SquasshAnalyst manual
|
|
|
- Imogen Fletcher
- 5 years ago
- Views:
Transcription
1 SquasshAnalyst manual Aurélien Rizk SquasshAnalyst graphical user interface is designed for intuitive use. For the shortest introduction to Squassh and SquasshAnalyst follow the installation instructions section 2 and the toy example analysis section 4.1. You will then be able to analyze your own data or further look into the provided datasets showcasing SquasshAnalyst features. The reference section lists and describes SquasshAnalyst functionalities. 1 Squassh and SquasshAnalyst software files Squassh and SquasshAnalyst are available as additional file with SquasshAnalyst paper and for download at: The.zip archive contains the following files : Lif_extractor.ijm is an ImageJ macro enabling extraction of.lif files form Leica s microscopes to a series of.tiff images. Squassh_.jar contains the Squassh plugin [1] for segmenting subcellular structures. SquasshAnalyst.zip contains the source code of SquasshAnalyst. 2 Installation and launch 2.1 Squassh Install Squassh in ImageJ by drag and dropping the Squassh_.jar file on ImageJ main window. For more information on Squassh and its use refer to [1]. ImageJ can be downloaded from 1
2 2.2 Lif extractor The Lif extractor is an ImageJ macro working in combination with Bio- Formats an ImageJ plugin allowing to open a large number of life sciences image file formats. Bio-Formats is required for the Lif extractor to function and can be obtained from bio-formats/5.1.1/. Use Plugins Macros Run in ImageJ menus and select the Lif_extractor.ijm file for a single use of the macro. Alternatively, for a permanent install of the Lif extractor macro, copy the content of the Lif_extractor.ijm file to the end of StartupMacros.txt contained in ImageJ s macros folder. This can be done by opening the Lif_extractor.ijm file in a text editor and copy and pasting its content. ImageJ s macro folder is located in ImageJ s installation folder, next to ImageJ s application binary. The macro is then available in Plugins Macros ImageJ menu. 2.3 SquasshAnalyst SquasshAnalyst is an R program running as an interactive web application using the web application framework for R Shiny [2]. It works with any modern browser and requires the free statistical software R to be installed on your system. Follow instructions at to install the latest R version. Shiny installation Before first launch of SquasshAnalyst it is necessary to install Shiny in R with the following R command: install.packages("shiny") SquasshAnalyst launch SquasshAnalyst is launched from R with: shiny::rungithub("squasshanalyst", "a-rizk") Note that this requires an internet connection. 2
3 Alternative launch method It is possible to launch SquasshAnalyst with no internet connection by decompressing SquasshAnalyst.zip, changing R working directory to the parent folder and running SquasshAnalyst with: shiny::runapp("squasshanalyst") 3 Example data We provide example datasets online on a cloud file sharing service. Toy example. This small toy example dataset allows to test the whole analysis workflow from the.lif extraction, Squassh segmentation to SquasshAnalyst analysis in less than fifteen minutes. The set provides two conditions with five 2D one channel fluorescence images in each condition. The images show COS cells 30 minutes after EGF stimulation and before EGF stimulation. The data is given as.lif files as would be obtained when exported from a Leica microscope. EGF RABx colocalization. This set investigates the colocalization of EGF with RAB5a and RAB11b in COS cells before and 30 minutes after EGF stimulations. Each condition contains five 2D images with three fluorescence channels. RABx localization. This set investigates the colocalization of four RAB GTPases with four subcellular markers in HEK cells. It contains 3D images with two fluorescence channels to examine the colocalization of RAB4a, RAB5a, RAB7a and RABB11b with subcellular markers for early endosomes (EEA1), late endosomes (LBPA), endoplasmic reticulum (PDI) and lysosomes (LBPA). The first channel is the subcellular marker while the second channel is the RAB. There are thus 4 4 conditions in this set. Each condition contains 20 to 25 images with a total of 330 images. Segmenting this whole set in Squassh takes approximately 24 hours on a quad core computer. We provide the raw data as well as already segmented Squassh output. colocalization of VEGF121a with RAB7a and RAB11b in live cell imaging. This example contains a single movie obtained after stimulation with VEGF121a. The three fluorescence channels correspond to RAB7a, RAB11b and VEGF121a. 3
4 The toy dataset (427KB) is available at: It contains raw data only. The three other datasets (637MB) are packed in a single.zip file. The file contains the segmentation results from Squassh for the three datasets. It is therefore possible to use them directly in SquasshAnalyst without segmenting the raw data in Squassh. It also contains the raw data of VEGF121a and EGF RABx datasets. In order to reduce file size, raw data (1.86Gb) for the RABx localization dataset is available in a separate file. Both are available at 4 Tutorial 4.1 Toy example Data preparation Download the toy example dataset at: Unzip the file. The obtained toy example folder contains two.lif files. Each.lif file contains the images corresponding to one experimental condition. Extract the.lif files using the.lif extractor macro. in ImageJ choose Plugins Macros Run and select the Lif_extractor.ijm file to launch the macro. Select the toy example folder. The Lif extractor will search recursively in the directory for.lif files and will create one new subdirectory for each.lif file containing the extracted.tiff images. Each.tiff image contains one image (with all stacks and channels if present). The toy example contains only one channel and one layer per image. Important note : when analyzing your own dataset the images should be organized in the same way as this toy example. They should either be extracted by the lif extractor with one.lif file per experimental condition or organized as a series of.tiff images with all.tiff images in a given folder corresponding to an experimental condition. Analyzing all experimental conditions in SquasshAnalyst is then done by selecting the parent folder in SquasshAnalyst data input tab. 4
5 Image analysis with Squassh Launch Squassh from ImageJ Plugins Squassh Click on Select File/Folder and select the toy example folder. In background subtraction options activate Remove background and set the rolling ball window size to 25. In the segmentation parameters panel set the regularization to and minimum object intensity for channel one to Check that intensity rescaling is active, subpixel segmentation inactive and intensity estimation is set to automatic. These settings can then be used as default settings for analyzing your own dataset. Adapt regularization and minimum intensity thresholds first to obtain a good segmentation. Refer to [1] for a description of further Squassh settings. Unselect all options in the cell masks and visualization options panels. Click Ok on the main Squassh window to launch the segmentation. Squassh will recursively analyze all the images in the folder. Data analysis with SquasshAnalyst Open the statistical analysis software R and launch SquasshAnalyst with: shiny::rungithub("squasshanalyst", "a-rizk") Under the data input tab select the toy example folder. Click on the Segmentation overview tab and select a condition for an overview of the segmentation made in Squassh. Select the Analysis tab and click on Select all under the experimental conditions selection field. Select Total object signal in the Subcellular object/image attribute field. This will display a comparison of the amount of internalized EGF before and after stimulation. 5
6 4.2 RABx subcellular localization Data selection From SquasshAnalyst Data input tab click Select Directory and select RABx colocalization directory. Rename channel one and two to marker and RABx. Colocalization of EEA1 with RABx Under Analysis tab in compare experimental conditions field select the four conditions containing EEA1. Select Colocalization (size) in Subcellular object / image attribute field. Select Percentage of channel.. Marker colocalizing with channel RABx to display the amount of EEA1 colocalizing with the four RABs. Sort the four RABs with sort conditions. RAB4a and RAB5a colocalize the most with EEA1 (early endosmes). EEA1 fluorescence channel is obtained with immunostaining. In the Subcellular objects and image filters panel increase the intensity threshold of the Marker channel to 0.3. This will remove low intensity objects of this channel from the analysis. Notice how it increases the colocalization scores of RAB4a and RAB5a with EEA1. This can be explained by the fact that in immunostaining low intensity objects can correspond to non specific staining. Under Most representative image select RAB5 EEA1 condition to obtain the most representative image of this condition. A preview of the image is displayed as well as the file name of the raw image. Statistical difference between conditions is given at the bottom of the page. 6
