: Work on assignment Feb 14: Student PowerPoint presentations.
The Bead. beadarray: : An R Package for Illumina BeadArrays. Bead Preparation and Array Production. Beads in Wells. Mark Dunning -
 beadarray: : An R Package for Illumina BeadArrays Mark Dunning - md392@cam.ac.uk PhD Student - Computational Biology Group, Department of Oncology - University of Cambridge Address The Bead Probe 23 b
beadarray: : An R Package for Illumina BeadArrays Mark Dunning - md392@cam.ac.uk PhD Student - Computational Biology Group, Department of Oncology - University of Cambridge Address The Bead Probe 23 b
Guidance on Using Scanning Software: Part 5. Epson Scan
 Guidance on Using Scanning Software: Part 5. Epson Scan Version of 4/29/2012 Epson Scan comes with Epson scanners and has simple manual adjustments, but requires vigilance to control the default settings
Guidance on Using Scanning Software: Part 5. Epson Scan Version of 4/29/2012 Epson Scan comes with Epson scanners and has simple manual adjustments, but requires vigilance to control the default settings
ROBOT VISION. Dr.M.Madhavi, MED, MVSREC
 ROBOT VISION Dr.M.Madhavi, MED, MVSREC Robotic vision may be defined as the process of acquiring and extracting information from images of 3-D world. Robotic vision is primarily targeted at manipulation
ROBOT VISION Dr.M.Madhavi, MED, MVSREC Robotic vision may be defined as the process of acquiring and extracting information from images of 3-D world. Robotic vision is primarily targeted at manipulation
Axioscan - Startup. 1. Turn on the Axioscan (button to the left) and turn on the computer. 2. Log on and start the ZEN Blue software from the desktop
 Axioscan - Startup 1. Turn on the Axioscan (button to the left) and turn on the computer 2. Log on and start the ZEN Blue software from the desktop 3. Press ZEN slidescan and Start System 4. Start by changing
Axioscan - Startup 1. Turn on the Axioscan (button to the left) and turn on the computer 2. Log on and start the ZEN Blue software from the desktop 3. Press ZEN slidescan and Start System 4. Start by changing
Akinori Mitani and Geoff Weiner BGGN 266 Spring 2013 Non-linear optics final report. Introduction and Background
 Akinori Mitani and Geoff Weiner BGGN 266 Spring 2013 Non-linear optics final report Introduction and Background Two-photon microscopy is a type of fluorescence microscopy using two-photon excitation. It
Akinori Mitani and Geoff Weiner BGGN 266 Spring 2013 Non-linear optics final report Introduction and Background Two-photon microscopy is a type of fluorescence microscopy using two-photon excitation. It
Non-Linear Optical Flow Cytometry Using a Scanned, Bessel Beam Light-Sheet
 1 Non-Linear Optical Flow Cytometry Using a Scanned, essel eam Light-Sheet Supplementary Information radley. Collier 1, Samir Awasthi 1,2, Deborah K. Lieu 3, James W. Chan 1,4* 1 Center for iophotonics,
1 Non-Linear Optical Flow Cytometry Using a Scanned, essel eam Light-Sheet Supplementary Information radley. Collier 1, Samir Awasthi 1,2, Deborah K. Lieu 3, James W. Chan 1,4* 1 Center for iophotonics,
How is the Digital Image Generated? Image Acquisition Devices
 In order for image analysis to be performed on a 2D gel, it must first be converted into digital data. Good image capture is critical to guarantee optimal performance of automated image analysis packages
In order for image analysis to be performed on a 2D gel, it must first be converted into digital data. Good image capture is critical to guarantee optimal performance of automated image analysis packages
Visioneer OneTouch Scanner. Installation Guide FOR WINDOWS
 Visioneer OneTouch Scanner Installation Guide FOR WINDOWS TABLE OF CONTENTS i TABLE OF CONTENTS Getting Started with your new Scanner....................... 1 Step 1: Installing the Scanner Software.......................
