KINALYZER, a computer program for reconstructing sibling groups
|
|
|
- Beverly Owen
- 6 years ago
- Views:
Transcription
1 Molecular Ecology Resources (2009) 9, doi: /j x Blackwell Publishing Ltd COMPUTER PROGRAM NOTE KINALYZER, a computer program for reconstructing sibling groups M. V. ASHLEY,* I. C. CABALLERO,* W. CHAOVALITWONGSE, B. DASGUPTA, P. GOVINDAN, S. I. SHEIKH and T. Y. BERGER-WOLF *Department of Biological Sciences, M/C 066, University of Illinois at Chicago, 845 W. Taylor Street, Chicago, IL 60607, USA, Department of Industrial and Systems Engineering, Rutgers University, CoRE Building, 96 Frelinghuysen Road, Piscataway, NJ 08854, USA, Department of Computer Science, M/C 152, University of Illinois at Chicago, 851 S. Morgan, Chicago, IL 60607, USA Abstract A software suite KINALYZER reconstructs full-sibling groups without parental information using data from codominant marker loci such as microsatellites. KINALYZER utilizes a new algorithm for sibling reconstruction in diploid organisms based on combinatorial optimization. KINALYZER makes use of a Minimum 2-Allele Set Cover approach based on Mendelian inheritance rules and finds the smallest number of sibling groups that contain all the individuals in the sample. Also available is a Greedy Consensus approach that reconstructs sibgroups using subsets of loci and finds the consensus of the partial solutions. Unlike likelihood methods for sibling reconstruction, KINALYZER does not require information about population allele frequencies and it makes no assumptions regarding the mating system of the species. KINALYZER is freely available as a web-based service. Keywords: combinatorial optimization, kinship, microsatellite DNA, sibgroup reconstruction, sibling Received 13 October 2008; revision accepted 19 December 2008 Kinship reconstruction using codominant markers such as DNA microsatellites has become an important component of many investigations of wild populations (e.g. Pemberton 2008). The aim of kinship or pedigree reconstruction is to identify family groups, including parents, siblings, and higher-order relationships. Several methods and software for parentage assignment (maternity and paternity) are widely used and available (reviewed in Blouin 2003). Sibgroup reconstruction, with no or only partial parental information, is conceptually and computationally more difficult than parentage assignment, and sibling reconstruction studies have lagged behind those that use parentage assignment. More accurate and efficient approaches for sibgroup reconstruction are needed for cases where field studies sample cohorts of offspring, but obtaining samples of some or all candidate parents is less feasible. Recent examples include sampling of juvenile lemon sharks from nursery lagoons (Feldheim et al. 2004), brood parasitic cowbird nestlings sampled from host nests (Strausberger & Ashley 2003; Strausberger & Ashley 2005), wood duck eggs Correspondence: Mary V. Ashley, Fax: (312) ; ashley@uic.edu and nestlings (Roy Nielsen et al. 2006) and kelp bass larval cohorts (Selkoe et al. 2006). More studies would likely employ sibling reconstruction in data analysis if more robust approaches were widely available. There have been several approaches taken for reconstructing full-sibling groups, although none has emerged as a clear favourite among molecular ecologists. Most sibgroup reconstruction methods use statistical likelihood models (Thomas & Hill 2000; Smith et al. 2001; Konovalov et al. 2004; Wang 2004) and, thus, rely on accurate estimates of underlying population allele frequencies, which may be difficult to obtain independently of the sample of potential siblings. The software Pedigree, available for use as on online service, employs Markov chain Monte Carlo (MCMC) methods for sib reconstruction by maximizing the joint likelihood of the entire sibship reconstruction rather than the pairwise relatedness ratio (Smith et al. 2001). Family Finder (Beyer & May 2003) uses a graph-based model, with edges representing pairwise sibling relationships that are weighted by the relationship likelihood (Goodnight & Queller 1999). Graph clusters corresponding to sibling groups are identified by finding light edge cuts. Most of the available methods do not allow for genotyping errors or
2 1128 COMPUTER PROGRAM NOTE mutations (Almudevar & Field 1999; Thomas & Hill 2000; Smith et al. 2001; Konovalov et al. 2004), yet errors are likely to occur at least at low frequencies in any large microsatellite data set. One exception is COLONY (Wang 2004). COL- ONY uses simulated annealing to exhaustively search for sibling reconstruction based on overall maximum likelihood, accounting for genotyping errors in the process. In a test of four methods using simulated data, Butler et al. (2004) conclude that none of the algorithms performed well over the range of conditions tested, which included varying number of loci and alleles, family distributions, and errors in the data. In our recent review (Ashley et al. 2008) testing sibling reconstruction methods, we found that among statistical methods, COLONY (Wang 2004) accurately reconstructed siblings when sufficient number of loci were sampled (at least six) and allele diversity was high. However, COLONY is limited by an assumption of one gender monogamy and is too computationally demanding for analysis of moderate to large data sets in a reasonable time. In contrast to statistical likelihood approaches, combinatorial approaches construct potential sibling groups using only Mendelian properties (Almudevar & Field 1999; Berger-Wolf et al. 2007; Sheikh et al. 2008) and search for the most parsimonious solution, such as the smallest number of mating pairs or parents. The method of Almudevar & Field (1999) uses a heuristic approach (rather than established computational optimization methods) to find a local optimum, but is not guaranteed to find the overall best solution (i.e. the smallest number of mating pairs). Alternatively, KINALYZER uses a combinatorial approach based upon a simple rule for allele inheritance in diploid organisms: an offspring inherits one allele from each of its parents