7 Colocalization of RAB5 with subcellular markers EEA1, LBPA, H4B4 and PDI Select all conditions containing RAB5a and repeat above given steps to obtain the colocalization of RAB5a with the four subcellular markers. 4.3 EGF RABx colocalization Data selection From SquasshAnalyst Data input tab click Select Directory and select EGF RABx colocalization directory. Rename channel one to three respectively to RAB5, RAB11 and EGF. Segmentation overview In segmentation overview tab select condition R5 R11 EGF T30 to overview segmentation of three channel images. Color legend of preview images is given on top : red for RAB5a, green for RAB11b and blue for EGF. Original image is on left side. Segmentation is on right side. EGF internalization before and after stimulation In Analysis tab select all conditions with Select all. Select Total object signal in image attribute selection and EGF in the channel selection field. By default box plots then display the total EGF signal present in both conditions. There is as expected no EGF present at time = 0 min. Colocalization of EGF with RAB5a and RAB11b Select only condition R5 R11 EGF T30 in condition selection field. Select Total object Volume Venn diagram in image attribute selection. The Venn diagram shows that in average over all images at time = 30 min objects with a total size of only 1.4 pixels are positive for RAB5a, RAB11b and EGF at the same time. EGF is colocalizing about twice as much with RAB5a than with RAB11b at this time point. 7
8 4.4 Live cell imaging Data selection From SquasshAnalyst Data input tab click Select Directory and select R7 R11 VEGF121a movie directory. Rename channel one to three respectively to RAB7, RAB11 and VEGF121a. Internalization of VEGF121a over time Under analysis tab select condition R7 R11 VEGF121a. Select Total object signal in image attribute selection. The curve displays the total amount of VEGF121a in segmented objects. This reflects VEGF121a internalization after stimulation. Data coming from movie analysis is often displaying high variability with time (from experimental noise or from cell stochasticity). It is possible to average over time with rolling mean. The slider controls the length of the sliding window used to average over time. Set it to 10 to smooth the curve. Colocalization of VEGF121a with RAB7 and RAB11 Select Colocalization (size) in image attribute selection and colocalization of VEGF121a with RAB7. Click Keep plot. Now select colocalization of VEGF121a with RAB7. The graph will display an overlay of two curves for the colocalization of VEGF121a with RAB7 and RAB11. This allows to monitor colocalization of the VEGF ligand with different RABs over time. 5 Reference 5.1 Data input Select a folder that contains Squassh analyzed fluorescence microscopy images. 8
9 Directory selection: When selecting a directory Squassh Analyst will recursively search for all.csv data files generated by Squassh in subdirectories and will display found conditions data. Each subdirectory should contain files corresponding to at most one conditon (contact sheet image files and.csv files). All conditions should contain the same type of data (2D/3D, one channel/two channels/three channels, fixed images/movie). When using Squassh Analyst on Windows computer the directory selection window opens in R. Movie analysis: For live cell imaging analysis data should be analyzed by Squassh as one.tiff file for all channels, slices and time points. Channel renaming: After folder selection and if Squassh data is found text fields for optional channel renaming appears. 5.2 Segmentation Overview Survey Squassh segmentation and exclude images from analysis Squassh parameters: After selecting a condition Squassh Analyst will display parameters used by Squassh for the segmentation. Images and segmentation preview: Squassh Analyst displays contact sheet previews of original microscopy images maximum intensity z-projections alongside Squassh segmentation z-projection. Cell mask if present is displayed on both images as white outline. Image features overview: Squassh Analyst displays next to contact sheets images a table containing mean number of objects in each channel and colocalization quantification (for two channel images). Exclude from further analysis bad images by ticking Ex- Image removal: clude image box. 5.3 Analysis Compute image features and compare across experimental conditions 9
10 Select wich image feature to com- Subcellular object/image attribute: pute and display. Mean object volume: computes the mean volume of subcellular objects. Mean is computed for each image over all its subcellular objects. For 2D images this computes the surface of subcellular objects. Mean object length: computes for each image the mean length of its subcellular objects. Mean object intensity: computes for each image the mean intensity of its subcellular objects. Intensities are normalized in each image between 0 and 1. Colocalization (number): percentage of objects in one channel that overlap with objects from the other channel. An object is considered overlapping with the other channel if at least 50% of its volume is overlapping with objects form the other channel. More precisely the colocalization of channel one with channel two is defined as: C number (O1 O2+ /O1) = {o O1 : O o > 0.5} O1 where O1 is the set of all objects detected in channel 1 and O2 the set of objects detected in channel 2. denotes the total number of objects in the given set. O o is the fraction of object o that overlaps with any other object from the other channel. The notation O1 O2+ intuitively means objects in channel one that are positive for objects in channel two. Colocalization (size): percentage of the total size of objects in one channel overlapping with objects from the other channel. The size-based colocalization coefficient of channel one with channel two is defined as: C size (O1 O2+ /O1) = o O1 S oo o o O1 S o 10
11 where S o is the size of object o given by the number of pixels belonging to it. Hence, o O1 S oo o is the total size of the colocalizing regions and o O1 S o is the total number of pixels covered by any object in that channel. Colocalization (signal): percentage of the total signal (=volume*intensity) of objects in one channel overlapping with objects from the other channel The intensity-based colocalization coefficients of channel one with channel two is: C signal (O1 O2+ /O1) = o O1 I os o O o o O1 I os o where I o is the estimated intensity of object o. Hence, o O1 I os o O o is the total signal in the colocalizing region in channel 1, while o O1 I os o is the total signal present in channel 1 altogether. Pearson correlation: pearson correlation coefficient between two channels Pearson correlation inside cell mask: computed inside cell masks only. pearson correlation coefficient Total object volume Venn diagram: displays a Venn diagramm representing the overlap between the objects volumes in the different channels. This attribute can only be displayed for one experimental condition at a time. The first condition selected in Compare experimental conditionss is used. Total object signal: total signal (=volume*intensity) of objects in an image. This quantitiy is extensive with the cell size : the larger a cell is, the more subcellular objects it can have. Total object signal / Cell Size: same as Total object signal but normalized with the cell size. Cell size is computed from the cell mask. Total object volume: total volume of objects in an image. Total object volume / Cell Size: same as total object volume but normalized with cell size. 11
12 Object number: total number of objects in an image. Object number / Cell Size: same as Object number but normalized with cell size. Channel selection: Select the channel in which the image attribute is computed. For three channel images, when a Pearson correlation attribute is selected the field turns into a two channel selection form. For a colocalization attribute the selection is ordered to be able to select either channnel 1 colocalizing with channel 2 or channel 2 colocalizing with channel 1. In the case of three channel images and colocalization attribute it is possible to compute the colocalization of one channel with two others by selecting one channel in the first field and two in the second field. Condition selection: Experimental conditions on which the image attribute will be computed. Each condition will be displayed in the chart as a separate bar or boxplot. The condition selection field is non reactive, use Apply conditions changes for modifications to take effect. Use Select all or Deselect all as shortcuts to select/ deselect all conditions. Excluded images: Images excluded in the segmentation overview tab are given here as a reminder. Chart type: Plot image attributes with Box plot, Strip chart, or Bar chart. Bar charts are displayed with mean values and standard error of the mean. Conditions can be sorted by increasing mean value by ticking Sort conditions. Movie analysis: In the case of movie analysis there is no chart type option. The plot displays the evolution over time of the selected image attribute. Figure export: Use Export pdf button to export current chart in.pdf format. For png format export, right click the chart and save as image. Export the data corresponding to the currently displayed chart in a spreadsheet file with Export.csv. 12