Visioneer OneTouch Scanner Installation Guide FOR WINDOWS TABLE OF CONTENTS i TABLE OF CONTENTS Getting Started with your new Scanner....................... 1 Step 1: Installing the Scanner Software.......................
Examination, TEN1, in courses SK2500/SK2501, Physics of Biomedical Microscopy,
 KTH Applied Physics Examination, TEN1, in courses SK2500/SK2501, Physics of Biomedical Microscopy, 2009-06-05, 8-13, FB51 Allowed aids: Compendium Imaging Physics (handed out) Compendium Light Microscopy
KTH Applied Physics Examination, TEN1, in courses SK2500/SK2501, Physics of Biomedical Microscopy, 2009-06-05, 8-13, FB51 Allowed aids: Compendium Imaging Physics (handed out) Compendium Light Microscopy
Chapter 17. Shape-Based Operations
 Chapter 17 Shape-Based Operations An shape-based operation identifies or acts on groups of pixels that belong to the same object or image component. We have already seen how components may be identified
Chapter 17 Shape-Based Operations An shape-based operation identifies or acts on groups of pixels that belong to the same object or image component. We have already seen how components may be identified
Segmentation of Microscopic Bone Images
 International Journal of Electronics Engineering, 2(1), 2010, pp. 11-15 Segmentation of Microscopic Bone Images Anand Jatti Research Scholar, Vishveshvaraiah Technological University, Belgaum, Karnataka
International Journal of Electronics Engineering, 2(1), 2010, pp. 11-15 Segmentation of Microscopic Bone Images Anand Jatti Research Scholar, Vishveshvaraiah Technological University, Belgaum, Karnataka
Opterra II Multipoint Scanning Confocal Microscope. Innovation with Integrity
 Opterra II Multipoint Scanning Confocal Microscope Enabling 4D Live-Cell Fluorescence Imaging through Speed, Sensitivity, Viability and Simplicity Innovation with Integrity Fluorescence Microscopy The
Opterra II Multipoint Scanning Confocal Microscope Enabling 4D Live-Cell Fluorescence Imaging through Speed, Sensitivity, Viability and Simplicity Innovation with Integrity Fluorescence Microscopy The
Confidence Intervals. Class 23. November 29, 2011
 Confidence Intervals Class 23 November 29, 2011 Last Time When sampling from a population in which 30% of individuals share a certain characteristic, we identified the reasonably likely values for the
Confidence Intervals Class 23 November 29, 2011 Last Time When sampling from a population in which 30% of individuals share a certain characteristic, we identified the reasonably likely values for the
Images and Graphics. 4. Images and Graphics - Copyright Denis Hamelin - Ryerson University
 Images and Graphics Images and Graphics Graphics and images are non-textual information that can be displayed and printed. Graphics (vector graphics) are an assemblage of lines, curves or circles with
Images and Graphics Images and Graphics Graphics and images are non-textual information that can be displayed and printed. Graphics (vector graphics) are an assemblage of lines, curves or circles with
Feature Level Data. Outline. Affymetrix GeneChip Design. Affymetrix GeneChip arrays Two color platforms
 Feature Level Data Outline Affymetrix GeneChip arrays Two color platforms Affymetrix GeneChip Design 5 3 Reference sequence TGTGATGGTGCATGATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT GTACTACCCAGTCTTCCGGAGGCTA
Feature Level Data Outline Affymetrix GeneChip arrays Two color platforms Affymetrix GeneChip Design 5 3 Reference sequence TGTGATGGTGCATGATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT GTACTACCCAGTCTTCCGGAGGCTA
CSSE463: Image Recognition Day 2
 CSSE463: Image Recognition Day 2 Roll call Announcements: Moodle has drop box for Lab 1 Next class: lots more Matlab how-to (bring your laptop) Questions? Today: Color and color features Do questions 1-2
CSSE463: Image Recognition Day 2 Roll call Announcements: Moodle has drop box for Lab 1 Next class: lots more Matlab how-to (bring your laptop) Questions? Today: Color and color features Do questions 1-2
ThermaViz. Operating Manual. The Innovative Two-Wavelength Imaging Pyrometer
 ThermaViz The Innovative Two-Wavelength Imaging Pyrometer Operating Manual The integration of advanced optical diagnostics and intelligent materials processing for temperature measurement and process control.