for each locus. This rule of Mendelian inheritance introduces a necessary constraint on full-sibling groups in the absence of genotyping errors or mutations: the 2-allele property (Berger-Wolf et al. 2007; Ashley et al. 2008; Sheikh et al. 2008). The 2-allele property states that there exists an assignment of individual alleles within a locus to maternal and paternal parents such that the number of distinct alleles assigned to each parent at this locus does not exceed two. Barring mutation or genotyping error, any sibling group must satisfy this constraint. Formally, a diploid individual i sampled at l loci is represented by its l pairs of alleles: i =[(a i1, b i1 ), (a i2, b i2 ),..., (a il, b il )]. A set of individuals S in a population sample U has the 2-allele property if for each individual i in S at each locus there exists an assignment of the two alleles a ij =c ij and b ij = «ij or a ij = «ij and b ij =c ij such that 1 j l: { c } 2and { c } 2 i S ij i S KINALYZER employs the Minimum 2-Allele Set Cover approach to find the smallest number of sibgroups S 1,..., S m such that each sibgroup consists of a subset of individuals ij in U, the 2-allele property is satisfied for every sibgroup, and every individual is contained in at least one sibgroup (US i = U). This smallest number of feasible sibgroups (that satisfy the 2-allele constraint) is found using a combinatorial optimization technique to select the fewest possible sibgroups. Combinatorial optimization is a class of problems where the qualitative (combinatorial) structure is more important than the numerical values. Such problems are defined by structural constraints on potential solutions and a cost associated with each solution. The objective is to find a solution which optimizes (minimizes or maximizes) the cost. For the Minimum 2-Allele Set Cover, a feasible solution is any partition of individuals into groups that satisfy the 2- allele property (the structural constraint). The cost of each solution is the number of groups, and the objective is to find the solution with the smallest number of groups. Such combinatorial optimization problems are typically provably hard (computationally infeasible) and the Minimum 2-Allele Set Cover is no exception (Ashley et al. 2008). There are a wide variety of computational techniques that solve combinatorial optimization problems (Cook et al. 1997; Papadimitriou & Steiglitz 1998). While any technique applied to any particular combinatorial problem may take a long, even infeasible, time to find a solution, such solution, when found, is guaranteed to be optimal. Combinatorial optimization in KINALYZER is based on the implementation of CPLEX, 1 a commercially available optimization software package. CPLEX employs multiple optimization algorithms including simplex, cutting plane, interior point, barrier, and branch-and-bound to solve difficult combinatorial optimization problems and is guaranteed to find the overall optimum. Note that while every optimization technique will find the overall optimum, some may take longer than others on any given problem. The main advantage of CPLEX is that the most efficient optimization algorithm is used based on the structure of the problem. The computational objective of minimizing the number of sibling groups is formally equivalent to minimizing the number of mating pairs, and provides the most parsimonious reconstruction goal. The solution satisfies Mendelian rules of inheritance and is guaranteed to be the optimal solution if the underlying objective of the smallest number of matings is correct. As we develop computational methods with different biological objectives, such as minimizing the number of fathers or maximizing family size, these will be added to the KINALYZER software suite. KINALYZER also includes a consensus-based approach ( Greedy Consensus ) that discards individual loci one at a time and reconstructs solutions using the remaining loci. The final solution output is a consensus of the partial solutions. The consensus is calculated by first computing the 1 CPLEX is a registered trademark of ILOG.
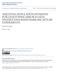
3 COMPUTER PROGRAM NOTE 1129 Fig. 1 Comparison of accuracy of different sib reconstruction approaches. A C show results using simulated data. Simulated data was generated by first randomly generating parent pairs based on population parameters (alleles per locus, number of loci), and then randomly generating their offspring. The number of male/female parents, families and the offspring per family were varied as indicated to generate the simulated populations. Each algorithm was run on simulated data sets created with the specified parameters until the mean and standard deviation of error rates were stable for 10 consecutive iterations. Accuracy was calculated by the Gusfield Partition Distance (Gusfield 2002) between the algorithm s reconstruction and the known sibling relationships (see Ashley et al for further details on simulations). 2-Allele refers to the minimum set cover implemented in KINALYZER. G. Consensus (Sheikh et al. 2008) refers to the consensus approach described in the text that is also available in KINALYZER. D shows analysis of real data where sibling relationships (from controlled crosses) were known for tiger shrimp Penaeus monodon (Jerry etal. 2006), Atlantic salmon (Herbinger et al. 1999) and the polygynous ants Leptothorax acervorum (Hammond et al. 1999). groups that are in common and then greedily (taking the best immediate, or local, solution) merging the nearest pair of groups iteratively. Distance is computed based on costs associated with errors and allelic information shared (see Sheikh et al for details). Using simulated and real data with known sibling relationships, we have compared available sibling reconstruction software (Berger-Wolf et al. 2007; Ashley et al. 2008; Sheikh et al. 2008). An example of a comparison of KINALYZER to three of the commonly used statistical methods, Pedigree (Smith et al. 2001), COLONY (Wang 2004) and Family Finder (Beyer & May 2003) is shown in Fig. 1. Error rates were calculated using the Gusfield partition