Abstract. comment reviews reports deposited research refereed research interactions information
|
|
|
- Rosalyn Dixon
- 5 years ago
- Views:
Transcription
1 Research Sources of nonlinearity in cdna microarray expression measurements Latha Ramdas*, Kevin R Coombes, Keith Baggerly, Lynne Abruzzo, W Edward Highsmith, Tammy Krogmann, Stanley R Hamilton* and Wei Zhang* Addresses: *Departments of Pathology, Biostatistics, Biomathematics and Hematopathology, Cancer Genomics Core Laboratory, University of Texas M D Anderson Cancer Center, Houston, TX 773, USA. Department of Pathology, University of Maryland, College Park, MD 2742, USA. Correspondence: Wei Zhang. wzhang@mdanderson.org Published: 1 October 21 Genome Biology 21, 2(11):research The electronic version of this article is the complete one and can be found online at 21 Ramdas et al., licensee BioMed Central Ltd (Print ISSN 145-9; Online ISSN ) Abstract Background: A key assumption in the analysis of microarray data is that the quantified signal intensities are linearly related to the expression levels of the corresponding genes. To test this assumption, we experimentally examined the relationship between signal and expression for the two types of microarrays we most commonly encounter: radioactively labeled cdnas on nylon membranes and fluorescently labeled cdnas on glass slides. Results: We uncovered two sources of nonlinearity. The first, which led to discrepancies in analysis affecting the fluorescent signals, was signal quenching associated with excessive dye concentrations. The second, affecting the radioactive signals, was a nonlinear transformation of the raw data introduced by the scanner. Correction for this transformation was made by some, but not all, image-quantification software packages. Conclusions: The second type of nonlinearity is more troublesome, because it could not have been predicted a priori. Both types of nonlinearities were detected by simple dilution series, which we recommend as a quality-control step. Background DNA microarray technology allows the simultaneous analyses of thousands of genes [1-3]. There are two major platforms for cdna microarrays: membrane-based arrays (porous surfaces like nylon) and chemically coated glassbased arrays. In both cases, thousands of cdna fragments are robotically deposited on the substrate. The nylon membrane microarrays are hybridized with 32 P or 33 P-labeled cdna targets, and microarrays on glass are hybridized with fluorescent dye-labeled cdna targets. After hybridization, Received: 23 July 21 Revised: 3 August 21 Accepted: 11 September 21 the radioactive or fluorescent signal intensities are measured using a phosphorimager or laser scanner, respectively. The signal intensities are surrogates for the expression levels of the genes in the samples under testing and are used to make biological inferences. A key assumption in the analysis of microarray data is that the quantified signal intensities are linearly related to the expression of the corresponding genes in the target sample. We experimentally examined this relationship. Our investigations
2 2 Genome Biology Vol 2 No 11 Ramdas et al. uncovered two sources of nonlinearity: signal quenching and a nonlinear (square-root) transformation of the raw data introduced by the scanner. Users presented with the same image but using different software packages may arrive at quite different conclusions about levels of differential expression. In both cases, the nonlinearities were revealed by serial dilution experiments. Given the lack of an absolute scale for microarray measurements, we recommend serial dilution experiments as a quality-control step. Results Measurement of fluorescent signals from glass-based microarrays To assess response linearity on glass slides, we designed two dilution experiments. In the first, a set of serially diluted (factor of two) Cy3-labeled oligonucleotide samples ranging from g/2l was arrayed on a slide in a 2 x 5 grid of x patches. Each row within a patch was a serial dilution; each patch contained eight replicates of the dilution. The slide was scanned with a laser scanner, and the image obtained (Figure 1a) was analyzed using the ArrayVision quantification software. If spot intensity is linearly related to the amount of labeled cdna, then a plot of the log (base 2) background-corrected signal intensity as a function of the serial dilution steps should have a slope of -1.. However, at this concentration range, a slope of -.4 was observed (Figure 1b). This means that when the concentration was halved, the intensity was consistently decreased by some factor less than one half. The total drop (across eight dilution levels) in concentration was 12-fold; the total drop in observed intensity was roughly nine-fold. This result is not surprising because fluorescence quenching is known to play a major role when the fluorescent material is present at such high concentrations. Quenching occurs when large numbers of fluorophores are highly concentrated so that photons emitted by one molecule can be reabsorbed by another molecule, thus artificially decreasing the detected signal [4]. A second experiment was designed using both Cy3- and Cy5- labeled oligonucleotides spotted in a much lower concentration range: from.1 to.7 2g/ml. Signals in this range were detected using scanning parameters similar to those we normally use for hybridization experiments. The two images are shown in Figure 2. The images (with ten separate patches and eight replicate sets in each patch representing replicates) were quantified with ArrayVision and plotted against the dilution steps. The average slope of the mean line for the Cy5-labeled oligonucleotide was -.9 and -.7 for Cy3-labeled oligonucleotide (Figure 3). A linear relationship is seen for both Cy3 and Cy5 in this concentration range, and the slopes are close to the expected value of -1.. The lack of perfect correlation to the actual signal intensity may be a result of quenching, and Cy3 may have more quenching effect than Cy5. Measurement of radioactive signals on a membrane array To assess response linearity on nylon, we carried out a dilution experiment where a serially diluted known amount of 32 P-ATP was spotted on a nylon membrane. After being exposed to a phosphorimager screen and measured by a STORM PhosphorImager, a GEL image file (Figure 4a) was produced and the signals analyzed using the ImageQuant analysis software (Molecular Dynamics, Sunnyvale, CA). As expected, the signals were linearly related to the powers of ½ that is, to 1, ½, ¼, and so on (Figure 5a). This result indicates that the ImageQuant software gives accurate readings of the signals from a GEL image file. However, ImageQuant was not designed to handle high-density microarray images that contain closely spaced spots. Thus, microarray images produced by the STORM PhosphorImager are often quantified using other software. We requantified our image file using two commercial software packages: GLEAMS and ArrayVision. The membrane had been scanned by the STORM Phosphor- Imager at a 45 angle (not by design). Because neither GLEAMS nor ArrayVision cope well with microarray images at this angle, we loaded the GEL file into ImageJ [5], an image-editing program available from the National Institutes of Health (NIH). We rotated and cropped the image, and saved it as a Tagged Image Format File (TIFF) (Figure 4b) which was loaded into both commercial software packages. The results from both packages indicated that the signal intensities were proportional to the square root of the true concentrations (Figure 5), in disagreement with both theory and the ImageQuant results. In fact, the pixel-bypixel intensity data are square-root-transformed before being saved as a GEL file. When an image-editing program (such as ImageJ) processes these data, tags describing this transformation are not preserved in the resulting TIFF file. To determine whether this square-root transformation could affect the results of a microarray experiment, we performed a hybridization experiment using a Research Genetics GF2 GeneFilter. Messenger RNA extracted from a GA-1 Burkitt lymphoma cell line was radioactively labeled with 33 P during reverse transcription, hybridized to the GF 2 GeneFilter, and exposed to a Molecular Dynamics STORM PhosphorImager. The image was saved as a GEL file, which was loaded directly into both GLEAMS and ArrayVision, without transforming it to a TIFF file. Each package quantified the mean intensity and the local background intensity at each spot and the results were compared graphically (Figure ). In each subpanel, the horizontal axis is the intensity reported by ArrayVision and the vertical axis is the result reported by GLEAMS. The most striking feature of Figure is that the most reliably measured spots - the spots where both software packages identify a gene that is expressed at high level - are not linearly related. The nonlinear relationship between the results was estimated by trial
3 Patch replicate dilution series, patch Dilution stage, 1 = initial, = most dilute dilution levels Replicate (row) number Figure 1 Dilution experiment with fluorescently labeled samples. A patch from the first dilution experiment with Cy3-labeled oligonucleotides on glass, showing eight replicates of a serial dilution. Log 2 background-corrected intensity values for the spots in the patch, going from left to right (dilution stages) and from top to bottom (replicates). The linear decrease shows that intensity is dropping as a power of concentration, but the slope suggests that this power is not what is expected. Note the consistency across replicates. svol is the background-corrected volume, where volume is the density of each spot multiplied by its area, and density represents the average of all the pixels in the spot. 2
4 4 Genome Biology Vol 2 No 11 Ramdas et al. Figure 2 Fluorescence dilution experiment at a lower concentration range. The second dilution experiment on glass, with much lower concentrations of dye used. Cy3; Cy5. The array design is a 2 x 5 grid of x patches. Each row in a patch is a serial dilution; thus the serial dilution has been repeated times on this glass slide. The spots are about 4 2m apart and about 2 2m in diameter. The Cy3 image was scanned at a gain of 45 and the Cy5 image at a gain of 5, as commonly done for the Cy3 and Cy5 images in our laboratory. and error to be y =15ax. The results reported by the two software packages followed the same curve on ten additional GEL image files (data not shown). Discussion This study was conducted to assess the response linearity of measurements from cdna microarray experiments using the two most frequently used systems. The study was performed not only because of the general need for quality control, but also because of the complexity of the process of acquiring data from microarrays. Images and data are often transferred between different computer programs, and many instruments used for microarray research are new and insufficiently tested. Thus, it is rather optimistic to take the numbers generated from a series of machines and software at face value. Simple dilution experiments revealed problems that have implications for the biological interpretation of gene expression data produced from microarray experiments. Our experiments on glass provided an assessment of the degree of signal quenching for the two fluorescent/glass microarrays. In dilute solutions fluorescence intensity is linearly proportional to the concentration with all other parameters being constant. However, in a sample with absorbance exceeding.5 at the emission wavelength, the relationship becomes nonlinear and the measurements are distorted (by self absorption, inner filter effect, quenching) [4,]. Fluorescence properties of such labeled DNA probes have been studied [7,]. Our experiments on membranes provide instances where different microarray-specific image-analysis programs were applied to the same images and produced divergent results. In each instance, at least one of the software packages produced results that were linearly related to the square root of the results produced by another package. The significance of this finding for the biological interpretation of gene expressions is very clear. Where users of software package 1 might detect, for example, a four-fold change in gene expression, users of software package 2 would see only a two-fold change. If two-fold change is set as a threshold, the same Dilution steps Dilution steps Figure 3 Log intensities plot as a function of concentration. Background-corrected log-intensities with the lower dye concentrations. Cy3; Cy5. In both cases, the slopes are much closer to the expected value of -1.