ThermaViz The Innovative Two-Wavelength Imaging Pyrometer Operating Manual The integration of advanced optical diagnostics and intelligent materials processing for temperature measurement and process control.
Volocity Tutorial Creating a Measurement Protocol in Volocity Software
 TUTORIAL NOTE Cellular Imaging and Analysis Volocity Tutorial Creating a Measurement Protocol in Volocity Software This tutorial will illustrate the process of creating a Measurement Protocol in Volocity
TUTORIAL NOTE Cellular Imaging and Analysis Volocity Tutorial Creating a Measurement Protocol in Volocity Software This tutorial will illustrate the process of creating a Measurement Protocol in Volocity
Enhancing the quality metric of protein microarray image *
 Wang et al. / J Zhejiang Univ SCI 2004 5(2):62-628 62 Journal of Zhejiang University SCIENCE ISSN 009-3095 http://www.zju.edu.cn/jzus E-mail: jzus@zju.edu.cn Enhancing the quality metric of protein microarray
Wang et al. / J Zhejiang Univ SCI 2004 5(2):62-628 62 Journal of Zhejiang University SCIENCE ISSN 009-3095 http://www.zju.edu.cn/jzus E-mail: jzus@zju.edu.cn Enhancing the quality metric of protein microarray
Experimental protocol PIPE
 Experimental protocol PIPE May 5, 2016 Abstract PIPE is a uorescence perturbation technique that works by measuring the expansion of a laser induced perturbation of photo convertible fused protein in the
Experimental protocol PIPE May 5, 2016 Abstract PIPE is a uorescence perturbation technique that works by measuring the expansion of a laser induced perturbation of photo convertible fused protein in the
Nature Methods: doi: /nmeth Supplementary Figure 1. Schematic of 2P-ISIM AO optical setup.
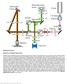 Supplementary Figure 1 Schematic of 2P-ISIM AO optical setup. Excitation from a femtosecond laser is passed through intensity control and shuttering optics (1/2 λ wave plate, polarizing beam splitting
Supplementary Figure 1 Schematic of 2P-ISIM AO optical setup. Excitation from a femtosecond laser is passed through intensity control and shuttering optics (1/2 λ wave plate, polarizing beam splitting
Center for Microscopy and Image Analysis Axio Scan.Z1 Operating Manual
 No index entries found. Center for Microscopy and Image Analysis Axio Scan.Z1 Operating Manual Table of contents 1. Starting procedure... 3 1.1. Turn on hardware... 3 1.2. Starting ZEN blue... 4 2. Load
No index entries found. Center for Microscopy and Image Analysis Axio Scan.Z1 Operating Manual Table of contents 1. Starting procedure... 3 1.1. Turn on hardware... 3 1.2. Starting ZEN blue... 4 2. Load
Supplementary Figure S1: Schematic view of the confocal laser scanning STED microscope used for STED-RICS. For a detailed description of our
 Supplementary Figure S1: Schematic view of the confocal laser scanning STED microscope used for STED-RICS. For a detailed description of our home-built STED microscope used for the STED-RICS experiments,
Supplementary Figure S1: Schematic view of the confocal laser scanning STED microscope used for STED-RICS. For a detailed description of our home-built STED microscope used for the STED-RICS experiments,
Reference and User Manual May, 2015 revision - 3
 Reference and User Manual May, 2015 revision - 3 Innovations Foresight 2015 - Powered by Alcor System 1 For any improvement and suggestions, please contact customerservice@innovationsforesight.com Some
Reference and User Manual May, 2015 revision - 3 Innovations Foresight 2015 - Powered by Alcor System 1 For any improvement and suggestions, please contact customerservice@innovationsforesight.com Some
MICROSCOPE LAB. Resolving Power How well specimen detail is preserved during the magnifying process.