distance (Gusfield 2002), the minimum number of individuals to remove in order to make the two partitions (the reconstructed sibgoups and the actual sibgroups) equivalent. Overall, KINALYZER performed as well or better than other methods on a wide range of data set parameters. It remains robust even when the allelic diversity is low (Fig. 1A), the number of loci sampled is small (Fig. 1B), and there are genotyping errors (Fig. 1C). It also performed well on three different biological data sets tested, while three other available methods were less consistent (Fig. 1D) (Berger-Wolf et al. 2007; Ashley et al. 2008). KINALYZER is a web-based program that requires an input file comprising the individuals and genotypes to analyse. Because many users will already be familiar with Kinship (Goodnight & Queller 1999), KinGroup (Konovalov et al. 2004), gerud ( Jones 2005) or Cervus (Marshall et al. 1998) input files, we have preserved these for KINALYZER with the exception that no population allele frequencies are needed to run the program. To upload the genotype data file, the user logs into the website and provides their name and address (Fig. 2) which are necessary to deliver the results. Currently KINALYZER only accepts.csv input files from Excel. Three- or two-digit coded alleles are automatically recognized by the program and missing data should be coded as 1 (failure to do so will prompt the program to display a message to correct the format). The columns should correspond to the identity of individuals and name of loci. The input file may contain extra columns not used by KINALYZER (i.e. sex, locality, group, etc.); the software has an option to disable them prior to uploading the file. There is no limit to sample sizes or number of loci. The web address for KINALYZER is kinalyzer.cs.uic.edu/. Because this is a web-based program, the user will be given an input file number and the results are delivered via
4 1130 COMPUTER PROGRAM NOTE Fig. 2 Screenshot of KINALYZER software interface, showing user login, data upload and formatting windows, and confirmation of submissions with information on receiving the results. . The time to analyse the data will depend on how many jobs the server is processing at that time. Users can find out about the status of the queue job online at any time (using the input file number). The output file will show individuals divided into full-sib groups (sets). Each one of these sets will list the individuals by the identification name (or number) that was provided in the input file. Because sibling reconstruction is still a developing field, we recommend that investigators try different approaches, and select an appropriate procedure based on their study systems and the assumptions and limitations of currently available methods. No single method is guaranteed to provide the correct answer, but we favour the 2-allele method implemented in KINALYZER because of the available methods, it makes the fewest number of assumptions and performs well over a wide range of data parameters. It is, therefore, a good general method, especially when few loci are sampled or the allelic diversity is low (Fig. 1). The Greedy Consensus method was found to be highly accurate in tests using benchmark data, especially when allelic variation was low, and was highly tolerant of genotyping errors and mutations (Sheikh et al. 2008). As mentioned above, other reconstruction objectives will also be added to KINALYZER as they are developed. Acknowledgements The development of KINALYZER was supported by NSF IIS and IIS (Berger-Wolf, Ashley, Chaovalitwongse, DasGupta), NSF CCF (Chaovalitwongse), NSF IIS CAREER (Berger-Wolf), NSF DBI (DasGupta), NSF IIS (DasGupta), DIMACS special focus on Computational and Mathematical Epidemiology (DasGupta) and a Fulbright Scholarship (Sheikh). Numerous people have shared their data for testing KINALYZER, including Jeffrey Connor, the Atlantic
5 COMPUTER PROGRAM NOTE 1131 Salmon Federation, Dean Jerry and Stuart Barker. We also thank Anthony Almudevar, Bernie May, and Dmitry Konovalov for sharing their software. References Almudevar A, Field C (1999) Estimation of single-generation sibling relationships based on DNA markers. Journal of Agricultural Biological and Environmental Statistics, 4, Ashley MV, Berger-Wolf TY, Caballero IC, Chaovalitwongse W, DasGupta B, Sheikh SI (2008) Full sibling reconstruction in wild populations from microsatellite genetic markers. In: Computational Biology: New Research. Nova Science Publishers, Hauppauge, New York. Berger-Wolf TY, Sheikh SI, DasGupta B, Ashley MV, Caballero IC, Chaovalitwongse W, Putrevu SL (2007) Reconstructing sibling relationships in wild populations. Bioinformatics, 23, I49 I56. Beyer J, May B (2003) A graph-theoretic approach to the partition of individuals into full-sib families. Molecular Ecology, 12, Blouin MS (2003) DNA-based methods for pedigree reconstruction and kinship analysis in natural populations. Trends in Ecology & Evolution, 18, Butler K, Field C, Herbinger CM, Smith BR (2004) Accuracy, efficiency and robustness of four algorithms allowing full sibship reconstruction from DNA marker data. Molecular Ecology, 13, Cook WJ, Cunningham WH, Pulleyblank WR, Schrijver A (1997) Combinatorial Optimization, 1st edn. John Wiley & Sons, New York. Feldheim KA, Gruber SH, Ashley MV (2004) Reconstruction of parental microsatellite genotypes reveals female polyandry and philopatry in the lemon shark, Negaprion brevirostris. Evolution, 58, Goodnight KF, Queller DC (1999) Computer software for performing likelihood tests of pedigree relationship using genetic markers. Molecular Ecology, 8, Gusfield D (2002) Partition-distance: a problem and class of perfect graphs arising in clustering. Information Processing Letters, 82, Hammond RL, Bourke AFG, Bruford MW (1999) Mating frequency and mating system of the polygynous ant, Leptothorax acervorum. Molecular Ecology, 10, Herbinger C, O Reilly P, Doyle R, Wright J, O'Flynn F (1999) Early growth performance of Atlantic salmon full-sib families reared in single family tanks or in mixed family tanks. Aquaculture, 173, Jerry D, Evans B, Kenway