5 data can be viewed as significant or insignificant, depending on which software package is used. ImageQuant intensity Figure 4 Dilution experiment with radiolabeled samples. TIFF images of the radioactively labeled dilution experiment before and after rotation and cropping. (c) 1, 2, 3, 4,..4. Theoretical intensity ArrayVision intensity 5, 15, The explanation for the divergent results in our experiments is simple: the hardware (scanner) applied a mathematical transformation to the data before writing them to the image file. The nature of this transformation was not communicated to the software (image-quantifying program) that analyzed the data. Consequently, the software assumed (incorrectly) that the values in the file were linearly related to the original intensity levels. In our case, the STORM PhosphorImager produced a GEL file. This file contained numerical values for each pixel, which need to be squared to exhibit the proper linear relationship. The problem lies with the fact that the internal structure of a GEL image file is essentially identical to that of a TIFF image file, so any program that can read a TIFF file can read a GEL file and even manipulate the contents as if it were a TIFF file. But, if the file is then saved as a TIFF file, its GEL file origins are lost. This leads to two scenarios for bad data. In the first scenario, a GEL file is loaded into two software packages. Software package 1 recognizes that a GEL file includes a nonlinear transformation and corrects for it. Software package 2 treats the GEL file as a TIFF file and does not correct for nonlinearity. The results from the two packages therefore disagree. In the second scenario, the GEL file is saved as a TIFF file after editing. Software package 1, which correctly dealt with a GEL file, now sees a TIFF file and applies no transformation because none is generally needed for TIFF files. Software package 2 sees a TIFF file and deals 2,, 1, GLEAMS intensity Theoretical intensity Figure 5 Comparison of software packages for image analysis. Each panel shows theoretical intensity levels (as fractions of the initial concentration in the dilution series) or measured intensity values (in arbitrary units) for the radioactively labeled dilution experiment. ImageQuant was used on the original GEL file; GLEAMS and ArrayVision were used on the TIFF file produced from the GEL file by rotation and cropping. ImageQuant values are linearly related to the theoretical values. The values reported by GLEAMS and ArrayVision are linearly related. (c) The values reported by ArrayVision are not linearly related to the theoretical values. In this case, both ArrayVision and GLEAMS provide measurements related to the square root of the theoretical values. ArrayVision intensity 5, 15,..4.
6 Vol 2 No 11 Intensities 1, 2, 3, 4, 2 15 x 2, 4,,, (c) (d) Close-up of intensities Log intensities 2, 4,,, 1, 12, 1 14 Backgrounds Ramdas et al. 1, 1,4 Genome Biology 4 1 2, 4,,, 1, Figure Comparison of signal intensity and background intensity. (a-d) Intensity and background values produced by software packages 1 and 2 (ArrayVision and GLEAMS respectively) were applied to the GEL file produced by the follow-up experiment with radioactive labeling. ArrayVision finds the appropriate transformation; GLEAMS does not. with it as before. The results from the two packages now agree, but both are wrong because we have removed the information the packages need to perform correctly. It is worth pointing out another common instance where a square-root transformation is applied to microarray data. In a two-color fluorescence experiment, the microarray is scanned twice, at different wavelengths corresponding to the different dyes used in the assay. Each scan is saved as a separate 1-bit grayscale image. It is possible to combine the two grayscale images into a single 24-bit color image, sometimes called a false-color image. One simply imports the first image into the red channel and the second image into the green channel. However, a 24-bit full color image allocates only bits to each channel. In order to pack a 1-bit number representing the scanned intensity into an -bit space, some information must be discarded. For instance, the software operating the GenePix 4A Microarray Scanner (Axon Instruments, Foster City, CA) provides four packing options (note that the Axon manual says that packing is a bad idea if investigators want to get numbers from the image later). The default option is to perform a square-root operation. The remaining options preserve linearity, but truncate the data, either by preserving low values, preserving high values or preserving middle values. Although it is tempting to discard the two grayscale images and save only the full-color image, doing so would unavoidably discard essential aspects of the data. The primary data produced by a microarray experiment is the original scanned image, which is stored as a computer file. Any processing of this image file has the potential to change, lose or otherwise corrupt data. We have seen that square-root transformations are incorporated in some programs. All general-purpose image-editing programs provide multitudes of additional transformations that can be used to
7 brighten, sharpen or smooth images. Even though the square-root transformation appears to be the only transformation in common use among current generations of scanners, it is conceivable that other transformations may be introduced in the future. In summary, when designing a protocol for a set of microarray experiments, researchers should perform dilution series as one of their standard calibration experiments. Processing of the array through the scanner and quantification software that will be used in the experiments can confirm that the reported results are linearly related to the known input values. Methods Fluorescent labeling For the experiments on glass, cyanine 3-labeled (Cy3), cyanine 5-labeled (Cy5) [7] and unlabeled 3mer oligonucleotides were synthesized (Synthegen, Houston, TX). Plain glass slides from Fisher Scientific were coated with polylysine according to the published procedure [9]. An arrayer from Genomic Solutions (Ann Arbor, MI) was used to spot the oligonucleotides onto the treated glass slides. A 4-pin head from Genomic Solutions was used to create an array design of a 2 x 5 grid of x patches with a spot spacing of about 4 2m. The slides were scanned on a GeneTac LS IV laser scanner (Genomic Solutions) with laser energy sources for measuring Cy3 and Cy5 fluorophore. Data from the dual-lasers are collected as separate TIFF files for each of the two lasers. The images were processed using the analysis software program ArrayVision, version 5.1 (Imaging Research, Inc., St Catherine s, Ontario, Canada) and GLEAMS version 2. (NuTec Sciences, Houston, TX). Background-corrected intensity was determined for each element of each array. Radioactive labeling For the experiments on membranes, 1 2l 32 P-,-dATP stock solution (NEN Life Science Products, Inc., Boston, MA) was first diluted 1 times, then 5 2l of this mixture was diluted two-fold by adding 5 2l water. This process was repeated to generate a serial dilution. Next, 1 2l of each diluted sample was spotted onto a nylon membrane. After hybridization, the nylon membrane was exposed to the STORM PhosphorImager from Molecular Dynamics (Sunnyvale, CA), which produced a GEL image file. ImageQuant analysis software (Molecular Dynamics) was used to quantify the images. For the follow-up experiment, a GF2 Human GeneFilter microarray was purchased from Research Genetics (Huntsville, AL). Total RNA was isolated from a GA-1 Burkitt lymphoma cell line (a kind gift of Aaron Rapaport, University of Maryland). Ten 2g total RNA were reversetranscribed and 33 P-labeled following the standard procedure. The labeled cdnas were hybridized to the GeneFilter. Acknowledgements We thank Kenneth Hess, Jing Wang, Mini Kapoor and David Stivers for their critical comments on this study and Marla Bordelon for editorial assistance. This work was partially supported by Tobacco Settlement Funds appropriated to M.D. Anderson Cancer Center by the Texas Legislature, a donation from the Kadoorie Foundation, and a grant from Texas Higher Education Coordinating Board under grant number References 1. Lockhart DJ, Winzler EA: Genomics, gene expression and DNA arrays. Nature 2, 45: DeRisi JL, Iyer VR: Genomics and array technology. Curr Opin Oncol 1999, 11: Khan J, Saal LH, Bittner ML, Chen Y, Trent JM, Meltzer PS: Expression profiling in cancer using cdna microarrays. Electrophoresis 1999, 2: Kubista M: Experimental correction for the inner-filter effect in fluorescence spectra. Analyst 1994, 119: ImageJ [ Cantor CR, Schimmel PR: Biophysical Chemistry: Techniques for the Study of Biological Stucture and Function. San Francisco: W.H. Freeman and Co; Mujumdar RB, Ernst LA, Mujumdar SR, Lewis CJ, Waggoner AS: Cyanine dye labeling reagents: sulfoindocyanine succinimidyl esters. Bioconjug Chem 1993, 4: Randolph JB, Waggoner AS: Stability, specificity and fluorescence brightness of multiply-labeled fluorescent DNA probes. Nucleic Acids Res 1997, 25: Schena M, Shalon D, Davis RW, Brown PO: Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science 1995, 27:47-47.