13 Filters: Remove subcellular objects from the analysis by setting thresholds on their minimum volume, maximum volume or minimum intensities. Remove images from the analysis by setting a threshold on the minimum number of subcellular objects an image has to contain. Most representative image: SuqasshAnalyst displays the image whose attribute value is closest to the median value of all images in the chosen condition. This thus provides the most representative image of the group for the calclated attribute. The file name of the image is given on top of its contact sheet preview. Statistical analysis: When at least two experimental conditions are selected statistical significance tests results are performed in this section. Two statistical analysis are possible, parametric or non parametric. The first one is one way ANOVA followed by Tukey Honest Significance difference test for multiple comparisons. The second one is the Kruskal-Wallis test followed by Dunn s test adjusted with the Bonferonni correction for multiple comparisons. ANOVA and Tukey are parametric, that is they require that samples come from Gaussian distributions. The non parametric tests Kruskal-Wallis and Dunn do not require this assumption but only take into account the rankings of the values and have thus less statistical power. Star ratings are given next to p values with **** for p value < , *** for < p value < 0.001, ** for < p value < 0.01,* for 0.01 < p value < 0.05 and ns for p value > References [1] Rizk et al. Segmentation and quantification of subcellular structures in fluorescence microscopy images using Squassh. Nat Protoc, 9 (3), [2] Winston Chang, Joe Cheng, JJ Allaire, Yihui Xie and Jonathan McPherson (2015). shiny: Web Application Framework for R. R package version
Dr. Bob on Colocalization or MSL Experiments In Learning Colocalization Using Image J
 Dr. Bob on Colocalization or MSL Experiments In Learning Colocalization Using Image J Confocal microscopy is used to test whether two fluorescently labeled molecules are associated with one another. If
Dr. Bob on Colocalization or MSL Experiments In Learning Colocalization Using Image J Confocal microscopy is used to test whether two fluorescently labeled molecules are associated with one another. If
Manual. Cell Border Tracker. Jochen Seebach Institut für Anatomie und Vaskuläre Biologie, WWU Münster
 Manual Cell Border Tracker Jochen Seebach Institut für Anatomie und Vaskuläre Biologie, WWU Münster 1 Cell Border Tracker 1. System Requirements The software requires Windows XP operating system or higher
Manual Cell Border Tracker Jochen Seebach Institut für Anatomie und Vaskuläre Biologie, WWU Münster 1 Cell Border Tracker 1. System Requirements The software requires Windows XP operating system or higher
inform ADVANCED IMAGE ANALYSIS SOFTWARE inform User Manual
 inform ADVANCED IMAGE ANALYSIS SOFTWARE inform User Manual Notice The information in this document is subject to change without notice and should not be construed as a commitment by PerkinElmer, Inc. PerkinElmer
inform ADVANCED IMAGE ANALYSIS SOFTWARE inform User Manual Notice The information in this document is subject to change without notice and should not be construed as a commitment by PerkinElmer, Inc. PerkinElmer
IncuCyte ZOOM Fluorescent Processing Overview
 IncuCyte ZOOM Fluorescent Processing Overview The IncuCyte ZOOM offers users the ability to acquire HD phase as well as dual wavelength fluorescent images of living cells producing multiplexed data that
IncuCyte ZOOM Fluorescent Processing Overview The IncuCyte ZOOM offers users the ability to acquire HD phase as well as dual wavelength fluorescent images of living cells producing multiplexed data that
User Manual for HoloStudio M4 2.5 with HoloMonitor M4. Phase Holographic Imaging
 User Manual for HoloStudio M4 2.5 with HoloMonitor M4 Phase Holographic Imaging 1 2 HoloStudio M4 2.5 Software instruction manual 2013 Phase Holographic Imaging AB 3 Contact us: Phase Holographic Imaging
User Manual for HoloStudio M4 2.5 with HoloMonitor M4 Phase Holographic Imaging 1 2 HoloStudio M4 2.5 Software instruction manual 2013 Phase Holographic Imaging AB 3 Contact us: Phase Holographic Imaging
MRI Grid. The MRI Grid is a tool in MRI Cell Image Analyzer, that can be used to associate measurements with labeled positions on a board.
 Abstract The is a tool in MRI Cell Image Analyzer, that can be used to associate measurements with labeled positions on a board. Illustration 2: A grid on a binary image. Illustration 1: The interface
Abstract The is a tool in MRI Cell Image Analyzer, that can be used to associate measurements with labeled positions on a board. Illustration 2: A grid on a binary image. Illustration 1: The interface
Protocols. Graphical programming for Icy. a.k.a. programming, for the rest of us
 Protocols Graphical programming for Icy a.k.a. programming, for the rest of us Foreword: Reproducible Research Quote: "Results aren't much if they can t be reproduced!" (your boss, your reviewers, your
Protocols Graphical programming for Icy a.k.a. programming, for the rest of us Foreword: Reproducible Research Quote: "Results aren't much if they can t be reproduced!" (your boss, your reviewers, your
Introduction to Image Analysis with
 Introduction to Image Analysis with PLEASE ENSURE FIJI IS INSTALLED CORRECTLY! WHAT DO WE HOPE TO ACHIEVE? Specifically, the workshop will cover the following topics: 1. Opening images with Bioformats
Introduction to Image Analysis with PLEASE ENSURE FIJI IS INSTALLED CORRECTLY! WHAT DO WE HOPE TO ACHIEVE? Specifically, the workshop will cover the following topics: 1. Opening images with Bioformats
Image Analysis for Fluorescence
 Image Analysis for Fluorescence Terminology Table Image Analysis Macro Colocalization Intensity Dye AFI The extraction of meaningful information from digital images by means of digital image processing
Image Analysis for Fluorescence Terminology Table Image Analysis Macro Colocalization Intensity Dye AFI The extraction of meaningful information from digital images by means of digital image processing
Ca ++ Imaging Analysis in MATLAB
 Students names: Ca ++ Imaging Analysis in MATLAB Universität Basel, Mrsic-Flogel Lab Introduction In this practical you will analyse -photon imaging data using MATLAB. This is the same approach neuroscientists
Students names: Ca ++ Imaging Analysis in MATLAB Universität Basel, Mrsic-Flogel Lab Introduction In this practical you will analyse -photon imaging data using MATLAB. This is the same approach neuroscientists
Excel Tool: Plots of Data Sets
 Excel Tool: Plots of Data Sets Excel makes it very easy for the scientist to visualize a data set. In this assignment, we learn how to produce various plots of data sets. Open a new Excel workbook, and
Excel Tool: Plots of Data Sets Excel makes it very easy for the scientist to visualize a data set. In this assignment, we learn how to produce various plots of data sets. Open a new Excel workbook, and
LSM 800 Confocal Microscope Standard Operation Protocol
 LSM 800 Confocal Microscope Standard Operation Protocol Turning on the system 1. Switch on the Main switch (labeled 1 and 2 ) mounted on the wall. 2. Turn the Laser Key (labeled 3 ) 90 clockwise for power
LSM 800 Confocal Microscope Standard Operation Protocol Turning on the system 1. Switch on the Main switch (labeled 1 and 2 ) mounted on the wall. 2. Turn the Laser Key (labeled 3 ) 90 clockwise for power
Excel Lab 2: Plots of Data Sets
 Excel Lab 2: Plots of Data Sets Excel makes it very easy for the scientist to visualize a data set. In this assignment, we learn how to produce various plots of data sets. Open a new Excel workbook, and
Excel Lab 2: Plots of Data Sets Excel makes it very easy for the scientist to visualize a data set. In this assignment, we learn how to produce various plots of data sets. Open a new Excel workbook, and
IMAGE PROCESSING PRACTICALS
 EPFL PTBIOP IMAGE PROCESSING PRACTICALS 14.03.2011-16.03.2011 ACKNOWLEDGEMENTS This presentation and the exercises are based on the script CMCI Image processing & Analysis Course Series I which was kindly
EPFL PTBIOP IMAGE PROCESSING PRACTICALS 14.03.2011-16.03.2011 ACKNOWLEDGEMENTS This presentation and the exercises are based on the script CMCI Image processing & Analysis Course Series I which was kindly
BacklightFly Manual.