M, Wilson K (2006) Development of a microsatellite DNA parentage marker suite for black tiger shrimp Penaeus monodon. Aquaculture, Jones AG (2005) gerud 2.0: a computer program for the reconstruction of parental genotypes from half-sib progeny arrays with known or unknown parents. Molecular Ecology Notes, 5, Konovalov DA, Manning C, Henshaw MT (2004) KinGroup: a program for pedigree relationship reconstruction and kin group assignments using genetic markers. Molecular Ecology Notes, 4, Marshall TC, Slate J, Kruuk LEB, Pemberton JM (1998) Statistical confidence for likelihood-based paternity inference in natural populations. Molecular Ecology, 7, Papadimitriou CH, Steiglitz K (1998) Combinatorial Optimization: Algorithms and Complexity. Dover Publications, Mineola, New York. Pemberton JM (2008) Wild pedigrees: the way forward. Proceedings of the Royal Society Series B, 275, Roy Nielsen C, Gates R, Parker P (2006) Intraspecific nest parasitism of wood ducks in natural cavities: Comparisons with nest boxes. Journal of Wildlife Management, 70, Selkoe KA, Gaines SD, Caselle JE, Warner RR (2006) Current shifts and kin aggregation explain genetic patchiness in fish recruits. Ecology, 87, Sheikh SI, Berger-Wolf TY, Chaovalitwongse W, Ashley MV (2008) Error-tolerant sibship reconstruction in wild populations 7th Annual International Conference on Computational Systems Bioinformatics. Smith BR, Herbinger CM, Merry HR (2001) Accurate partition of individuals into full-sib families from genetic data without parental information. Genetics, 158, Strausberger BM, Ashley MV (2003) Breeding biology of brood parasitic brown-headed cowbirds (Molothrus ater) characterized by parent-offspring and sibling-group reconstruction. Auk, 120, Strausberger BM, Ashley MV (2005) Host use strategies of individual female brown-headed cowbirds Molothrus ater in a diverse avian community. Journal of Avian Biology, 36, Thomas SC, Hill WG (2000) Estimating quantitative genetic parameters using sibships reconstructed from marker data. Genetics, 155, Wang JL (2004) Sibship reconstruction from genetic data with typing errors. Genetics, 166,
COMBINATORIAL RECONSTRUCTION OF HALF-SIBLING GROUPS
 COMBINATORIAL RECONSTRUCTION OF HALF-SIBLING GROUPS Saad I. Sheikh, Tanya Y. Berger-Wolf, Ashfaq A. Khokhar Department of Computer Science, University of Illinois at Chicago, 851 S. Morgan St (M/C 152),
COMBINATORIAL RECONSTRUCTION OF HALF-SIBLING GROUPS Saad I. Sheikh, Tanya Y. Berger-Wolf, Ashfaq A. Khokhar Department of Computer Science, University of Illinois at Chicago, 851 S. Morgan St (M/C 152),
COMBINATORIAL RECONSTRUCTION OF HALF-SIBLING GROUPS
 COMBINATORIAL RECONSTRUCTION OF HALF-SIBLING GROUPS Saad I. Sheikh, Tanya Y. Berger-Wolf, Ashfaq A. Khokhar Dept. of Computer Science, University of Illinois at Chicago, 851 S. Morgan St (M/C 152), Chicago,
COMBINATORIAL RECONSTRUCTION OF HALF-SIBLING GROUPS Saad I. Sheikh, Tanya Y. Berger-Wolf, Ashfaq A. Khokhar Dept. of Computer Science, University of Illinois at Chicago, 851 S. Morgan St (M/C 152), Chicago,
Methods of Parentage Analysis in Natural Populations
 Methods of Parentage Analysis in Natural Populations Using molecular markers, estimates of genetic maternity or paternity can be achieved by excluding as parents all adults whose genotypes are incompatible
Methods of Parentage Analysis in Natural Populations Using molecular markers, estimates of genetic maternity or paternity can be achieved by excluding as parents all adults whose genotypes are incompatible
Lecture 6: Inbreeding. September 10, 2012
 Lecture 6: Inbreeding September 0, 202 Announcements Hari s New Office Hours Tues 5-6 pm Wed 3-4 pm Fri 2-3 pm In computer lab 3306 LSB Last Time More Hardy-Weinberg Calculations Merle Patterning in Dogs:
Lecture 6: Inbreeding September 0, 202 Announcements Hari s New Office Hours Tues 5-6 pm Wed 3-4 pm Fri 2-3 pm In computer lab 3306 LSB Last Time More Hardy-Weinberg Calculations Merle Patterning in Dogs:
Pedigree Reconstruction using Identity by Descent
 Pedigree Reconstruction using Identity by Descent Bonnie Kirkpatrick Electrical Engineering and Computer Sciences University of California at Berkeley Technical Report No. UCB/EECS-2010-43 http://www.eecs.berkeley.edu/pubs/techrpts/2010/eecs-2010-43.html
Pedigree Reconstruction using Identity by Descent Bonnie Kirkpatrick Electrical Engineering and Computer Sciences University of California at Berkeley Technical Report No. UCB/EECS-2010-43 http://www.eecs.berkeley.edu/pubs/techrpts/2010/eecs-2010-43.html
Revising how the computer program
 Molecular Ecology (2007) 6, 099 06 doi: 0./j.365-294X.2007.03089.x Revising how the computer program Blackwell Publishing Ltd CERVUS accommodates genotyping error increases success in paternity assignment
Molecular Ecology (2007) 6, 099 06 doi: 0./j.365-294X.2007.03089.x Revising how the computer program Blackwell Publishing Ltd CERVUS accommodates genotyping error increases success in paternity assignment
Detection of Misspecified Relationships in Inbred and Outbred Pedigrees
 Detection of Misspecified Relationships in Inbred and Outbred Pedigrees Lei Sun 1, Mark Abney 1,2, Mary Sara McPeek 1,2 1 Department of Statistics, 2 Department of Human Genetics, University of Chicago,
Detection of Misspecified Relationships in Inbred and Outbred Pedigrees Lei Sun 1, Mark Abney 1,2, Mary Sara McPeek 1,2 1 Department of Statistics, 2 Department of Human Genetics, University of Chicago,
BIOINFORMATICS ORIGINAL PAPER
 BIOINFORMATICS ORIGINAL PAPER Vol. 25 no. 6 29, pages 234 239 doi:.93/bioinformatics/btp64 Genetics and population analysis FRANz: reconstruction of wild multi-generation pedigrees Markus Riester,, Peter
BIOINFORMATICS ORIGINAL PAPER Vol. 25 no. 6 29, pages 234 239 doi:.93/bioinformatics/btp64 Genetics and population analysis FRANz: reconstruction of wild multi-generation pedigrees Markus Riester,, Peter
Coalescence. Outline History. History, Model, and Application. Coalescence. The Model. Application
 Coalescence History, Model, and Application Outline History Origins of theory/approach Trace the incorporation of other s ideas Coalescence Definition and descriptions The Model Assumptions and Uses Application