Automated cdna microarray image segmentation
 Automated cdna microarray image segmentation Author Liew, Alan Wee-Chung, Yan, Hong Published 2007 Conference Title Proceedings of the International Symposium on Computational Models for Life Sciences
Automated cdna microarray image segmentation Author Liew, Alan Wee-Chung, Yan, Hong Published 2007 Conference Title Proceedings of the International Symposium on Computational Models for Life Sciences
Automatic gene expression estimation from microarray images. Daniel O. Dantas Adviser: : Junior Barrera
 Automatic gene expression estimation from microarray images Daniel O. Dantas Adviser: : Junior Barrera IME-USP Summary Introduction Problem definition Solution strategy Image segmentation Signal estimation
Automatic gene expression estimation from microarray images Daniel O. Dantas Adviser: : Junior Barrera IME-USP Summary Introduction Problem definition Solution strategy Image segmentation Signal estimation
GenePix Application Note
 GenePix Application Note Determining the Signal-to-Noise Ratio and Optimal Photomultiplier gain setting in the GenePix 4000B Siobhan Pickett, M.S., Sean Carriedo, Ph.D. and Chang Wang, Ph.D. Axon Instruments,
GenePix Application Note Determining the Signal-to-Noise Ratio and Optimal Photomultiplier gain setting in the GenePix 4000B Siobhan Pickett, M.S., Sean Carriedo, Ph.D. and Chang Wang, Ph.D. Axon Instruments,
GenePix Application Note
 GenePix Application Note Biological Relevance of GenePix Results Shawn Handran, Ph.D. and Jack Y. Zhai, Ph.D. Axon Instruments, Inc. 3280 Whipple Road, Union City, CA 94587 Last Updated: Aug 22, 2003.
GenePix Application Note Biological Relevance of GenePix Results Shawn Handran, Ph.D. and Jack Y. Zhai, Ph.D. Axon Instruments, Inc. 3280 Whipple Road, Union City, CA 94587 Last Updated: Aug 22, 2003.
Quality control of microarrays
 Quality control of microarrays Solveig Mjelstad Angelskår Intoduction to Microarray technology September 2009 Overview of the presentation 1. Image analysis 2. Quality Control (QC) general concepts 3.
Quality control of microarrays Solveig Mjelstad Angelskår Intoduction to Microarray technology September 2009 Overview of the presentation 1. Image analysis 2. Quality Control (QC) general concepts 3.
Product Information. Introduction
 Microarray Scanner Calibration Slide To quantitatively analyze scanners performance and output, adjust and fine-tune scanners, and perform comparative analysis for multiple scanner units. To verify scanners
Microarray Scanner Calibration Slide To quantitatively analyze scanners performance and output, adjust and fine-tune scanners, and perform comparative analysis for multiple scanner units. To verify scanners
Scanning and Image Processing -by Steve Clough
 Scanning and Image Processing -by Steve Clough cdna microarrays use two dyes with well separated emission spectra such as Cy3 and Cy5 to allow direct comparisons on single slide GSI Lumonics Ratio of Expression
Scanning and Image Processing -by Steve Clough cdna microarrays use two dyes with well separated emission spectra such as Cy3 and Cy5 to allow direct comparisons on single slide GSI Lumonics Ratio of Expression
Image Capture TOTALLAB
 1 Introduction In order for image analysis to be performed on a gel or Western blot, it must first be converted into digital data. Good image capture is critical to guarantee optimal performance of automated
1 Introduction In order for image analysis to be performed on a gel or Western blot, it must first be converted into digital data. Good image capture is critical to guarantee optimal performance of automated
Influence of Dictionary Size on the Lossless Compression of Microarray Images
 Influence of Dictionary Size on the Lossless Compression of Microarray Images Robert Bierman 1, Rahul Singh 1 Department of Computer Science, San Francisco State University, San Francisco, CA bierman@sfsu.edu,
Influence of Dictionary Size on the Lossless Compression of Microarray Images Robert Bierman 1, Rahul Singh 1 Department of Computer Science, San Francisco State University, San Francisco, CA bierman@sfsu.edu,
How is the Digital Image Generated? Image Acquisition Devices
 In order for image analysis to be performed on a 2D gel, it must first be converted into digital data. Good image capture is critical to guarantee optimal performance of automated image analysis packages
In order for image analysis to be performed on a 2D gel, it must first be converted into digital data. Good image capture is critical to guarantee optimal performance of automated image analysis packages
Developed by BioDiscovery, Inc. DualChip evaluation software User Manual Version 1.1
 Developed by BioDiscovery, Inc. DualChip evaluation software User Manual Version 1.1 1 Table of contents 1. INTRODUCTION...3 2. SCOPE OF DELIVERY...4 3. INSTALLATION PROCEDURES...5 3.1. SYSTEM REQUIREMENTS...
Developed by BioDiscovery, Inc. DualChip evaluation software User Manual Version 1.1 1 Table of contents 1. INTRODUCTION...3 2. SCOPE OF DELIVERY...4 3. INSTALLATION PROCEDURES...5 3.1. SYSTEM REQUIREMENTS...
Preparation of Sample Hybridization Scanning and Image Analysis
 Preparation of Sample Hybridization Scanning and Image Analysis Sample preparation 1. Design experiment 2. Perform experiment Question? Replicates? Test? mutant wild type 3. Precipitate RNA 4. Label RNA
Preparation of Sample Hybridization Scanning and Image Analysis Sample preparation 1. Design experiment 2. Perform experiment Question? Replicates? Test? mutant wild type 3. Precipitate RNA 4. Label RNA
Development of a Next-Generation Laser-Scanner System for Life Science Research
 Development of a Next-Generation Laser-Scanner System for Life Science Research Masaki TAKAMATSU* Yasutake TANAKA* Takashi KOBAYASHI* Hiromi ISHIKAWA* and Akira YAMAGUCHI* Abstract We developed a next-generation
Development of a Next-Generation Laser-Scanner System for Life Science Research Masaki TAKAMATSU* Yasutake TANAKA* Takashi KOBAYASHI* Hiromi ISHIKAWA* and Akira YAMAGUCHI* Abstract We developed a next-generation
ScanArray Overview. Principle of Operation. Instrument Components
 ScanArray Overview The GSI Lumonics ScanArrayÒ Microarray Analysis System is a scanning laser confocal fluorescence microscope that is used to determine the fluorescence intensity of a two-dimensional
ScanArray Overview The GSI Lumonics ScanArrayÒ Microarray Analysis System is a scanning laser confocal fluorescence microscope that is used to determine the fluorescence intensity of a two-dimensional
Instructions for Howto Scan µarrays
 Instructions for Howto Scan µarrays Introduction After probing the µarray slides with samples, one is now ready to scan them. To scan a µarrays slide is too convert the biological information trapped on
Instructions for Howto Scan µarrays Introduction After probing the µarray slides with samples, one is now ready to scan them. To scan a µarrays slide is too convert the biological information trapped on
In our previous lecture, we understood the vital parameters to be taken into consideration before data acquisition and scanning.