 AP BIOLOGY Cells ACTIVITY #2 MICROSCOPE LAB OBJECTIVES 1. Demonstrate proper care and use of a compound microscope. 2. Identify the parts of the microscope and describe the function of each part. 3. Compare
AP BIOLOGY Cells ACTIVITY #2 MICROSCOPE LAB OBJECTIVES 1. Demonstrate proper care and use of a compound microscope. 2. Identify the parts of the microscope and describe the function of each part. 3. Compare
DIGITAL-MICROSCOPY CAMERA SOLUTIONS USB 3.0
 DIGITAL-MICROSCOPY CAMERA SOLUTIONS USB 3.0 PixeLINK for Microscopy Applications PixeLINK will work with you to choose and integrate the optimal USB 3.0 camera for your microscopy project. Ideal for use
DIGITAL-MICROSCOPY CAMERA SOLUTIONS USB 3.0 PixeLINK for Microscopy Applications PixeLINK will work with you to choose and integrate the optimal USB 3.0 camera for your microscopy project. Ideal for use
Ultrafast Technique of Impulsive Noise Removal with Application to Microarray Image Denoising
 Ultrafast Technique of Impulsive Noise Removal with Application to Microarray Image Denoising Bogdan Smolka 1, and Konstantinos N. Plataniotis 2 1 Silesian University of Technology, Department of Automatic
Ultrafast Technique of Impulsive Noise Removal with Application to Microarray Image Denoising Bogdan Smolka 1, and Konstantinos N. Plataniotis 2 1 Silesian University of Technology, Department of Automatic
WHITE PAPER FAST PROTEIN INTERACTION BINDING CURVES WITH INO S F-HS CONFOCAL MICROSCOPE
 WHITE PAPER FAST PROTEIN INTERACTION BINDING CURVES WITH INO S F-HS CONFOCAL MICROSCOPE Christian Tardif, Jean-Pierre Bouchard Pascal Gallant, Sebastien Roy, Ozzy Mermut September 2017 Introduction Protein-protein
WHITE PAPER FAST PROTEIN INTERACTION BINDING CURVES WITH INO S F-HS CONFOCAL MICROSCOPE Christian Tardif, Jean-Pierre Bouchard Pascal Gallant, Sebastien Roy, Ozzy Mermut September 2017 Introduction Protein-protein
SmartDoc 2.0 E5001-SDB Instruction Manual
 SmartDoc 2.0 E5001-SDB Instruction Manual Version 11.16 1 Table of Contents 1. Introduction 3 2. Warnings. 3 3. Unpacking.. 4 4. SmartDoc 2.0 Overview 4 5. Setting up the SmartDoc 2.0 5 6. Gel Viewing
SmartDoc 2.0 E5001-SDB Instruction Manual Version 11.16 1 Table of Contents 1. Introduction 3 2. Warnings. 3 3. Unpacking.. 4 4. SmartDoc 2.0 Overview 4 5. Setting up the SmartDoc 2.0 5 6. Gel Viewing
CLx QUANTITATIVE WESTERN BLOTS HIGH SENSITIVITY WIDE, LINEAR DYNAMIC RANGE NO FILM OR DARKROOM WIDE RANGE OF APPLICATIONS
 CLx QUANTITATIVE WESTERN BLOTS HIGH SENSITIVITY WIDE, LINEAR DYNAMIC RANGE NO FILM OR DARKROOM WIDE RANGE OF APPLICATIONS 2 Odyssey Infrared Imaging Systems ODYSSEY CLx Infrared Imaging System The Standard
CLx QUANTITATIVE WESTERN BLOTS HIGH SENSITIVITY WIDE, LINEAR DYNAMIC RANGE NO FILM OR DARKROOM WIDE RANGE OF APPLICATIONS 2 Odyssey Infrared Imaging Systems ODYSSEY CLx Infrared Imaging System The Standard
An Activity in Computed Tomography
 Pre-lab Discussion An Activity in Computed Tomography X-rays X-rays are high energy electromagnetic radiation with wavelengths smaller than those in the visible spectrum (0.01-10nm and 4000-800nm respectively).