Coalescence History, Model, and Application Outline History Origins of theory/approach Trace the incorporation of other s ideas Coalescence Definition and descriptions The Model Assumptions and Uses Application
KINSHIP ANALYSIS AND HUMAN IDENTIFICATION IN MASS DISASTERS: THE USE OF MDKAP FOR THE WORLD TRADE CENTER TRAGEDY
 1 KINSHIP ANALYSIS AND HUMAN IDENTIFICATION IN MASS DISASTERS: THE USE OF MDKAP FOR THE WORLD TRADE CENTER TRAGEDY Benoît Leclair 1, Steve Niezgoda 2, George R. Carmody 3 and Robert C. Shaler 4 1 Myriad
1 KINSHIP ANALYSIS AND HUMAN IDENTIFICATION IN MASS DISASTERS: THE USE OF MDKAP FOR THE WORLD TRADE CENTER TRAGEDY Benoît Leclair 1, Steve Niezgoda 2, George R. Carmody 3 and Robert C. Shaler 4 1 Myriad
ICMP DNA REPORTS GUIDE
 ICMP DNA REPORTS GUIDE Distribution: General Sarajevo, 16 th December 2010 GUIDE TO ICMP DNA REPORTS 1. Purpose of This Document 1. The International Commission on Missing Persons (ICMP) endeavors to secure
ICMP DNA REPORTS GUIDE Distribution: General Sarajevo, 16 th December 2010 GUIDE TO ICMP DNA REPORTS 1. Purpose of This Document 1. The International Commission on Missing Persons (ICMP) endeavors to secure
Relative accuracy of three common methods of parentage analysis in natural populations
 Molecular Ecology (13) 22, 1158 117 doi: 1.1111/mec.12138 Relative accuracy of three common methods of parentage analysis in natural populations HUGO B. HARRISON,* 1 PABLO SAENZ-AGUDELO, 1 SERGE PLANES,
Molecular Ecology (13) 22, 1158 117 doi: 1.1111/mec.12138 Relative accuracy of three common methods of parentage analysis in natural populations HUGO B. HARRISON,* 1 PABLO SAENZ-AGUDELO, 1 SERGE PLANES,
Comparative method, coalescents, and the future
 Comparative method, coalescents, and the future Joe Felsenstein Depts. of Genome Sciences and of Biology, University of Washington Comparative method, coalescents, and the future p.1/36 Correlation of
Comparative method, coalescents, and the future Joe Felsenstein Depts. of Genome Sciences and of Biology, University of Washington Comparative method, coalescents, and the future p.1/36 Correlation of
Optimum contribution selection conserves genetic diversity better than random selection in small populations with overlapping generations
 Optimum contribution selection conserves genetic diversity better than random selection in small populations with overlapping generations K. Stachowicz 12*, A. C. Sørensen 23 and P. Berg 3 1 Department
Optimum contribution selection conserves genetic diversity better than random selection in small populations with overlapping generations K. Stachowicz 12*, A. C. Sørensen 23 and P. Berg 3 1 Department
Large scale kinship:familial Searching and DVI. Seoul, ISFG workshop
 Large scale kinship:familial Searching and DVI Seoul, ISFG workshop 29 August 2017 Large scale kinship Familial Searching: search for a relative of an unidentified offender whose profile is available in
Large scale kinship:familial Searching and DVI Seoul, ISFG workshop 29 August 2017 Large scale kinship Familial Searching: search for a relative of an unidentified offender whose profile is available in
Population Structure and Genealogies
 Population Structure and Genealogies One of the key properties of Kingman s coalescent is that each pair of lineages is equally likely to coalesce whenever a coalescent event occurs. This condition is
Population Structure and Genealogies One of the key properties of Kingman s coalescent is that each pair of lineages is equally likely to coalesce whenever a coalescent event occurs. This condition is
Kinship and Population Subdivision
 Kinship and Population Subdivision Henry Harpending University of Utah The coefficient of kinship between two diploid organisms describes their overall genetic similarity to each other relative to some
Kinship and Population Subdivision Henry Harpending University of Utah The coefficient of kinship between two diploid organisms describes their overall genetic similarity to each other relative to some
Chapter 2: Genes in Pedigrees
 Chapter 2: Genes in Pedigrees Chapter 2-0 2.1 Pedigree definitions and terminology 2-1 2.2 Gene identity by descent (ibd) 2-5 2.3 ibd of more than 2 genes 2-14 2.4 Data on relatives 2-21 2.1.1 GRAPHICAL
Chapter 2: Genes in Pedigrees Chapter 2-0 2.1 Pedigree definitions and terminology 2-1 2.2 Gene identity by descent (ibd) 2-5 2.3 ibd of more than 2 genes 2-14 2.4 Data on relatives 2-21 2.1.1 GRAPHICAL
Maximum likelihood pedigree reconstruction using integer programming
 Maximum likelihood pedigree reconstruction using integer programming James Dept of Computer Science & York Centre for Complex Systems Analysis University of York, York, YO10 5DD, UK jc@cs.york.ac.uk Abstract
Maximum likelihood pedigree reconstruction using integer programming James Dept of Computer Science & York Centre for Complex Systems Analysis University of York, York, YO10 5DD, UK jc@cs.york.ac.uk Abstract
Lecture 1: Introduction to pedigree analysis
 Lecture 1: Introduction to pedigree analysis Magnus Dehli Vigeland NORBIS course, 8 th 12 th of January 2018, Oslo Outline Part I: Brief introductions Pedigrees symbols and terminology Some common relationships
Lecture 1: Introduction to pedigree analysis Magnus Dehli Vigeland NORBIS course, 8 th 12 th of January 2018, Oslo Outline Part I: Brief introductions Pedigrees symbols and terminology Some common relationships
Comparative method, coalescents, and the future. Correlation of states in a discrete-state model
 Comparative method, coalescents, and the future Joe Felsenstein Depts. of Genome Sciences and of Biology, University of Washington Comparative method, coalescents, and the future p.1/28 Correlation of
Comparative method, coalescents, and the future Joe Felsenstein Depts. of Genome Sciences and of Biology, University of Washington Comparative method, coalescents, and the future p.1/28 Correlation of
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1122655/dc1 Supporting Online Material for Finding Criminals Through DNA of Their Relatives Frederick R. Bieber,* Charles H. Brenner, David Lazer *Author for correspondence.