 BacklightFly Manual http://www.febees.com/ Contents Start... 3 Installation... 3 Registration... 7 BacklightFly 1-2-3... 9 Overview... 10 Layers... 14 Layer Container... 14 Layer... 16 Density and Design
BacklightFly Manual http://www.febees.com/ Contents Start... 3 Installation... 3 Registration... 7 BacklightFly 1-2-3... 9 Overview... 10 Layers... 14 Layer Container... 14 Layer... 16 Density and Design
Introduction to BioImage Analysis
 Introduction to BioImage Analysis Qi Gao CellNetworks Math-Clinic core facility 22-23.02.2018 MATH- CLINIC Math-Clinic core facility Data analysis services on bioimage analysis & bioinformatics: 1-to-1
Introduction to BioImage Analysis Qi Gao CellNetworks Math-Clinic core facility 22-23.02.2018 MATH- CLINIC Math-Clinic core facility Data analysis services on bioimage analysis & bioinformatics: 1-to-1
Recitation 2 Introduction to Photoshop
 Recitation 2 Introduction to Photoshop What is Adobe Photoshop? Adobe Photoshop is a tool for creating digital graphics either by starting with a scanned photograph or artwork or by creating the graphics
Recitation 2 Introduction to Photoshop What is Adobe Photoshop? Adobe Photoshop is a tool for creating digital graphics either by starting with a scanned photograph or artwork or by creating the graphics
ImageJ: Introduction to Image Analysis 3 May 2012 Jacqui Ross
 Biomedical Imaging Research Unit School of Medical Sciences Faculty of Medical and Health Sciences The University of Auckland Private Bag 92019 Auckland 1142, NZ Ph: 373 7599 ext. 87438 http://www.fmhs.auckland.ac.nz/sms/biru/.
Biomedical Imaging Research Unit School of Medical Sciences Faculty of Medical and Health Sciences The University of Auckland Private Bag 92019 Auckland 1142, NZ Ph: 373 7599 ext. 87438 http://www.fmhs.auckland.ac.nz/sms/biru/.
Batch Counting of Foci
 Batch Counting of Foci Getting results from Z stacks of images. 1. First it is necessary to determine suitable CHARM parameters to be used for batch counting. First drag a stack of images taken with the
Batch Counting of Foci Getting results from Z stacks of images. 1. First it is necessary to determine suitable CHARM parameters to be used for batch counting. First drag a stack of images taken with the
Locating Molecules Using GSD Technology Project Folders: Organization of Experiment Files...1
 .....................................1 1 Project Folders: Organization of Experiment Files.................................1 2 Steps........................................................................2
.....................................1 1 Project Folders: Organization of Experiment Files.................................1 2 Steps........................................................................2
Volocity Tutorial Creating a Measurement Protocol in Volocity Software
 TUTORIAL NOTE Cellular Imaging and Analysis Volocity Tutorial Creating a Measurement Protocol in Volocity Software This tutorial will illustrate the process of creating a Measurement Protocol in Volocity
TUTORIAL NOTE Cellular Imaging and Analysis Volocity Tutorial Creating a Measurement Protocol in Volocity Software This tutorial will illustrate the process of creating a Measurement Protocol in Volocity
Introduction to BioImage Analysis using Fiji
 Introduction to BioImage Analysis using Fiji CellNetworks Math-Clinic core facility Qi Gao Carlo A. Beretta 12.05.2017 Math-Clinic core facility Data analysis services on bioinformatics & bioimage analysis:
Introduction to BioImage Analysis using Fiji CellNetworks Math-Clinic core facility Qi Gao Carlo A. Beretta 12.05.2017 Math-Clinic core facility Data analysis services on bioinformatics & bioimage analysis:
Importing and processing gel images
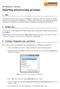 BioNumerics Tutorial: Importing and processing gel images 1 Aim Comprehensive tools for the processing of electrophoresis fingerprints, both from slab gels and capillary sequencers are incorporated into
BioNumerics Tutorial: Importing and processing gel images 1 Aim Comprehensive tools for the processing of electrophoresis fingerprints, both from slab gels and capillary sequencers are incorporated into
Version 2 Image Clarification Tool for Avid Editing Systems. Part of the dtective suite of forensic video analysis tools from Ocean Systems
 By Version 2 Image Clarification Tool for Avid Editing Systems Part of the dtective suite of forensic video analysis tools from Ocean Systems User Guide www.oceansystems.com www.dtectivesystem.com Page
By Version 2 Image Clarification Tool for Avid Editing Systems Part of the dtective suite of forensic video analysis tools from Ocean Systems User Guide www.oceansystems.com www.dtectivesystem.com Page
Introduction to Autodesk Inventor for F1 in Schools (Australian Version)
 Introduction to Autodesk Inventor for F1 in Schools (Australian Version) F1 in Schools race car In this course you will be introduced to Autodesk Inventor, which is the centerpiece of Autodesk s Digital
Introduction to Autodesk Inventor for F1 in Schools (Australian Version) F1 in Schools race car In this course you will be introduced to Autodesk Inventor, which is the centerpiece of Autodesk s Digital
Downloading Imagery & LIDAR
 Downloading Imagery & LIDAR 333 Earth Explorer The USGS is a great source for downloading many different GIS data products for the entire US and Canada and much of the world. Below are instructions for
Downloading Imagery & LIDAR 333 Earth Explorer The USGS is a great source for downloading many different GIS data products for the entire US and Canada and much of the world. Below are instructions for
EPFL BIOP Image Processing Practicals R. Guiet, O. Burri
 EPFL BIOP Image Processing Practicals 23-25.03.2015 R. Guiet, O. Burri Overview DAY 1 Intensity/Histogram Look up table (LUT) Contrast Image Depth RGB images Image Math File Formats Resizing Images Regions
EPFL BIOP Image Processing Practicals 23-25.03.2015 R. Guiet, O. Burri Overview DAY 1 Intensity/Histogram Look up table (LUT) Contrast Image Depth RGB images Image Math File Formats Resizing Images Regions
Chapter 6: TVA MR and Cardiac Function
 Chapter 6 Cardiac MR Introduction Chapter 6: TVA MR and Cardiac Function The Time-Volume Analysis (TVA) optional module calculates time-dependent behavior of volumes in multi-phase studies from MR. An
Chapter 6 Cardiac MR Introduction Chapter 6: TVA MR and Cardiac Function The Time-Volume Analysis (TVA) optional module calculates time-dependent behavior of volumes in multi-phase studies from MR. An
Sheet Metal Punch ifeatures
 Lesson 5 Sheet Metal Punch ifeatures Overview This lesson describes punch ifeatures and their use in sheet metal parts. You use punch ifeatures to simplify the creation of common and specialty cut and
Lesson 5 Sheet Metal Punch ifeatures Overview This lesson describes punch ifeatures and their use in sheet metal parts. You use punch ifeatures to simplify the creation of common and specialty cut and
Definiens. Tissue Studio 4.2. Tutorial 1: Composer and Nuclear Markers
 Definiens Tissue Studio 4.2 Tutorial 1: Composer and Nuclear Markers Tutorial 1: Composer and Nuclear Markers Imprint and Version Copyright 2015 Definiens AG. All rights reserved. This document may be
Definiens Tissue Studio 4.2 Tutorial 1: Composer and Nuclear Markers Tutorial 1: Composer and Nuclear Markers Imprint and Version Copyright 2015 Definiens AG. All rights reserved. This document may be
Point Spread Function Estimation Tool, Alpha Version. A Plugin for ImageJ
 Tutorial Point Spread Function Estimation Tool, Alpha Version A Plugin for ImageJ Benedikt Baumgartner Jo Helmuth jo.helmuth@inf.ethz.ch MOSAIC Lab, ETH Zurich www.mosaic.ethz.ch This tutorial explains
Tutorial Point Spread Function Estimation Tool, Alpha Version A Plugin for ImageJ Benedikt Baumgartner Jo Helmuth jo.helmuth@inf.ethz.ch MOSAIC Lab, ETH Zurich www.mosaic.ethz.ch This tutorial explains