 Interactomics: Protein Arrays & Label Free Biosensors Professor Sanjeeva Srivastava MOOC NPTEL Course Indian Institute of Technology Bombay Module 7 Lecture No 34 Software for Image scanning and data processing
Interactomics: Protein Arrays & Label Free Biosensors Professor Sanjeeva Srivastava MOOC NPTEL Course Indian Institute of Technology Bombay Module 7 Lecture No 34 Software for Image scanning and data processing
Scan slides (Axon Genepix 4200AL)
 Page 1 Scan slides (Axon Genepix 4200AL) We need to scan the slides on both channels (Cy3 and Cy5) to obtain a 16-bit grayscale TIFF file for each. Typically these files are about 20-26Mb per channel,
Page 1 Scan slides (Axon Genepix 4200AL) We need to scan the slides on both channels (Cy3 and Cy5) to obtain a 16-bit grayscale TIFF file for each. Typically these files are about 20-26Mb per channel,
Products - Microarray Scanners - Laser Scanners - InnoScan 900 Series and MAPIX Software
 Products - Microarray Scanners - Laser Scanners - InnoScan 900 Series and MAPIX Software Arrayit offers the world s only next generation microarray scanning technology, with proprietary rotary motion control,
Products - Microarray Scanners - Laser Scanners - InnoScan 900 Series and MAPIX Software Arrayit offers the world s only next generation microarray scanning technology, with proprietary rotary motion control,
EmbryoCellect. RHS Scanning and Analysis Instructions. for. Genepix Pro Software
 EmbryoCellect RHS Scanning and Analysis Instructions for Genepix Pro Software EmbryoCellect Genepix Pro Scanning and Analysis Technical Data Sheet Version 1.0 October 2015 1 Copyright Reproductive Health
EmbryoCellect RHS Scanning and Analysis Instructions for Genepix Pro Software EmbryoCellect Genepix Pro Scanning and Analysis Technical Data Sheet Version 1.0 October 2015 1 Copyright Reproductive Health
NPTEL VIDEO COURSE PROTEOMICS PROF. SANJEEVA SRIVASTAVA
 HANDOUT LECTURE-31 MICROARRAY WORK-FLOW: IMAGE SCANNING AND DATA PROCESSING Slide 1: This module contains the summary of the discussion with Mr. Pankaj Khanna, an application specialist from Spinco Biotech,
HANDOUT LECTURE-31 MICROARRAY WORK-FLOW: IMAGE SCANNING AND DATA PROCESSING Slide 1: This module contains the summary of the discussion with Mr. Pankaj Khanna, an application specialist from Spinco Biotech,
Computational Genomics. High-throughput experimental biology
 Computational Genomics 10-810/02 810/02-710, Spring 2009 Gene Expression Analysis Data pre-processing processing Eric Xing Lecture 15, March 4, 2009 Reading: class assignment Eric Xing @ CMU, 2005-2009
Computational Genomics 10-810/02 810/02-710, Spring 2009 Gene Expression Analysis Data pre-processing processing Eric Xing Lecture 15, March 4, 2009 Reading: class assignment Eric Xing @ CMU, 2005-2009
MICROARRAY IMAGE ANALYSIS PROGRAM
 Revision submitted for publication to Loyola Schools Review, 13 November 2002 MICROARRAY IMAGE ANALYSIS PROGRAM Paul Ignatius D. Echevarria Jerome C. Punzalan John Paul C. Vergara Department of Information
Revision submitted for publication to Loyola Schools Review, 13 November 2002 MICROARRAY IMAGE ANALYSIS PROGRAM Paul Ignatius D. Echevarria Jerome C. Punzalan John Paul C. Vergara Department of Information
NPTEL VIDEO COURSE PROTEOMICS PROF. SANJEEVA SRIVASTAVA
 LECTURE-31 MICROARRAY WORK-FLOW: IMAGE SACNNING AND DATA PROCESSING TRANSCRIPT Welcome to the proteomics course. In today s lecture we will talk about microarray work-flow the image scanning and processing.
LECTURE-31 MICROARRAY WORK-FLOW: IMAGE SACNNING AND DATA PROCESSING TRANSCRIPT Welcome to the proteomics course. In today s lecture we will talk about microarray work-flow the image scanning and processing.
Multiplexing as Essential Tool for Modern Biology
 Multiplexing as Essential Tool for Modern Biology Bio-Plex Seminar, Debrecen, 2012. Gyula Csanádi, PhD. The "Age of "-omics" Studying interrelationships at different level of complexity Genes - Unveiling
Multiplexing as Essential Tool for Modern Biology Bio-Plex Seminar, Debrecen, 2012. Gyula Csanádi, PhD. The "Age of "-omics" Studying interrelationships at different level of complexity Genes - Unveiling
III III 0 IIOI DID IIO 1101 I II 0II II 100 III IID II DI II
 (19) United States III III 0 IIOI DID IIO 1101 I0 1101 0II 0II II 100 III IID II DI II US 200902 19549A1 (12) Patent Application Publication (10) Pub. No.: US 2009/0219549 Al Nishizaka et al. (43) Pub.
(19) United States III III 0 IIOI DID IIO 1101 I0 1101 0II 0II II 100 III IID II DI II US 200902 19549A1 (12) Patent Application Publication (10) Pub. No.: US 2009/0219549 Al Nishizaka et al. (43) Pub.
Steps involved in microarray analysis after the experiments
 Steps involved in microarray analysis after the experiments Scanning slides to create images Conversion of images to numerical data Processing of raw numerical data Further analysis Clustering Integration
Steps involved in microarray analysis after the experiments Scanning slides to create images Conversion of images to numerical data Processing of raw numerical data Further analysis Clustering Integration
GALILEO TMA CK 4500 HTS Tissue Microarray Platform
 GALILEO TMA CK 4500 HTS Tissue Microarray Platform Tissue Microarray (TMA) A Block Of Samples From Hundreds Of Blocks (S. M. Hewitt, M.D., Ph.D., Tissue Array Research Program, LP, CCR, NCI, NIH) TMA technology
GALILEO TMA CK 4500 HTS Tissue Microarray Platform Tissue Microarray (TMA) A Block Of Samples From Hundreds Of Blocks (S. M. Hewitt, M.D., Ph.D., Tissue Array Research Program, LP, CCR, NCI, NIH) TMA technology
The Bead. beadarray: : An R Package for Illumina BeadArrays. Bead Preparation and Array Production. Beads in Wells. Mark Dunning -
 beadarray: : An R Package for Illumina BeadArrays Mark Dunning - md392@cam.ac.uk PhD Student - Computational Biology Group, Department of Oncology - University of Cambridge Address The Bead Probe 23 b
beadarray: : An R Package for Illumina BeadArrays Mark Dunning - md392@cam.ac.uk PhD Student - Computational Biology Group, Department of Oncology - University of Cambridge Address The Bead Probe 23 b
LSM 780 Confocal Microscope Standard Operation Protocol
 LSM 780 Confocal Microscope Standard Operation Protocol Basic Operation Turning on the system 1. Sign on log sheet according to Actual start time 2. Check Compressed Air supply for the air table 3. Switch
LSM 780 Confocal Microscope Standard Operation Protocol Basic Operation Turning on the system 1. Sign on log sheet according to Actual start time 2. Check Compressed Air supply for the air table 3. Switch
INSTRUMENTATION BREADBOARDING (VERSION 1.3)
 Instrumentation Breadboarding, Page 1 INSTRUMENTATION BREADBOARDING (VERSION 1.3) I. BACKGROUND The purpose of this experiment is to provide you with practical experience in building electronic circuits
Instrumentation Breadboarding, Page 1 INSTRUMENTATION BREADBOARDING (VERSION 1.3) I. BACKGROUND The purpose of this experiment is to provide you with practical experience in building electronic circuits
3) Start ImageJ, install CM Engine as a macro (instructions here:
 Instructions for CM Engine use 1) Download CM Engine from SourceForge (http://cm- engine.sourceforge.net/) or from the Rothstein Lab website (http://www.rothsteinlab.com/cm- engine.zip ). 2) Download ImageJ
Instructions for CM Engine use 1) Download CM Engine from SourceForge (http://cm- engine.sourceforge.net/) or from the Rothstein Lab website (http://www.rothsteinlab.com/cm- engine.zip ). 2) Download ImageJ
Microarray Data Pre-processing. Ana H. Barragan Lid
 Microarray Data Pre-processing Ana H. Barragan Lid Hybridized Microarray Imaged in a microarray scanner Scanner produces fluorescence intensity measurements Intensities correspond to levels of hybridization
Microarray Data Pre-processing Ana H. Barragan Lid Hybridized Microarray Imaged in a microarray scanner Scanner produces fluorescence intensity measurements Intensities correspond to levels of hybridization
Supporting Information: Electron Microscopic Visualization of Protein Assemblies on Flattened DNA Origami
 Supporting Information: Electron Microscopic Visualization of Protein Assemblies on Flattened DNA Origami Leena Mallik, Soma Dhakal, Joseph Nichols, Jacob Mahoney, Anne M. Dosey, Shuoxing Jiang ǂ, Roger