www.sciencemag.org/cgi/content/full/1122655/dc1 Supporting Online Material for Finding Criminals Through DNA of Their Relatives Frederick R. Bieber,* Charles H. Brenner, David Lazer *Author for correspondence.
Ancestral Recombination Graphs
 Ancestral Recombination Graphs Ancestral relationships among a sample of recombining sequences usually cannot be accurately described by just a single genealogy. Linked sites will have similar, but not
Ancestral Recombination Graphs Ancestral relationships among a sample of recombining sequences usually cannot be accurately described by just a single genealogy. Linked sites will have similar, but not
Gene coancestry in pedigrees and populations
 Gene coancestry in pedigrees and populations Thompson, Elizabeth University of Washington, Department of Statistics Box 354322 Seattle, WA 98115-4322, USA E-mail: eathomp@uw.edu Glazner, Chris University
Gene coancestry in pedigrees and populations Thompson, Elizabeth University of Washington, Department of Statistics Box 354322 Seattle, WA 98115-4322, USA E-mail: eathomp@uw.edu Glazner, Chris University
Pedigree reconstruction from SNP data: parentage assignment, sibship clustering and beyond
 Molecular Ecology Resources (2017) 17, 1009 1024 doi: 10.1111/1755-0998.12665 Pedigree reconstruction from SNP data: parentage assignment, sibship clustering and beyond JISCA HUISMAN Ashworth Laboratories,
Molecular Ecology Resources (2017) 17, 1009 1024 doi: 10.1111/1755-0998.12665 Pedigree reconstruction from SNP data: parentage assignment, sibship clustering and beyond JISCA HUISMAN Ashworth Laboratories,
University of Washington, TOPMed DCC July 2018
 Module 12: Comput l Pipeline for WGS Relatedness Inference from Genetic Data Timothy Thornton (tathornt@uw.edu) & Stephanie Gogarten (sdmorris@uw.edu) University of Washington, TOPMed DCC July 2018 1 /
Module 12: Comput l Pipeline for WGS Relatedness Inference from Genetic Data Timothy Thornton (tathornt@uw.edu) & Stephanie Gogarten (sdmorris@uw.edu) University of Washington, TOPMed DCC July 2018 1 /
AFDAA 2012 WINTER MEETING Population Statistics Refresher Course - Lecture 3: Statistics of Kinship Analysis
 AFDAA 2012 WINTER MEETING Population Statistics Refresher Course - Lecture 3: Statistics of Kinship Analysis Ranajit Chakraborty, PhD Center for Computational Genomics Institute of Applied Genetics Department
AFDAA 2012 WINTER MEETING Population Statistics Refresher Course - Lecture 3: Statistics of Kinship Analysis Ranajit Chakraborty, PhD Center for Computational Genomics Institute of Applied Genetics Department
Algorithms for Genetics: Basics of Wright Fisher Model and Coalescent Theory
 Algorithms for Genetics: Basics of Wright Fisher Model and Coalescent Theory Vineet Bafna Harish Nagarajan and Nitin Udpa 1 Disclaimer Please note that a lot of the text and figures here are copied from
Algorithms for Genetics: Basics of Wright Fisher Model and Coalescent Theory Vineet Bafna Harish Nagarajan and Nitin Udpa 1 Disclaimer Please note that a lot of the text and figures here are copied from
Population Genetics using Trees. Peter Beerli Genome Sciences University of Washington Seattle WA
 Population Genetics using Trees Peter Beerli Genome Sciences University of Washington Seattle WA Outline 1. Introduction to the basic coalescent Population models The coalescent Likelihood estimation of
Population Genetics using Trees Peter Beerli Genome Sciences University of Washington Seattle WA Outline 1. Introduction to the basic coalescent Population models The coalescent Likelihood estimation of
1) Using the sightings data, determine who moved from one area to another and fill this data in on the data sheet.
 Parentage and Geography 5. The Life of Lulu the Lioness: A Heroine s Story Name: Objective Using genotypes from many individuals, determine maternity, paternity, and relatedness among a group of lions.
Parentage and Geography 5. The Life of Lulu the Lioness: A Heroine s Story Name: Objective Using genotypes from many individuals, determine maternity, paternity, and relatedness among a group of lions.