Quick Guide for Zeiss 710 Laser Scanning Confocal MGH Cancer Center
 Quick Guide for Zeiss 710 Laser Scanning Confocal MGH Cancer Center For any questions or concerns, please contact: Linda Nieman lnieman@mgh.harvard.edu Office: (617) 643-9684 Cell: (512) 565-8076 Chenyue
Quick Guide for Zeiss 710 Laser Scanning Confocal MGH Cancer Center For any questions or concerns, please contact: Linda Nieman lnieman@mgh.harvard.edu Office: (617) 643-9684 Cell: (512) 565-8076 Chenyue
ADMS 5 MapInfo Link. User Guide CERC
 ADMS 5 MapInfo Link User Guide CERC ADMS 5 MapInfo Link User Guide November 2012 Cambridge Environmental Research Consultants Ltd 3 King s Parade Cambridge CB2 1SJ Telephone: +44 (0)1223 357773 Fax: +44
ADMS 5 MapInfo Link User Guide CERC ADMS 5 MapInfo Link User Guide November 2012 Cambridge Environmental Research Consultants Ltd 3 King s Parade Cambridge CB2 1SJ Telephone: +44 (0)1223 357773 Fax: +44
Instruction Manual. Mark Deimund, Zuyi (Jacky) Huang, Juergen Hahn
 Instruction Manual Mark Deimund, Zuyi (Jacky) Huang, Juergen Hahn This manual is for the program that implements the image analysis method presented in our paper: Z. Huang, F. Senocak, A. Jayaraman, and
Instruction Manual Mark Deimund, Zuyi (Jacky) Huang, Juergen Hahn This manual is for the program that implements the image analysis method presented in our paper: Z. Huang, F. Senocak, A. Jayaraman, and
AstroImageJ User Guide
 AstroImageJ User Guide Introduction AstroImageJ (AIJ) is simply ImageJ (IJ) with some customizations to the base code and a packaged set of astronomy specific plugins. The plugins are based on the Astronomy
AstroImageJ User Guide Introduction AstroImageJ (AIJ) is simply ImageJ (IJ) with some customizations to the base code and a packaged set of astronomy specific plugins. The plugins are based on the Astronomy
Microsoft Excel. Creating a Pie Chart on a Picture. 1. In order to create a pie chart on a picture, you need to first find
 Microsoft Excel Creating a Pie Chart on a Picture Name Date 1. In order to create a pie chart on a picture, you need to first find the picture you want to use. Click on the Internet Explorer icon. 2. When
Microsoft Excel Creating a Pie Chart on a Picture Name Date 1. In order to create a pie chart on a picture, you need to first find the picture you want to use. Click on the Internet Explorer icon. 2. When
Quick Guide. NucleoCounter NC-3000
 Quick Guide NucleoCounter NC-3000 Table of contents Setting up the FlexiCyte Protocol 2 Editing Image Capture and Analysis Parameters 3 Optimizing Exposure Time 4 Compensation for Spectral Overlap 6 Creating
Quick Guide NucleoCounter NC-3000 Table of contents Setting up the FlexiCyte Protocol 2 Editing Image Capture and Analysis Parameters 3 Optimizing Exposure Time 4 Compensation for Spectral Overlap 6 Creating
Using Adobe Photoshop
 Using Adobe Photoshop 6 One of the most useful features of applications like Photoshop is the ability to work with layers. allow you to have several pieces of images in the same file, which can be arranged
Using Adobe Photoshop 6 One of the most useful features of applications like Photoshop is the ability to work with layers. allow you to have several pieces of images in the same file, which can be arranged
Leica SPEII confocal microscope. Short Manual
 Leica SPEII confocal microscope Short Manual Switching ON sequence: 1. Turn on the Workstation under the bench (top, far right). 2. Turn on the Supply Unit - Laser box (big green switch first and then
Leica SPEII confocal microscope Short Manual Switching ON sequence: 1. Turn on the Workstation under the bench (top, far right). 2. Turn on the Supply Unit - Laser box (big green switch first and then
Assignment 5 due Monday, May 7
 due Monday, May 7 Simulations and the Law of Large Numbers Overview In both parts of the assignment, you will be calculating a theoretical probability for a certain procedure. In other words, this uses
due Monday, May 7 Simulations and the Law of Large Numbers Overview In both parts of the assignment, you will be calculating a theoretical probability for a certain procedure. In other words, this uses
PixInsight Workflow. Revision 1.2 March 2017
 Revision 1.2 March 2017 Contents 1... 1 1.1 Calibration Workflow... 2 1.2 Create Master Calibration Frames... 3 1.2.1 Create Master Dark & Bias... 3 1.2.2 Create Master Flat... 5 1.3 Calibration... 8
Revision 1.2 March 2017 Contents 1... 1 1.1 Calibration Workflow... 2 1.2 Create Master Calibration Frames... 3 1.2.1 Create Master Dark & Bias... 3 1.2.2 Create Master Flat... 5 1.3 Calibration... 8
ARC HYDRO GROUNDWATER TUTORIALS
 ARC HYDRO GROUNDWATER TUTORIALS Subsurface Analyst Creating ArcMap cross sections from existing cross section images Arc Hydro Groundwater (AHGW) is a geodatabase design for representing groundwater datasets
ARC HYDRO GROUNDWATER TUTORIALS Subsurface Analyst Creating ArcMap cross sections from existing cross section images Arc Hydro Groundwater (AHGW) is a geodatabase design for representing groundwater datasets
Stitching distortion-free mosaic images for QWA using PTGui. Georg von Arx
 Stitching distortion-free mosaic images for QWA using PTGui Georg von Arx Index A. Introduction and overview... 2 B. Taking microscopic images... 2 C. Installing PTGui... 3 D. Initial Setup... 3 E. Preparing
Stitching distortion-free mosaic images for QWA using PTGui Georg von Arx Index A. Introduction and overview... 2 B. Taking microscopic images... 2 C. Installing PTGui... 3 D. Initial Setup... 3 E. Preparing
CONCEPTS EXPLAINED CONCEPTS (IN ORDER)
 CONCEPTS EXPLAINED This reference is a companion to the Tutorials for the purpose of providing deeper explanations of concepts related to game designing and building. This reference will be updated with
CONCEPTS EXPLAINED This reference is a companion to the Tutorials for the purpose of providing deeper explanations of concepts related to game designing and building. This reference will be updated with
Automatic data analysis
 NOVA technical note #1 1 Automatic data analysis Case study: automatic IV curve and power curve from fuel cell measurements Fuel cell characterization is usually performed by measuring the IV and power
NOVA technical note #1 1 Automatic data analysis Case study: automatic IV curve and power curve from fuel cell measurements Fuel cell characterization is usually performed by measuring the IV and power
Introduction to ImageJ 8 Sept 2009
 Biomedical Imaging Research Unit School of Medical Sciences Faculty of Medical and Health Sciences The University of Auckland Private Bag 92019 Auckland, NZ Ph: 373 7599 ext. 87438 http://www.auckland.ac.nz/biru/
Biomedical Imaging Research Unit School of Medical Sciences Faculty of Medical and Health Sciences The University of Auckland Private Bag 92019 Auckland, NZ Ph: 373 7599 ext. 87438 http://www.auckland.ac.nz/biru/
AxioVision User's Guide. Release 4.1
 AxioVision User's Guide Release 4.1 Number of this manual: B 48-0038 e 10.2003 Date of issue: 10.2003 Carl Zeiss Vision draws the User's attention to the fact that the information and references contained
AxioVision User's Guide Release 4.1 Number of this manual: B 48-0038 e 10.2003 Date of issue: 10.2003 Carl Zeiss Vision draws the User's attention to the fact that the information and references contained
Digital Imaging - Photoshop
 Digital Imaging - Photoshop A digital image is a computer representation of a photograph. It is composed of a grid of tiny squares called pixels (picture elements). Each pixel has a position on the grid
Digital Imaging - Photoshop A digital image is a computer representation of a photograph. It is composed of a grid of tiny squares called pixels (picture elements). Each pixel has a position on the grid
Managing Your Workflow Using Coloured Filters with Snapper.Photo s PhotoManager Welcome to the World of S napper.photo