Supporting Information: Electron Microscopic Visualization of Protein Assemblies on Flattened DNA Origami Leena Mallik, Soma Dhakal, Joseph Nichols, Jacob Mahoney, Anne M. Dosey, Shuoxing Jiang ǂ, Roger
Low-level Analysis. cdna Microarrays. Lecture 2 Low Level Gene Expression Data Analysis. Stat 697K, CS 691K, Microbio 690K
 Lecture 2 Low Level Gene Expression Data nalysis Stat 697K, CS 691K, icrobio 690K Statistical Challenges odel variation of data not related to gene expression Compare expression for the same gene across
Lecture 2 Low Level Gene Expression Data nalysis Stat 697K, CS 691K, icrobio 690K Statistical Challenges odel variation of data not related to gene expression Compare expression for the same gene across
Microarray Image Analysis: Background Estimation using Region and Filtering Techniques
 Microarray Image Analysis: Background Estimation using Region and Filtering Techniques Anders Bengtsson December 9, 2003 Abstract This report examines properties of two main methods used for local background
Microarray Image Analysis: Background Estimation using Region and Filtering Techniques Anders Bengtsson December 9, 2003 Abstract This report examines properties of two main methods used for local background
STORM/ PALM ANSWER KEY
 STORM/ PALM ANSWER KEY Phys598BP Spring 2016 University of Illinois at Urbana-Champaign Questions for Lab Report 1. How do you define a resolution in STORM imaging? If you are given a STORM setup, how
STORM/ PALM ANSWER KEY Phys598BP Spring 2016 University of Illinois at Urbana-Champaign Questions for Lab Report 1. How do you define a resolution in STORM imaging? If you are given a STORM setup, how
ECEN. Spectroscopy. Lab 8. copy. constituents HOMEWORK PR. Figure. 1. Layout of. of the
 ECEN 4606 Lab 8 Spectroscopy SUMMARY: ROBLEM 1: Pedrotti 3 12-10. In this lab, you will design, build and test an optical spectrum analyzer and use it for both absorption and emission spectroscopy. The
ECEN 4606 Lab 8 Spectroscopy SUMMARY: ROBLEM 1: Pedrotti 3 12-10. In this lab, you will design, build and test an optical spectrum analyzer and use it for both absorption and emission spectroscopy. The
Game Mechanics Minesweeper is a game in which the player must correctly deduce the positions of
 Table of Contents Game Mechanics...2 Game Play...3 Game Strategy...4 Truth...4 Contrapositive... 5 Exhaustion...6 Burnout...8 Game Difficulty... 10 Experiment One... 12 Experiment Two...14 Experiment Three...16
Table of Contents Game Mechanics...2 Game Play...3 Game Strategy...4 Truth...4 Contrapositive... 5 Exhaustion...6 Burnout...8 Game Difficulty... 10 Experiment One... 12 Experiment Two...14 Experiment Three...16
Feature Level Data. Outline. Affymetrix GeneChip Design. Affymetrix GeneChip arrays Two color platforms
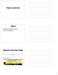 Feature Level Data Outline Affymetrix GeneChip arrays Two color platforms Affymetrix GeneChip Design 5 3 Reference sequence TGTGATGGTGCATGATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT GTACTACCCAGTCTTCCGGAGGCTA
Feature Level Data Outline Affymetrix GeneChip arrays Two color platforms Affymetrix GeneChip Design 5 3 Reference sequence TGTGATGGTGCATGATGGGTCAGAAGGCCTCCGATGCGCCGATTGAGAAT GTACTACCCAGTCTTCCGGAGGCTA
Analysing data from Illumina BeadArrays
 The bead Analysing data from Illumina BeadArrays Each silica bead is 3 microns in diameter Matt Ritchie Department of Oncology University of Cambridge, UK 4th September 008 700,000 copies of same probe
The bead Analysing data from Illumina BeadArrays Each silica bead is 3 microns in diameter Matt Ritchie Department of Oncology University of Cambridge, UK 4th September 008 700,000 copies of same probe
Introduction to Image Analysis with
 Introduction to Image Analysis with PLEASE ENSURE FIJI IS INSTALLED CORRECTLY! WHAT DO WE HOPE TO ACHIEVE? Specifically, the workshop will cover the following topics: 1. Opening images with Bioformats
Introduction to Image Analysis with PLEASE ENSURE FIJI IS INSTALLED CORRECTLY! WHAT DO WE HOPE TO ACHIEVE? Specifically, the workshop will cover the following topics: 1. Opening images with Bioformats
ImageJ, A Useful Tool for Image Processing and Analysis Joel B. Sheffield
 ImageJ, A Useful Tool for Image Processing and Analysis Joel B. Sheffield Temple University Dedicated to the memory of Dan H. Moore (1909-2008) Presented at the 2008 meeting of the Microscopy and Microanalytical
ImageJ, A Useful Tool for Image Processing and Analysis Joel B. Sheffield Temple University Dedicated to the memory of Dan H. Moore (1909-2008) Presented at the 2008 meeting of the Microscopy and Microanalytical
Dr. Bob on Colocalization or MSL Experiments In Learning Colocalization Using Image J
 Dr. Bob on Colocalization or MSL Experiments In Learning Colocalization Using Image J Confocal microscopy is used to test whether two fluorescently labeled molecules are associated with one another. If
Dr. Bob on Colocalization or MSL Experiments In Learning Colocalization Using Image J Confocal microscopy is used to test whether two fluorescently labeled molecules are associated with one another. If
Spotxel 1.7 Microarray Image and Data Analysis Software User s Guide
 Spotxel 1.7 Microarray Image and Data Analysis Software User s Guide 27 April 2017 - Rev 7 Spotxel is only intended for research and not intended or approved for diagnosis of disease in humans or animals.
Spotxel 1.7 Microarray Image and Data Analysis Software User s Guide 27 April 2017 - Rev 7 Spotxel is only intended for research and not intended or approved for diagnosis of disease in humans or animals.
Seishi IKAMI* Takashi KOBAYASHI** Yasutake TANAKA* and Akira YAMAGUCHI* Abstract. 2. System configuration. 1. Introduction
 Development of a Next-generation CCD Imager for Life Sciences Research Seishi IKAMI* Takashi KOBAYASHI** Yasutake TANAKA* and Akira YAMAGUCHI* Abstract We have developed a next-generation CCD-based imager
Development of a Next-generation CCD Imager for Life Sciences Research Seishi IKAMI* Takashi KOBAYASHI** Yasutake TANAKA* and Akira YAMAGUCHI* Abstract We have developed a next-generation CCD-based imager
IncuCyte ZOOM Fluorescent Processing Overview
 IncuCyte ZOOM Fluorescent Processing Overview The IncuCyte ZOOM offers users the ability to acquire HD phase as well as dual wavelength fluorescent images of living cells producing multiplexed data that
IncuCyte ZOOM Fluorescent Processing Overview The IncuCyte ZOOM offers users the ability to acquire HD phase as well as dual wavelength fluorescent images of living cells producing multiplexed data that
SEPTEMBER VOL. 38, NO. 9 ELECTRONIC DEFENSE SIMULTANEOUS SIGNAL ERRORS IN WIDEBAND IFM RECEIVERS WIDE, WIDER, WIDEST SYNTHETIC APERTURE ANTENNAS
 r SEPTEMBER VOL. 38, NO. 9 ELECTRONIC DEFENSE SIMULTANEOUS SIGNAL ERRORS IN WIDEBAND IFM RECEIVERS WIDE, WIDER, WIDEST SYNTHETIC APERTURE ANTENNAS CONTENTS, P. 10 TECHNICAL FEATURE SIMULTANEOUS SIGNAL
r SEPTEMBER VOL. 38, NO. 9 ELECTRONIC DEFENSE SIMULTANEOUS SIGNAL ERRORS IN WIDEBAND IFM RECEIVERS WIDE, WIDER, WIDEST SYNTHETIC APERTURE ANTENNAS CONTENTS, P. 10 TECHNICAL FEATURE SIMULTANEOUS SIGNAL
Chem466 Lecture Notes. Spring, 2004
 Chem466 Lecture Notes Spring, 004 Overview of the course: Many of you will use instruments for chemical analyses in lab. settings. Some of you will go into careers (medicine, pharmacology, forensic science,
Chem466 Lecture Notes Spring, 004 Overview of the course: Many of you will use instruments for chemical analyses in lab. settings. Some of you will go into careers (medicine, pharmacology, forensic science,
Locating Molecules Using GSD Technology Project Folders: Organization of Experiment Files...1
 .....................................1 1 Project Folders: Organization of Experiment Files.................................1 2 Steps........................................................................2
.....................................1 1 Project Folders: Organization of Experiment Files.................................1 2 Steps........................................................................2
TO PLOT OR NOT TO PLOT?