Bayesian parentage analysis with systematic accountability of genotyping error, missing data, and false matching
 Genetics and population analysis Bayesian parentage analysis with systematic accountability of genotyping error, missing data, and false matching Mark R. Christie 1,*, Jacob A. Tennessen 1 and Michael
Genetics and population analysis Bayesian parentage analysis with systematic accountability of genotyping error, missing data, and false matching Mark R. Christie 1,*, Jacob A. Tennessen 1 and Michael
Pedigree Reconstruction Using Identity by Descent
 Pedigree Reconstruction Using Identity by Descent Bonnie Kirkpatrick 1, Shuai Cheng Li 2, Richard M. Karp 3, and Eran Halperin 4 1 Electrical Engineering and Computer Sciences, University of California,
Pedigree Reconstruction Using Identity by Descent Bonnie Kirkpatrick 1, Shuai Cheng Li 2, Richard M. Karp 3, and Eran Halperin 4 1 Electrical Engineering and Computer Sciences, University of California,
ville, VA Associate Editor: XXXXXXX Received on XXXXX; revised on XXXXX; accepted on XXXXX
 Robust Relationship Inference in Genome Wide Association Studies Ani Manichaikul 1,2, Josyf Mychaleckyj 1, Stephen S. Rich 1, Kathy Daly 3, Michele Sale 1,4,5 and Wei- Min Chen 1,2,* 1 Center for Public
Robust Relationship Inference in Genome Wide Association Studies Ani Manichaikul 1,2, Josyf Mychaleckyj 1, Stephen S. Rich 1, Kathy Daly 3, Michele Sale 1,4,5 and Wei- Min Chen 1,2,* 1 Center for Public
A Review on Genetic Algorithm and Its Applications
 2017 IJSRST Volume 3 Issue 8 Print ISSN: 2395-6011 Online ISSN: 2395-602X Themed Section: Science and Technology A Review on Genetic Algorithm and Its Applications Anju Bala Research Scholar, Department
2017 IJSRST Volume 3 Issue 8 Print ISSN: 2395-6011 Online ISSN: 2395-602X Themed Section: Science and Technology A Review on Genetic Algorithm and Its Applications Anju Bala Research Scholar, Department
Parsimony II Search Algorithms
 Parsimony II Search Algorithms Genome 373 Genomic Informatics Elhanan Borenstein Raw distance correction As two DNA sequences diverge, it is easy to see that their maximum raw distance is ~0.75 (assuming
Parsimony II Search Algorithms Genome 373 Genomic Informatics Elhanan Borenstein Raw distance correction As two DNA sequences diverge, it is easy to see that their maximum raw distance is ~0.75 (assuming
Forensic use of the genomic relationship matrix to validate and discover livestock. pedigrees
 Forensic use of the genomic relationship matrix to validate and discover livestock pedigrees K. L. Moore*, C. Vilela*, K. Kaseja*, R, Mrode* and M. Coffey* * Scotland s Rural College (SRUC), Easter Bush,
Forensic use of the genomic relationship matrix to validate and discover livestock pedigrees K. L. Moore*, C. Vilela*, K. Kaseja*, R, Mrode* and M. Coffey* * Scotland s Rural College (SRUC), Easter Bush,
Popstats Parentage Statistics Strength of Genetic Evidence In Parentage Testing
 Popstats Parentage Statistics Strength of Genetic Evidence In Parentage Testing Arthur J. Eisenberg, Ph.D. Director DNA Identity Laboratory UNT-Health Science Center eisenber@hsc.unt.edu PATERNITY TESTING
Popstats Parentage Statistics Strength of Genetic Evidence In Parentage Testing Arthur J. Eisenberg, Ph.D. Director DNA Identity Laboratory UNT-Health Science Center eisenber@hsc.unt.edu PATERNITY TESTING
4. Kinship Paper Challenge
 4. António Amorim (aamorim@ipatimup.pt) Nádia Pinto (npinto@ipatimup.pt) 4.1 Approach After a woman dies her child claims for a paternity test of the man who is supposed to be his father. The test is carried
4. António Amorim (aamorim@ipatimup.pt) Nádia Pinto (npinto@ipatimup.pt) 4.1 Approach After a woman dies her child claims for a paternity test of the man who is supposed to be his father. The test is carried
BIOL Evolution. Lecture 8
 BIOL 432 - Evolution Lecture 8 Expected Genotype Frequencies in the Absence of Evolution are Determined by the Hardy-Weinberg Equation. Assumptions: 1) No mutation 2) Random mating 3) Infinite population
BIOL 432 - Evolution Lecture 8 Expected Genotype Frequencies in the Absence of Evolution are Determined by the Hardy-Weinberg Equation. Assumptions: 1) No mutation 2) Random mating 3) Infinite population
Objective: Why? 4/6/2014. Outlines:
 Objective: Develop mathematical models that quantify/model resemblance between relatives for phenotypes of a quantitative trait : - based on pedigree - based on markers Outlines: Causal model for covariances
Objective: Develop mathematical models that quantify/model resemblance between relatives for phenotypes of a quantitative trait : - based on pedigree - based on markers Outlines: Causal model for covariances
NON-RANDOM MATING AND INBREEDING
 Instructor: Dr. Martha B. Reiskind AEC 495/AEC592: Conservation Genetics DEFINITIONS Nonrandom mating: Mating individuals are more closely related or less closely related than those drawn by chance from
Instructor: Dr. Martha B. Reiskind AEC 495/AEC592: Conservation Genetics DEFINITIONS Nonrandom mating: Mating individuals are more closely related or less closely related than those drawn by chance from
DNA: Statistical Guidelines
 Frequency calculations for STR analysis When a probative association between an evidence profile and a reference profile is made, a frequency estimate is calculated to give weight to the association. Frequency
Frequency calculations for STR analysis When a probative association between an evidence profile and a reference profile is made, a frequency estimate is calculated to give weight to the association. Frequency
TDT vignette Use of snpstats in family based studies
 TDT vignette Use of snpstats in family based studies David Clayton April 30, 2018 Pedigree data The snpstats package contains some tools for analysis of family-based studies. These assume that a subject
TDT vignette Use of snpstats in family based studies David Clayton April 30, 2018 Pedigree data The snpstats package contains some tools for analysis of family-based studies. These assume that a subject
Some of these slides have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks!