 Managing Your Workflow Using Coloured Filters with Snapper.Photo s PhotoManager Welcome to the World of S napper.photo Get there with a click Click on an Index Line to go directly there Click on the home
Managing Your Workflow Using Coloured Filters with Snapper.Photo s PhotoManager Welcome to the World of S napper.photo Get there with a click Click on an Index Line to go directly there Click on the home
CloneSelect Imager OPERATOR MANUAL SOFTWARE RELEASE
 CloneSelect Imager SOFTWARE RELEASE 1.3.73.1073 Control #: 05MAN1070.A5 Effective Date: xx-xx-xx ECO #: 3092 Contents Introduction... 5 CloneSelect Imager... 5 System Features... 5 Before Starting... 5
CloneSelect Imager SOFTWARE RELEASE 1.3.73.1073 Control #: 05MAN1070.A5 Effective Date: xx-xx-xx ECO #: 3092 Contents Introduction... 5 CloneSelect Imager... 5 System Features... 5 Before Starting... 5
Spreadsheets 3: Charts and Graphs
 Spreadsheets 3: Charts and Graphs Name: Main: When you have finished this handout, you should have the following skills: Setting up data correctly Labeling axes, legend, scale, title Editing symbols, colors,
Spreadsheets 3: Charts and Graphs Name: Main: When you have finished this handout, you should have the following skills: Setting up data correctly Labeling axes, legend, scale, title Editing symbols, colors,
RETRO 3D MOVIE EFFECT
 RETRO 3D MOVIE EFFECT Long before Avatar transported us to the breathtakingly beautiful world of Pandora with its state of the art 3D technology, movie audiences in the 1950 s were wearing cheap cardboard
RETRO 3D MOVIE EFFECT Long before Avatar transported us to the breathtakingly beautiful world of Pandora with its state of the art 3D technology, movie audiences in the 1950 s were wearing cheap cardboard
SKF TKTI. Thermal Camera Software. Instructions for use
 SKF TKTI Thermal Camera Software Instructions for use Table of contents 1. Introduction...4 1.1 Installing and starting the Software... 5 2. Usage Notes...6 3. Image Properties...7 3.1 Loading images
SKF TKTI Thermal Camera Software Instructions for use Table of contents 1. Introduction...4 1.1 Installing and starting the Software... 5 2. Usage Notes...6 3. Image Properties...7 3.1 Loading images
1 ImageBrowser Software User Guide 5.1
 1 ImageBrowser Software User Guide 5.1 Table of Contents (1/2) Chapter 1 What is ImageBrowser? Chapter 2 What Can ImageBrowser Do?... 5 Guide to the ImageBrowser Windows... 6 Downloading and Printing Images
1 ImageBrowser Software User Guide 5.1 Table of Contents (1/2) Chapter 1 What is ImageBrowser? Chapter 2 What Can ImageBrowser Do?... 5 Guide to the ImageBrowser Windows... 6 Downloading and Printing Images
Geomatica OrthoEngine v10.2 Tutorial DEM Extraction of WorldView-1 Data
 Geomatica OrthoEngine v10.2 Tutorial DEM Extraction of WorldView-1 Data WorldView 1, launched on September 18, 2007, offers a panchromatic imagery at a very high resolution of 50 cm at nadir. The key benefits
Geomatica OrthoEngine v10.2 Tutorial DEM Extraction of WorldView-1 Data WorldView 1, launched on September 18, 2007, offers a panchromatic imagery at a very high resolution of 50 cm at nadir. The key benefits
Photoshop CS2. Step by Step Instructions Using Layers. Adobe. About Layers:
 About Layers: Layers allow you to work on one element of an image without disturbing the others. Think of layers as sheets of acetate stacked one on top of the other. You can see through transparent areas
About Layers: Layers allow you to work on one element of an image without disturbing the others. Think of layers as sheets of acetate stacked one on top of the other. You can see through transparent areas
Printer Driver. This guide describes how to set up the Printer Driver for Windows 7.
 4-187-187-12 (1) Printer Driver Setup Guide This guide describes how to set up the Printer Driver for Windows 7. Before Using this Software Before using the printer driver, be sure to read the Readme file.
4-187-187-12 (1) Printer Driver Setup Guide This guide describes how to set up the Printer Driver for Windows 7. Before Using this Software Before using the printer driver, be sure to read the Readme file.
Excel Manual X Axis Label Below Chart 2010 >>>CLICK HERE<<<
 Excel Manual X Axis Label Below Chart 2010 When the X-axis is crowded with labels one way to solve the problem is to split the labels for to use two rows of labels enter the two rows of X-axis labels as
Excel Manual X Axis Label Below Chart 2010 When the X-axis is crowded with labels one way to solve the problem is to split the labels for to use two rows of labels enter the two rows of X-axis labels as
Chanalyzer by MetaGeek USER GUIDE page 1
 Chanalyzer 5 Chanalyzer by MetaGeek USER GUIDE page 1 Chanalyzer 5 spectrum analysis software Table of Contents Introduction What is Wi-Spy? What is Chanalyzer? Installation Choose a Wireless Network Interface
Chanalyzer 5 Chanalyzer by MetaGeek USER GUIDE page 1 Chanalyzer 5 spectrum analysis software Table of Contents Introduction What is Wi-Spy? What is Chanalyzer? Installation Choose a Wireless Network Interface
Adobe Studio on Adobe Photoshop CS2 Enhance scientific and medical images. 2 Hide the original layer.
 1 Adobe Studio on Adobe Photoshop CS2 Light, shadow and detail interact in wild and mysterious ways in microscopic photography, posing special challenges for the researcher and educator. With Adobe Photoshop
1 Adobe Studio on Adobe Photoshop CS2 Light, shadow and detail interact in wild and mysterious ways in microscopic photography, posing special challenges for the researcher and educator. With Adobe Photoshop
Customized Foam for Tools
 Table of contents Make sure that you have the latest version before using this document. o o o o o o o Overview of services offered and steps to follow (p.3) 1. Service : Cutting of foam for tools 2. Service
Table of contents Make sure that you have the latest version before using this document. o o o o o o o Overview of services offered and steps to follow (p.3) 1. Service : Cutting of foam for tools 2. Service
IncuCyte ZOOM Scratch Wound Processing Overview
 IncuCyte ZOOM Scratch Wound Processing Overview The IncuCyte ZOOM Scratch Wound assay utilizes the WoundMaker-IncuCyte ZOOM-ImageLock Plate system to analyze both 2D-migration and 3D-invasion in label-free,
IncuCyte ZOOM Scratch Wound Processing Overview The IncuCyte ZOOM Scratch Wound assay utilizes the WoundMaker-IncuCyte ZOOM-ImageLock Plate system to analyze both 2D-migration and 3D-invasion in label-free,
Submittals Quick Reference Guide
 This topic provides a reference for the Project Center Submittals activity center. Purpose The Submittals activity center in Newforma Contract Management enables you to effectively log submittals and track
This topic provides a reference for the Project Center Submittals activity center. Purpose The Submittals activity center in Newforma Contract Management enables you to effectively log submittals and track
Laboratory 2: Graphing
 Purpose It is often said that a picture is worth 1,000 words, or for scientists we might rephrase it to say that a graph is worth 1,000 words. Graphs are most often used to express data in a clear, concise
Purpose It is often said that a picture is worth 1,000 words, or for scientists we might rephrase it to say that a graph is worth 1,000 words. Graphs are most often used to express data in a clear, concise
Camera Base. User Guide. Version Mathias Tobler
 Camera Base Version 1.5.1 User Guide Mathias Tobler September, 2012 Table of contents 1 Introduction... 3 2 License... 4 3 Overview... 5 3.1 Data Management... 5 3.2 Analysis and outputs... 5 4 Installation...
Camera Base Version 1.5.1 User Guide Mathias Tobler September, 2012 Table of contents 1 Introduction... 3 2 License... 4 3 Overview... 5 3.1 Data Management... 5 3.2 Analysis and outputs... 5 4 Installation...