 Graphic Examples This document provides examples of a number of graphs that might be used in understanding or presenting data. Comments with each example are intended to help you understand why the data
Graphic Examples This document provides examples of a number of graphs that might be used in understanding or presenting data. Comments with each example are intended to help you understand why the data
A Hue-Based Method for ph Determination
 University of Portland Pilot Scholars Chemistry Undergraduate Publications, Presentations and Projects Chemistry Spring 2015 A Hue-Based Method for ph Determination Blair Pearson Kelly Ramzy Ryan Bergio
University of Portland Pilot Scholars Chemistry Undergraduate Publications, Presentations and Projects Chemistry Spring 2015 A Hue-Based Method for ph Determination Blair Pearson Kelly Ramzy Ryan Bergio
SmartDoc 2.0 E5001-SDB Instruction Manual
 SmartDoc 2.0 E5001-SDB Instruction Manual Version 11.16 1 Table of Contents 1. Introduction 3 2. Warnings. 3 3. Unpacking.. 4 4. SmartDoc 2.0 Overview 4 5. Setting up the SmartDoc 2.0 5 6. Gel Viewing
SmartDoc 2.0 E5001-SDB Instruction Manual Version 11.16 1 Table of Contents 1. Introduction 3 2. Warnings. 3 3. Unpacking.. 4 4. SmartDoc 2.0 Overview 4 5. Setting up the SmartDoc 2.0 5 6. Gel Viewing
LSM 710 Confocal Microscope Standard Operation Protocol
 LSM 710 Confocal Microscope Standard Operation Protocol Basic Operation Turning on the system 1. Switch on Main power switch 2. Switch on System / PC power button 3. Switch on Components power button 4.
LSM 710 Confocal Microscope Standard Operation Protocol Basic Operation Turning on the system 1. Switch on Main power switch 2. Switch on System / PC power button 3. Switch on Components power button 4.
(Quantitative Imaging for) Colocalisation Analysis
 (Quantitative Imaging for) Colocalisation Analysis or Why Colour Merge / Overlay Images are EVIL! Special course for DIGS-BB PhD program What is an Image anyway..? An image is a representation of reality
(Quantitative Imaging for) Colocalisation Analysis or Why Colour Merge / Overlay Images are EVIL! Special course for DIGS-BB PhD program What is an Image anyway..? An image is a representation of reality
Laser Beam Analysis Using Image Processing
 Journal of Computer Science 2 (): 09-3, 2006 ISSN 549-3636 Science Publications, 2006 Laser Beam Analysis Using Image Processing Yas A. Alsultanny Computer Science Department, Amman Arab University for
Journal of Computer Science 2 (): 09-3, 2006 ISSN 549-3636 Science Publications, 2006 Laser Beam Analysis Using Image Processing Yas A. Alsultanny Computer Science Department, Amman Arab University for
National Science Education Standards, Content Standard 5-8, Correlation with IPS and FM&E
 National Science Education Standards, Content Standard 5-8, Correlation with and Standard Science as Inquiry Fundamental Concepts Scientific Principles Abilities necessary to do Identify questions that
National Science Education Standards, Content Standard 5-8, Correlation with and Standard Science as Inquiry Fundamental Concepts Scientific Principles Abilities necessary to do Identify questions that
Laboratory 1: Uncertainty Analysis
 University of Alabama Department of Physics and Astronomy PH101 / LeClair May 26, 2014 Laboratory 1: Uncertainty Analysis Hypothesis: A statistical analysis including both mean and standard deviation can
University of Alabama Department of Physics and Astronomy PH101 / LeClair May 26, 2014 Laboratory 1: Uncertainty Analysis Hypothesis: A statistical analysis including both mean and standard deviation can
The Next Generation Science Standards Grades 6-8
 A Correlation of The Next Generation Science Standards Grades 6-8 To Oregon Edition A Correlation of to Interactive Science, Oregon Edition, Chapter 1 DNA: The Code of Life Pages 2-41 Performance Expectations
A Correlation of The Next Generation Science Standards Grades 6-8 To Oregon Edition A Correlation of to Interactive Science, Oregon Edition, Chapter 1 DNA: The Code of Life Pages 2-41 Performance Expectations
Fast Raman Spectral Imaging Using Chirped Femtosecond Lasers
 Fast Raman Spectral Imaging Using Chirped Femtosecond Lasers Dan Fu 1, Gary Holtom 1, Christian Freudiger 1, Xu Zhang 2, Xiaoliang Sunney Xie 1 1. Department of Chemistry and Chemical Biology, Harvard
Fast Raman Spectral Imaging Using Chirped Femtosecond Lasers Dan Fu 1, Gary Holtom 1, Christian Freudiger 1, Xu Zhang 2, Xiaoliang Sunney Xie 1 1. Department of Chemistry and Chemical Biology, Harvard
JCB Feature. What s in a picture? The temptation of image manipulation. The Journal of Cell Biology
 JCB Feature What s in a picture? The temptation of image manipulation Mike Rossner 1 and Kenneth M. Yamada 2 1 Managing Editor, The Journal of Cell Biology 2 Editor, The Journal of Cell Biology, and the
JCB Feature What s in a picture? The temptation of image manipulation Mike Rossner 1 and Kenneth M. Yamada 2 1 Managing Editor, The Journal of Cell Biology 2 Editor, The Journal of Cell Biology, and the
Multi-channel imaging cytometry with a single detector
 Multi-channel imaging cytometry with a single detector Sarah Locknar 1, John Barton 1, Mark Entwistle 2, Gary Carver 1 and Robert Johnson 1 1 Omega Optical, Brattleboro, VT 05301 2 Philadelphia Lightwave,
Multi-channel imaging cytometry with a single detector Sarah Locknar 1, John Barton 1, Mark Entwistle 2, Gary Carver 1 and Robert Johnson 1 1 Omega Optical, Brattleboro, VT 05301 2 Philadelphia Lightwave,
Resting pulse After exercise Resting pulse After exercise. Trial Trial Trial Trial. Subject Subject
 EXERCISE 2.3 Data Presentation Objectives After completing this exercise, you should be able to 1. Explain the difference between discrete and continuous variables and give examples. 2. Use one given data
EXERCISE 2.3 Data Presentation Objectives After completing this exercise, you should be able to 1. Explain the difference between discrete and continuous variables and give examples. 2. Use one given data
Experiment G: Introduction to Graphical Representation of Data & the Use of Excel
 Experiment G: Introduction to Graphical Representation of Data & the Use of Excel Scientists answer posed questions by performing experiments which provide information about a given problem. After collecting
Experiment G: Introduction to Graphical Representation of Data & the Use of Excel Scientists answer posed questions by performing experiments which provide information about a given problem. After collecting
Assessments Using Spike-In Experiments
 Assessments Using Spike-In Experiments Rafael A Irizarry, Department of Biostatistics JHU rafa@jhu.edu http://www.biostat.jhsph.edu/~ririzarr http://www.bioconductor.org A probe set = 11-20 PM,MM pairs
Assessments Using Spike-In Experiments Rafael A Irizarry, Department of Biostatistics JHU rafa@jhu.edu http://www.biostat.jhsph.edu/~ririzarr http://www.bioconductor.org A probe set = 11-20 PM,MM pairs
Evaluating Commercial Scanners for Astronomical Images. The underlying technology of the scanners: Pixel sizes:
 Evaluating Commercial Scanners for Astronomical Images Robert J. Simcoe Associate Harvard College Observatory rjsimcoe@cfa.harvard.edu Introduction: Many organizations have expressed interest in using
Evaluating Commercial Scanners for Astronomical Images Robert J. Simcoe Associate Harvard College Observatory rjsimcoe@cfa.harvard.edu Introduction: Many organizations have expressed interest in using
Instructions for Mapping * µarray Images using GenePix 5.0
 Instructions for Mapping * µarray Images using GenePix 5.0 Preliminary Information Make sure that the GenePix 5.0 software has been installed on your computer and you have the USB hardware dongle that
Instructions for Mapping * µarray Images using GenePix 5.0 Preliminary Information Make sure that the GenePix 5.0 software has been installed on your computer and you have the USB hardware dongle that
Supplementary Information. Stochastic Optical Reconstruction Microscopy Imaging of Microtubule Arrays in Intact Arabidopsis thaliana Seedling Roots
 Supplementary Information Stochastic Optical Reconstruction Microscopy Imaging of Microtubule Arrays in Intact Arabidopsis thaliana Seedling Roots Bin Dong 1,, Xiaochen Yang 2,, Shaobin Zhu 1, Diane C.