Pre-lab Discussion An Activity in Computed Tomography X-rays X-rays are high energy electromagnetic radiation with wavelengths smaller than those in the visible spectrum (0.01-10nm and 4000-800nm respectively).
Mascot Distiller Hot Topics
 Mascot Distiller Hot Topics 1 Mascot Distiller Hot Topics Quantitation FAQ from 2009 Label-free Quantitation Large Multi-File Projects Arg-Pro Conversion Automation This presentation will touch on some
Mascot Distiller Hot Topics 1 Mascot Distiller Hot Topics Quantitation FAQ from 2009 Label-free Quantitation Large Multi-File Projects Arg-Pro Conversion Automation This presentation will touch on some
Enriching Beads Oligo (dt) Magnetic Beads for mrna Purification
 Enriching Beads Oligo (dt) Magnetic Beads for mrna Purification Isolate the mrna transcriptome in 15 minutes User Guidance Enriching Biotechnology Rev. 1.0 October 25th. 2018 Why choose Enriching Beads
Enriching Beads Oligo (dt) Magnetic Beads for mrna Purification Isolate the mrna transcriptome in 15 minutes User Guidance Enriching Biotechnology Rev. 1.0 October 25th. 2018 Why choose Enriching Beads
MAKE SURE YOUR SLIDES ARE CLEAN (TOP & BOTTOM) BEFORE LOADING DO NOT LOAD SLIDES DURING SOFTWARE INITIALIZATION
 Olympus VS120-L100 Slide Scanner Standard Operating Procedure Startup 1) Red power bar switch (behind monitor) 2) Computer 3) Login: UserVS120 account (no password) 4) Double click: WAIT FOR INITIALIZATION
Olympus VS120-L100 Slide Scanner Standard Operating Procedure Startup 1) Red power bar switch (behind monitor) 2) Computer 3) Login: UserVS120 account (no password) 4) Double click: WAIT FOR INITIALIZATION
An Activity in Computed Tomography
 Pre-lab Discussion An Activity in Computed Tomography X-rays X-rays are high energy electromagnetic radiation with wavelengths smaller than those in the visible spectrum (0.01-10nm and 4000-800nm respectively).
Pre-lab Discussion An Activity in Computed Tomography X-rays X-rays are high energy electromagnetic radiation with wavelengths smaller than those in the visible spectrum (0.01-10nm and 4000-800nm respectively).
Introduction to BioImage Analysis
 Introduction to BioImage Analysis Qi Gao CellNetworks Math-Clinic core facility 22-23.02.2018 MATH- CLINIC Math-Clinic core facility Data analysis services on bioimage analysis & bioinformatics: 1-to-1
Introduction to BioImage Analysis Qi Gao CellNetworks Math-Clinic core facility 22-23.02.2018 MATH- CLINIC Math-Clinic core facility Data analysis services on bioimage analysis & bioinformatics: 1-to-1
ImageJ: Introduction to Image Analysis 3 May 2012 Jacqui Ross
 Biomedical Imaging Research Unit School of Medical Sciences Faculty of Medical and Health Sciences The University of Auckland Private Bag 92019 Auckland 1142, NZ Ph: 373 7599 ext. 87438 http://www.fmhs.auckland.ac.nz/sms/biru/.
Biomedical Imaging Research Unit School of Medical Sciences Faculty of Medical and Health Sciences The University of Auckland Private Bag 92019 Auckland 1142, NZ Ph: 373 7599 ext. 87438 http://www.fmhs.auckland.ac.nz/sms/biru/.
Amplitudes Variation of GPR Rebar Reflection Due to the Influence of Concrete Aggregate Scattering
 More Info at Open Access Database www.ndt.net/?id=18402 Amplitudes Variation of GPR Rebar Reflection Due to the Influence of Concrete Aggregate Scattering Thomas KIND Federal Institute for Materials Research
More Info at Open Access Database www.ndt.net/?id=18402 Amplitudes Variation of GPR Rebar Reflection Due to the Influence of Concrete Aggregate Scattering Thomas KIND Federal Institute for Materials Research