Stratigraphy Modeling Boreholes and Cross. Become familiar with boreholes and borehole cross sections in GMS
 v. 10.3 GMS 10.3 Tutorial Stratigraphy Modeling Boreholes and Cross Sections Become familiar with boreholes and borehole cross sections in GMS Objectives Learn how to import borehole data, construct a
v. 10.3 GMS 10.3 Tutorial Stratigraphy Modeling Boreholes and Cross Sections Become familiar with boreholes and borehole cross sections in GMS Objectives Learn how to import borehole data, construct a
Origami. for Joomla! Theme Documentation. Version 1.0 Last Updated: November 4, gothemeteam.com
 Origami for Joomla! Theme Documentation Version 1.0 Last Updated: November 4, 2011 gothemeteam.com Table of Contents Installation...3 Overview & Requirements...3 Quickstart Package...4 Site Logo...7 Changing
Origami for Joomla! Theme Documentation Version 1.0 Last Updated: November 4, 2011 gothemeteam.com Table of Contents Installation...3 Overview & Requirements...3 Quickstart Package...4 Site Logo...7 Changing
This tutorial will lead you through step-by-step to make the plot below using Excel.
 GES 131 Making Plots with Excel 1 / 6 This tutorial will lead you through step-by-step to make the plot below using Excel. Number of Non-Student Tickets vs. Student Tickets Y, Number of Non-Student Tickets
GES 131 Making Plots with Excel 1 / 6 This tutorial will lead you through step-by-step to make the plot below using Excel. Number of Non-Student Tickets vs. Student Tickets Y, Number of Non-Student Tickets
Section 3 Correlation and Regression - Worksheet
 The data are from the paper: Exploring Relationships in Body Dimensions Grete Heinz and Louis J. Peterson San José State University Roger W. Johnson and Carter J. Kerk South Dakota School of Mines and
The data are from the paper: Exploring Relationships in Body Dimensions Grete Heinz and Louis J. Peterson San José State University Roger W. Johnson and Carter J. Kerk South Dakota School of Mines and
How to generate different file formats
 How to generate different file formats Different mediums print, web, and video require different file formats. This guide describes how to generate appropriate file formats for these mediums by using Adobe
How to generate different file formats Different mediums print, web, and video require different file formats. This guide describes how to generate appropriate file formats for these mediums by using Adobe
GIS Module GMS 7.0 TUTORIALS. 1 Introduction. 1.1 Contents
 GMS 7.0 TUTORIALS 1 Introduction The GIS module can be used to display data from a GIS database directly in GMS without having to convert that data to GMS data types. Native GMS data such as grids and
GMS 7.0 TUTORIALS 1 Introduction The GIS module can be used to display data from a GIS database directly in GMS without having to convert that data to GMS data types. Native GMS data such as grids and
Grid Assembly. User guide. A plugin developed for microscopy non-overlapping images stitching, for the public-domain image analysis package ImageJ
 BIOIMAGING AND OPTIC PLATFORM Grid Assembly A plugin developed for microscopy non-overlapping images stitching, for the public-domain image analysis package ImageJ User guide March 2008 Introduction In
BIOIMAGING AND OPTIC PLATFORM Grid Assembly A plugin developed for microscopy non-overlapping images stitching, for the public-domain image analysis package ImageJ User guide March 2008 Introduction In
Overview. The Game Idea
 Page 1 of 19 Overview Even though GameMaker:Studio is easy to use, getting the hang of it can be a bit difficult at first, especially if you have had no prior experience of programming. This tutorial is
Page 1 of 19 Overview Even though GameMaker:Studio is easy to use, getting the hang of it can be a bit difficult at first, especially if you have had no prior experience of programming. This tutorial is
Technical Note. How to Use the Image Studio Software Small Animal Image Analysis. Developed for: Image Studio Software
 Technical Note How to Use the Image Studio Software Small Animal Image Analysis Developed for: Image Studio Software Please refer to your manual to confirm that this protocol is appropriate for the applications
Technical Note How to Use the Image Studio Software Small Animal Image Analysis Developed for: Image Studio Software Please refer to your manual to confirm that this protocol is appropriate for the applications
Financial Disclosure Management Release Notes
 Financial Disclosure Management 8.1.0.9 Release Notes Publication 1.0 October 2017 INTRODUCTION... 1 FDM 8.1.0.9 ENHANCEMENTS... 1 MANAGEMENT REPORTS... 2 NEW PILOT PROGRAM! MANAGEMENT REPORTS... 2 Key
Financial Disclosure Management 8.1.0.9 Release Notes Publication 1.0 October 2017 INTRODUCTION... 1 FDM 8.1.0.9 ENHANCEMENTS... 1 MANAGEMENT REPORTS... 2 NEW PILOT PROGRAM! MANAGEMENT REPORTS... 2 Key
Image Classification (Decision Rules and Classification)
 Exercise #5D Image Classification (Decision Rules and Classification) Objective Choose how pixels will be allocated to classes Learn how to evaluate the classification Once signatures have been defined
Exercise #5D Image Classification (Decision Rules and Classification) Objective Choose how pixels will be allocated to classes Learn how to evaluate the classification Once signatures have been defined
Zeiss Deconvolution Microscope: A Quick Guide
 Zeiss Deconvolution Microscope: A Quick Guide Start-up Uncover microscope. Do not put dust cover on the floor. Plug in both cameras. The default camera is the AxioCam HRm (monochrome camera) for fluorescence
Zeiss Deconvolution Microscope: A Quick Guide Start-up Uncover microscope. Do not put dust cover on the floor. Plug in both cameras. The default camera is the AxioCam HRm (monochrome camera) for fluorescence
New Features in Release 2.2
 Release 2.2 New Features 1 New Features in Release 2.2 This short manual gives an overview over the new features in the Software v. 2.2.0.8, Analysis Software v. 1.2.0.4). release v. 2.2 (Acquisition 2
Release 2.2 New Features 1 New Features in Release 2.2 This short manual gives an overview over the new features in the Software v. 2.2.0.8, Analysis Software v. 1.2.0.4). release v. 2.2 (Acquisition 2
KODAK DIGITAL ROC Professional Plug-In 2.1
 KODAK DIGITAL ROC Professional Plug-In 2.1 Installing Kodak's DIGITAL ROC Professional Plug-In If you have not downloaded and installed DIGITAL ROC Professional, go to: http://www.asf.com/download/ Download
KODAK DIGITAL ROC Professional Plug-In 2.1 Installing Kodak's DIGITAL ROC Professional Plug-In If you have not downloaded and installed DIGITAL ROC Professional, go to: http://www.asf.com/download/ Download
v. 8.0 GMS 8.0 Tutorial GIS Module Shapefile import, display, and conversion Prerequisite Tutorials None Time minutes
 v. 8.0 GMS 8.0 Tutorial Shapefile import, display, and conversion Objectives Learn how to import and display shapefiles with and without ArcObjects. Convert the shapefiles to GMS feature objects. Prerequisite
v. 8.0 GMS 8.0 Tutorial Shapefile import, display, and conversion Objectives Learn how to import and display shapefiles with and without ArcObjects. Convert the shapefiles to GMS feature objects. Prerequisite
1. What is SENSE Batch
 1. What is SENSE Batch 1.1. Introduction SENSE Batch is processing software for thermal images and sequences. It is a modern software which automates repetitive tasks with thermal images. The most important
1. What is SENSE Batch 1.1. Introduction SENSE Batch is processing software for thermal images and sequences. It is a modern software which automates repetitive tasks with thermal images. The most important
Photoshop CC 2018 Essential Skills
 Photoshop CC 2018 Essential Skills Adobe Photoshop Creative Cloud 2018 University Information Technology Services Learning Technology, Training, Audiovisual and Outreach Copyright 2018 KSU Division of
Photoshop CC 2018 Essential Skills Adobe Photoshop Creative Cloud 2018 University Information Technology Services Learning Technology, Training, Audiovisual and Outreach Copyright 2018 KSU Division of
LSM 780 Confocal Microscope Standard Operation Protocol
 LSM 780 Confocal Microscope Standard Operation Protocol Basic Operation Turning on the system 1. Sign on log sheet according to Actual start time 2. Check Compressed Air supply for the air table 3. Switch
LSM 780 Confocal Microscope Standard Operation Protocol Basic Operation Turning on the system 1. Sign on log sheet according to Actual start time 2. Check Compressed Air supply for the air table 3. Switch
ADDING A RAINBOW TO A PHOTOGRAPH
 ADDING A RAINBOW TO A PHOTOGRAPH This assignment will cover how to add a simple rainbow (or if you want to go crazy, a double rainbow) to any photograph. This will give us some great work with gradients,
ADDING A RAINBOW TO A PHOTOGRAPH This assignment will cover how to add a simple rainbow (or if you want to go crazy, a double rainbow) to any photograph. This will give us some great work with gradients,
How to blend, feather, and smooth
 How to blend, feather, and smooth Quite often, you need to select part of an image to modify it. When you select uniform geometric areas squares, circles, ovals, rectangles you don t need to worry too
How to blend, feather, and smooth Quite often, you need to select part of an image to modify it. When you select uniform geometric areas squares, circles, ovals, rectangles you don t need to worry too
Tube Formation Analysis in the Automated Cellular Analysis System
 Tube Formation Analysis in the Automated Cellular Analysis System 1 ibidi GmbH, Version 1.0, 2017-07-14 Table of Content Specifications... 3 Step-by-Step Guide... 4 Analysis... 6 Example Report Job...