Supplementary Information Stochastic Optical Reconstruction Microscopy Imaging of Microtubule Arrays in Intact Arabidopsis thaliana Seedling Roots Bin Dong 1,, Xiaochen Yang 2,, Shaobin Zhu 1, Diane C.
Enhancing the quality metric of protein microarray image *
 Wang et al. / J Zhejiang Univ SCI 2004 5(2):62-628 62 Journal of Zhejiang University SCIENCE ISSN 009-3095 http://www.zju.edu.cn/jzus E-mail: jzus@zju.edu.cn Enhancing the quality metric of protein microarray
Wang et al. / J Zhejiang Univ SCI 2004 5(2):62-628 62 Journal of Zhejiang University SCIENCE ISSN 009-3095 http://www.zju.edu.cn/jzus E-mail: jzus@zju.edu.cn Enhancing the quality metric of protein microarray
Acoustic resolution. photoacoustic Doppler velocimetry. in blood-mimicking fluids. Supplementary Information
 Acoustic resolution photoacoustic Doppler velocimetry in blood-mimicking fluids Joanna Brunker 1, *, Paul Beard 1 Supplementary Information 1 Department of Medical Physics and Biomedical Engineering, University
Acoustic resolution photoacoustic Doppler velocimetry in blood-mimicking fluids Joanna Brunker 1, *, Paul Beard 1 Supplementary Information 1 Department of Medical Physics and Biomedical Engineering, University
Computer Graphics Si Lu Fall /27/2016
 Computer Graphics Si Lu Fall 2017 09/27/2016 Announcement Class mailing list https://groups.google.com/d/forum/cs447-fall-2016 2 Demo Time The Making of Hallelujah with Lytro Immerge https://vimeo.com/213266879
Computer Graphics Si Lu Fall 2017 09/27/2016 Announcement Class mailing list https://groups.google.com/d/forum/cs447-fall-2016 2 Demo Time The Making of Hallelujah with Lytro Immerge https://vimeo.com/213266879
Supplemental Information
 Optically Activated Delayed Fluorescence Blake C. Fleischer, Jeffrey T. Petty, Jung-Cheng Hsiang, Robert M. Dickson, * School of Chemistry & Biochemistry and Petit Institute for Bioengineering and Bioscience,
Optically Activated Delayed Fluorescence Blake C. Fleischer, Jeffrey T. Petty, Jung-Cheng Hsiang, Robert M. Dickson, * School of Chemistry & Biochemistry and Petit Institute for Bioengineering and Bioscience,
Oscilloscope Measurements
 PC1143 Physics III Oscilloscope Measurements 1 Purpose Investigate the fundamental principles and practical operation of the oscilloscope using signals from a signal generator. Measure sine and other waveform
PC1143 Physics III Oscilloscope Measurements 1 Purpose Investigate the fundamental principles and practical operation of the oscilloscope using signals from a signal generator. Measure sine and other waveform
Confocal Microscopy. Kristin Jensen
 Confocal Microscopy Kristin Jensen 17.11.05 References Cell Biological Applications of Confocal Microscopy, Brian Matsumoto, chapter 1 Studying protein dynamics in living cells,, Jennifer Lippincott-Schwartz
Confocal Microscopy Kristin Jensen 17.11.05 References Cell Biological Applications of Confocal Microscopy, Brian Matsumoto, chapter 1 Studying protein dynamics in living cells,, Jennifer Lippincott-Schwartz
Page 21 GRAPHING OBJECTIVES:
 Page 21 GRAPHING OBJECTIVES: 1. To learn how to present data in graphical form manually (paper-and-pencil) and using computer software. 2. To learn how to interpret graphical data by, a. determining the
Page 21 GRAPHING OBJECTIVES: 1. To learn how to present data in graphical form manually (paper-and-pencil) and using computer software. 2. To learn how to interpret graphical data by, a. determining the
Ph 3455 The Photoelectric Effect
 Ph 3455 The Photoelectric Effect Required background reading Tipler, Llewellyn, section 3-3 Prelab Questions 1. In this experiment you will be using a mercury lamp as the source of photons. At the yellow
Ph 3455 The Photoelectric Effect Required background reading Tipler, Llewellyn, section 3-3 Prelab Questions 1. In this experiment you will be using a mercury lamp as the source of photons. At the yellow
Observing Microorganisms through a Microscope LIGHT MICROSCOPY: This type of microscope uses visible light to observe specimens. Compound Light Micros
 PHARMACEUTICAL MICROBIOLOGY JIGAR SHAH INSTITUTE OF PHARMACY NIRMA UNIVERSITY Observing Microorganisms through a Microscope LIGHT MICROSCOPY: This type of microscope uses visible light to observe specimens.
PHARMACEUTICAL MICROBIOLOGY JIGAR SHAH INSTITUTE OF PHARMACY NIRMA UNIVERSITY Observing Microorganisms through a Microscope LIGHT MICROSCOPY: This type of microscope uses visible light to observe specimens.
The History and Future of Measurement Technology in Sumitomo Electric
 ANALYSIS TECHNOLOGY The History and Future of Measurement Technology in Sumitomo Electric Noritsugu HAMADA This paper looks back on the history of the development of measurement technology that has contributed
ANALYSIS TECHNOLOGY The History and Future of Measurement Technology in Sumitomo Electric Noritsugu HAMADA This paper looks back on the history of the development of measurement technology that has contributed
DNA Size Selection Magnetic Beads
 DNA Size Selection Magnetic Beads Catalog #: 801-117 User Manual Last revised July 30 th, 2018 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
DNA Size Selection Magnetic Beads Catalog #: 801-117 User Manual Last revised July 30 th, 2018 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
An Evaluation of MTF Determination Methods for 35mm Film Scanners
 An Evaluation of Determination Methods for 35mm Film Scanners S. Triantaphillidou, R. E. Jacobson, R. Fagard-Jenkin Imaging Technology Research Group, University of Westminster Watford Road, Harrow, HA1
An Evaluation of Determination Methods for 35mm Film Scanners S. Triantaphillidou, R. E. Jacobson, R. Fagard-Jenkin Imaging Technology Research Group, University of Westminster Watford Road, Harrow, HA1
Akinori Mitani and Geoff Weiner BGGN 266 Spring 2013 Non-linear optics final report. Introduction and Background
 Akinori Mitani and Geoff Weiner BGGN 266 Spring 2013 Non-linear optics final report Introduction and Background Two-photon microscopy is a type of fluorescence microscopy using two-photon excitation. It
Akinori Mitani and Geoff Weiner BGGN 266 Spring 2013 Non-linear optics final report Introduction and Background Two-photon microscopy is a type of fluorescence microscopy using two-photon excitation. It
Sizing of nano structures below the diffraction limit using laser scanning microscopy
 Sizing of nano structures below the diffraction limit using laser scanning microscopy JAN BERGSTRAND Master s Thesis Supervisor: Stefan Wennmalm Examiner: Jerker Widengren trita? Abstract The resolution
Sizing of nano structures below the diffraction limit using laser scanning microscopy JAN BERGSTRAND Master s Thesis Supervisor: Stefan Wennmalm Examiner: Jerker Widengren trita? Abstract The resolution
Fig Color spectrum seen by passing white light through a prism.
 1. Explain about color fundamentals. Color of an object is determined by the nature of the light reflected from it. When a beam of sunlight passes through a glass prism, the emerging beam of light is not
1. Explain about color fundamentals. Color of an object is determined by the nature of the light reflected from it. When a beam of sunlight passes through a glass prism, the emerging beam of light is not
Shreyash Tandon M.S. III Year
 Shreyash Tandon M.S. III Year 20091015 Confocal microscopy is a powerful tool for generating high-resolution images and 3-D reconstructions of a specimen by using point illumination and a spatial pinhole
Shreyash Tandon M.S. III Year 20091015 Confocal microscopy is a powerful tool for generating high-resolution images and 3-D reconstructions of a specimen by using point illumination and a spatial pinhole
SAE AE-2 Lightning Committee White Paper
 SAE AE-2 Lightning Committee White Paper Recommended Camera Calibration and Image Evaluation Methods for Detection of Ignition Sources Rev. NEW January 2018 1 Table of Contents Executive Summary... 3 1.
SAE AE-2 Lightning Committee White Paper Recommended Camera Calibration and Image Evaluation Methods for Detection of Ignition Sources Rev. NEW January 2018 1 Table of Contents Executive Summary... 3 1.
Supplemental Reference Guide
 Supplemental Reference Guide QuantiGene ViewRNA mirna ISH Cell Assay P/N 19167 Rev.A 120623 For research use only. Not for use in diagnostic procedures. Trademarks Affymetrix and, and QuantiGene are trademarks
Supplemental Reference Guide QuantiGene ViewRNA mirna ISH Cell Assay P/N 19167 Rev.A 120623 For research use only. Not for use in diagnostic procedures. Trademarks Affymetrix and, and QuantiGene are trademarks
Fast, high-contrast imaging of animal development with scanned light sheet based structured-illumination microscopy
 nature methods Fast, high-contrast imaging of animal development with scanned light sheet based structured-illumination microscopy Philipp J Keller, Annette D Schmidt, Anthony Santella, Khaled Khairy,
nature methods Fast, high-contrast imaging of animal development with scanned light sheet based structured-illumination microscopy Philipp J Keller, Annette D Schmidt, Anthony Santella, Khaled Khairy,
Light, Color, Spectra 05/30/2006. Lecture 17 1
 What do we see? Light Our eyes can t t detect intrinsic light from objects (mostly infrared), unless they get red hot The light we see is from the sun or from artificial light When we see objects, we see
What do we see? Light Our eyes can t t detect intrinsic light from objects (mostly infrared), unless they get red hot The light we see is from the sun or from artificial light When we see objects, we see
Texture characterization in DIRSIG
 Rochester Institute of Technology RIT Scholar Works Theses Thesis/Dissertation Collections 2001 Texture characterization in DIRSIG Christy Burtner Follow this and additional works at: http://scholarworks.rit.edu/theses
Rochester Institute of Technology RIT Scholar Works Theses Thesis/Dissertation Collections 2001 Texture characterization in DIRSIG Christy Burtner Follow this and additional works at: http://scholarworks.rit.edu/theses
Standard Operating Procedure
 Standard Operating Procedure Nanosurf Atomic Force Microscopy Operation Facility NCCRD Nanotechnology Center for Collaborative Research and Development Department of Chemistry and Engineering Physics The
Standard Operating Procedure Nanosurf Atomic Force Microscopy Operation Facility NCCRD Nanotechnology Center for Collaborative Research and Development Department of Chemistry and Engineering Physics The
NAME SECTION PERFORMANCE TASK # 3. Part I. Qualitative Relationships
 NAME SECTION PARTNERS DATE PERFORMANCE TASK # 3 You must work in teams of three or four (ask instructor) and will turn in ONE report. Answer all questions. Write in complete sentences. You must hand this
NAME SECTION PARTNERS DATE PERFORMANCE TASK # 3 You must work in teams of three or four (ask instructor) and will turn in ONE report. Answer all questions. Write in complete sentences. You must hand this
RESISTANCE & OHM S LAW (PART I
 RESISTANCE & OHM S LAW (PART I and II) Objectives: To understand the relationship between potential and current in a resistor and to verify Ohm s Law. To understand the relationship between potential and
RESISTANCE & OHM S LAW (PART I and II) Objectives: To understand the relationship between potential and current in a resistor and to verify Ohm s Law. To understand the relationship between potential and
Aqualog. Water Quality Measurements Made Easy FLUORESCENCE
 Aqualog Water Quality Measurements Made Easy FLUORESCENCE Water quality measurements made easy The only simultaneous absorbance and fluorescence system for water quality analysis! The new Aqualog is the
Aqualog Water Quality Measurements Made Easy FLUORESCENCE Water quality measurements made easy The only simultaneous absorbance and fluorescence system for water quality analysis! The new Aqualog is the
Aqualog. Water Quality Measurements Made Easy PARTICLE CHARACTERIZATION ELEMENTAL ANALYSIS FLUORESCENCE
 Aqualog Water Quality Measurements Made Easy ELEMENTAL ANALYSIS FLUORESCENCE GRATINGS & OEM SPECTROMETERS OPTICAL COMPONENTS PARTICLE CHARACTERIZATION RAMAN SPECTROSCOPIC ELLIPSOMETRY SPR IMAGING Water
Aqualog Water Quality Measurements Made Easy ELEMENTAL ANALYSIS FLUORESCENCE GRATINGS & OEM SPECTROMETERS OPTICAL COMPONENTS PARTICLE CHARACTERIZATION RAMAN SPECTROSCOPIC ELLIPSOMETRY SPR IMAGING Water
Accuracy Estimation of Microwave Holography from Planar Near-Field Measurements
 Accuracy Estimation of Microwave Holography from Planar Near-Field Measurements Christopher A. Rose Microwave Instrumentation Technologies River Green Parkway, Suite Duluth, GA 9 Abstract Microwave holography
Accuracy Estimation of Microwave Holography from Planar Near-Field Measurements Christopher A. Rose Microwave Instrumentation Technologies River Green Parkway, Suite Duluth, GA 9 Abstract Microwave holography
Comparison of the Analysis Capabilities of Beckman Coulter MoFlo XDP and Becton Dickinson FACSAria I and II
 Comparison of the Analysis Capabilities of Beckman Coulter MoFlo XDP and Becton Dickinson FACSAria I and II Dr. Carley Ross, Angela Vandergaw, Katherine Carr, Karen Helm Flow Cytometry Business Center,
Comparison of the Analysis Capabilities of Beckman Coulter MoFlo XDP and Becton Dickinson FACSAria I and II Dr. Carley Ross, Angela Vandergaw, Katherine Carr, Karen Helm Flow Cytometry Business Center,
Cellular Bioengineering Boot Camp. Image Analysis
 Cellular Bioengineering Boot Camp Image Analysis Overview of the Lab Exercises Microscopy and Cellular Imaging The purpose of this laboratory exercise is to develop an understanding of the measurements
Cellular Bioengineering Boot Camp Image Analysis Overview of the Lab Exercises Microscopy and Cellular Imaging The purpose of this laboratory exercise is to develop an understanding of the measurements
Structural, optical, and electrical properties of phasecontrolled cesium lead iodide nanowires
 Electronic Supplementary Material Structural, optical, and electrical properties of phasecontrolled cesium lead iodide nanowires Minliang Lai 1, Qiao Kong 1, Connor G. Bischak 1, Yi Yu 1,2, Letian Dou
Electronic Supplementary Material Structural, optical, and electrical properties of phasecontrolled cesium lead iodide nanowires Minliang Lai 1, Qiao Kong 1, Connor G. Bischak 1, Yi Yu 1,2, Letian Dou
ImageJ. Once ImageJ is installed, open it up and open your scanned film file. 2. Under Image>Type click on 8-bit to convert the image to grayscale.
 ImageJ The homepage for ImageJ is here: http://rsb.info.nih.gov/ij/index.html wherein you can find links to the download, documentation, additional plugins and so on. Once ImageJ is installed, open it
ImageJ The homepage for ImageJ is here: http://rsb.info.nih.gov/ij/index.html wherein you can find links to the download, documentation, additional plugins and so on. Once ImageJ is installed, open it
Additional reagents and materials that are not supplied
 sparq PureMag Beads Cat. No. 95196-005 Size: 5 ml Store at 2 C to 8 C 95196-060 60 ml 95196-450 450 ml Description sparq PureMag Beads uses reversible nucleic acid-binding properties of magnetic beads
sparq PureMag Beads Cat. No. 95196-005 Size: 5 ml Store at 2 C to 8 C 95196-060 60 ml 95196-450 450 ml Description sparq PureMag Beads uses reversible nucleic acid-binding properties of magnetic beads
Camera Test Protocol. Introduction TABLE OF CONTENTS. Camera Test Protocol Technical Note Technical Note
 Technical Note CMOS, EMCCD AND CCD CAMERAS FOR LIFE SCIENCES Camera Test Protocol Introduction The detector is one of the most important components of any microscope system. Accurate detector readings
Technical Note CMOS, EMCCD AND CCD CAMERAS FOR LIFE SCIENCES Camera Test Protocol Introduction The detector is one of the most important components of any microscope system. Accurate detector readings