Tube Formation Analysis in the Automated Cellular Analysis System 1 ibidi GmbH, Version 1.0, 2017-07-14 Table of Content Specifications... 3 Step-by-Step Guide... 4 Analysis... 6 Example Report Job...
How to Make a Run Chart in Excel
 How to Make a Run Chart in Excel While there are some statistical programs that you can use to make a run chart, it is simple to make in Excel, using Excel s built-in chart functions. The following are
How to Make a Run Chart in Excel While there are some statistical programs that you can use to make a run chart, it is simple to make in Excel, using Excel s built-in chart functions. The following are
MY ASTROPHOTOGRAPHY WORKFLOW Scott J. Davis June 21, 2012
 Table of Contents Image Acquisition Types 2 Image Acquisition Exposure 3 Image Acquisition Some Extra Notes 4 Stacking Setup 5 Stacking 7 Preparing for Post Processing 8 Preparing your Photoshop File 9
Table of Contents Image Acquisition Types 2 Image Acquisition Exposure 3 Image Acquisition Some Extra Notes 4 Stacking Setup 5 Stacking 7 Preparing for Post Processing 8 Preparing your Photoshop File 9
Contents Downloading and installing IrfanView.. 1
 Contents Downloading and installing IrfanView.. 1 Rotating Images. 2 Resizing/Resample Images.. 3 Cropping Images.... 4 Thumbnail View.. 5 Effects.. 6 Downloading and installing IrfanView IrFanView is
Contents Downloading and installing IrfanView.. 1 Rotating Images. 2 Resizing/Resample Images.. 3 Cropping Images.... 4 Thumbnail View.. 5 Effects.. 6 Downloading and installing IrfanView IrFanView is
Excel Manual X Axis Scale Start At Graph
 Excel Manual X Axis Scale Start At 0 2010 Graph But when I plot them by XY chart in Excel (2003), it looks like a rectangle, even if I havesame for both X, and Y axes, and I can see the X and Y data maximum
Excel Manual X Axis Scale Start At 0 2010 Graph But when I plot them by XY chart in Excel (2003), it looks like a rectangle, even if I havesame for both X, and Y axes, and I can see the X and Y data maximum
MicroLab 500-series Getting Started
 MicroLab 500-series Getting Started 2 Contents CHAPTER 1: Getting Started Connecting the Hardware....6 Installing the USB driver......6 Installing the Software.....8 Starting a new Experiment...8 CHAPTER
MicroLab 500-series Getting Started 2 Contents CHAPTER 1: Getting Started Connecting the Hardware....6 Installing the USB driver......6 Installing the Software.....8 Starting a new Experiment...8 CHAPTER
Applying mathematics to digital image processing using a spreadsheet
 Jeff Waldock Applying mathematics to digital image processing using a spreadsheet Jeff Waldock Department of Engineering and Mathematics Sheffield Hallam University j.waldock@shu.ac.uk Introduction When
Jeff Waldock Applying mathematics to digital image processing using a spreadsheet Jeff Waldock Department of Engineering and Mathematics Sheffield Hallam University j.waldock@shu.ac.uk Introduction When
CONTENT INTRODUCTION BASIC CONCEPTS Creating an element of a black-and white line drawing DRAWING STROKES...
 USER MANUAL CONTENT INTRODUCTION... 3 1 BASIC CONCEPTS... 3 2 QUICK START... 7 2.1 Creating an element of a black-and white line drawing... 7 3 DRAWING STROKES... 15 3.1 Creating a group of strokes...
USER MANUAL CONTENT INTRODUCTION... 3 1 BASIC CONCEPTS... 3 2 QUICK START... 7 2.1 Creating an element of a black-and white line drawing... 7 3 DRAWING STROKES... 15 3.1 Creating a group of strokes...
Infographics: Display Data for Easy Interpretation
 Infographics: Display Data for Easy Interpretation Course objectives: Create new infographics Customise layouts Edit content using text, images, media, charts and maps Publish, Present and Print Student
Infographics: Display Data for Easy Interpretation Course objectives: Create new infographics Customise layouts Edit content using text, images, media, charts and maps Publish, Present and Print Student
10.2. Scanning Document Camera Scoring. Page 1 of 5. How do I score answer sheets using a document camera? STEP 1
 Step by Step How do I score answer sheets using a document camera? STEP 1 Click on the Assessment icon in the top navigation bar. STEP 2 To locate your assessment in an assessment list, first select the
Step by Step How do I score answer sheets using a document camera? STEP 1 Click on the Assessment icon in the top navigation bar. STEP 2 To locate your assessment in an assessment list, first select the
11 Advanced Layer Techniques
 11 Advanced Layer Techniques After you ve learned basic layer techniques, you can create more complex effects in your artwork using layer masks, path groups, filters, adjustment layers, and more style
11 Advanced Layer Techniques After you ve learned basic layer techniques, you can create more complex effects in your artwork using layer masks, path groups, filters, adjustment layers, and more style
Using Adobe Photoshop to enhance the image quality. Assistant course web site:
 Using Adobe Photoshop to enhance the image quality Assistant course web site: http://www.arches.uga.edu/~skwang/edit6170/course.htm Content Introduction 2 Unit1: Scan images 3 Lesson 1-1: Preparations
Using Adobe Photoshop to enhance the image quality Assistant course web site: http://www.arches.uga.edu/~skwang/edit6170/course.htm Content Introduction 2 Unit1: Scan images 3 Lesson 1-1: Preparations
(RGB images only) Ctrl-click (Windows) or Command-click (Mac OS) a pixel in the image.
 PHOTOSHOP TOOLS USING CURVES: To adjust tonality with Curves, do one of the following: Choose Image > Adjustments > Curves. Choose Layer > New Adjustment Layer > Curves. Click OK in the New Layer dialog
PHOTOSHOP TOOLS USING CURVES: To adjust tonality with Curves, do one of the following: Choose Image > Adjustments > Curves. Choose Layer > New Adjustment Layer > Curves. Click OK in the New Layer dialog
Image Processing Tutorial Basic Concepts
 Image Processing Tutorial Basic Concepts CCDWare Publishing http://www.ccdware.com 2005 CCDWare Publishing Table of Contents Introduction... 3 Starting CCDStack... 4 Creating Calibration Frames... 5 Create
Image Processing Tutorial Basic Concepts CCDWare Publishing http://www.ccdware.com 2005 CCDWare Publishing Table of Contents Introduction... 3 Starting CCDStack... 4 Creating Calibration Frames... 5 Create
Drawing with precision
 Drawing with precision Welcome to Corel DESIGNER, a comprehensive vector-based drawing application for creating technical graphics. Precision is essential in creating technical graphics. This tutorial
Drawing with precision Welcome to Corel DESIGNER, a comprehensive vector-based drawing application for creating technical graphics. Precision is essential in creating technical graphics. This tutorial