 Some of these slides have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks! Paul has many great tools for teaching phylogenetics at his web site: http://hydrodictyon.eeb.uconn.edu/people/plewis
Some of these slides have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks! Paul has many great tools for teaching phylogenetics at his web site: http://hydrodictyon.eeb.uconn.edu/people/plewis
Genetic Analysis for Spring- and Fall- Run San Joaquin River Chinook Salmon for the San Joaquin River Restoration Program
 Study 49 Genetic Analysis for Spring- and Fall- Run San Joaquin River Chinook Salmon for the San Joaquin River Restoration Program Final 2015 Monitoring and Analysis Plan January 2015 Statement of Work
Study 49 Genetic Analysis for Spring- and Fall- Run San Joaquin River Chinook Salmon for the San Joaquin River Restoration Program Final 2015 Monitoring and Analysis Plan January 2015 Statement of Work
Chromosome X haplotyping in deficiency paternity testing principles and case report
 International Congress Series 1239 (2003) 815 820 Chromosome X haplotyping in deficiency paternity testing principles and case report R. Szibor a, *, I. Plate a, J. Edelmann b, S. Hering c, E. Kuhlisch
International Congress Series 1239 (2003) 815 820 Chromosome X haplotyping in deficiency paternity testing principles and case report R. Szibor a, *, I. Plate a, J. Edelmann b, S. Hering c, E. Kuhlisch
Inbreeding and self-fertilization
 Inbreeding and self-fertilization Introduction Remember that long list of assumptions associated with derivation of the Hardy-Weinberg principle that I went over a couple of lectures ago? Well, we re about
Inbreeding and self-fertilization Introduction Remember that long list of assumptions associated with derivation of the Hardy-Weinberg principle that I went over a couple of lectures ago? Well, we re about
Exercise 4 Exploring Population Change without Selection
 Exercise 4 Exploring Population Change without Selection This experiment began with nine Avidian ancestors of identical fitness; the mutation rate is zero percent. Since descendants can never differ in
Exercise 4 Exploring Population Change without Selection This experiment began with nine Avidian ancestors of identical fitness; the mutation rate is zero percent. Since descendants can never differ in
Developing Conclusions About Different Modes of Inheritance
 Pedigree Analysis Introduction A pedigree is a diagram of family relationships that uses symbols to represent people and lines to represent genetic relationships. These diagrams make it easier to visualize
Pedigree Analysis Introduction A pedigree is a diagram of family relationships that uses symbols to represent people and lines to represent genetic relationships. These diagrams make it easier to visualize
Primer on Human Pedigree Analysis:
 Primer on Human Pedigree Analysis: Criteria for the selection and collection of appropriate Family Reference Samples John V. Planz. Ph.D. UNT Center for Human Identification Successful Missing Person ID
Primer on Human Pedigree Analysis: Criteria for the selection and collection of appropriate Family Reference Samples John V. Planz. Ph.D. UNT Center for Human Identification Successful Missing Person ID
Inbreeding and self-fertilization
 Inbreeding and self-fertilization Introduction Remember that long list of assumptions associated with derivation of the Hardy-Weinberg principle that we just finished? Well, we re about to begin violating
Inbreeding and self-fertilization Introduction Remember that long list of assumptions associated with derivation of the Hardy-Weinberg principle that we just finished? Well, we re about to begin violating
Analysis of geographically structured populations: Estimators based on coalescence
 Analysis of geographically structured populations: Estimators based on coalescence Peter Beerli Department of Genetics, Box 357360, University of Washington, Seattle WA 9895-7360, Email: beerli@genetics.washington.edu
Analysis of geographically structured populations: Estimators based on coalescence Peter Beerli Department of Genetics, Box 357360, University of Washington, Seattle WA 9895-7360, Email: beerli@genetics.washington.edu
Detecting inbreeding depression is difficult in captive endangered species
 Animal Conservation (1999) 2, 131 136 1999 The Zoological Society of London Printed in the United Kingdom Detecting inbreeding depression is difficult in captive endangered species Steven T. Kalinowski
Animal Conservation (1999) 2, 131 136 1999 The Zoological Society of London Printed in the United Kingdom Detecting inbreeding depression is difficult in captive endangered species Steven T. Kalinowski
An Optimal Algorithm for Automatic Genotype Elimination
 Am. J. Hum. Genet. 65:1733 1740, 1999 An Optimal Algorithm for Automatic Genotype Elimination Jeffrey R. O Connell 1,2 and Daniel E. Weeks 1 1 Department of Human Genetics, University of Pittsburgh, Pittsburgh,
Am. J. Hum. Genet. 65:1733 1740, 1999 An Optimal Algorithm for Automatic Genotype Elimination Jeffrey R. O Connell 1,2 and Daniel E. Weeks 1 1 Department of Human Genetics, University of Pittsburgh, Pittsburgh,
Determining Relatedness from a Pedigree Diagram
 Kin structure & relatedness Francis L. W. Ratnieks Aims & Objectives Aims 1. To show how to determine regression relatedness among individuals using a pedigree diagram. Social Insects: C1139 2. To show
Kin structure & relatedness Francis L. W. Ratnieks Aims & Objectives Aims 1. To show how to determine regression relatedness among individuals using a pedigree diagram. Social Insects: C1139 2. To show
CONGEN. Inbreeding vocabulary
 CONGEN Inbreeding vocabulary Inbreeding Mating between relatives. Inbreeding depression Reduction in fitness due to inbreeding. Identical by descent Alleles that are identical by descent are direct descendents
CONGEN Inbreeding vocabulary Inbreeding Mating between relatives. Inbreeding depression Reduction in fitness due to inbreeding. Identical by descent Alleles that are identical by descent are direct descendents
Introduction to Autosomal DNA Tools
 GENETIC GENEALOGY JOURNEY Debbie Parker Wayne, CG, CGL Introduction to Autosomal DNA Tools Just as in the old joke about a new genealogist walking into the library and asking for the book that covers my
GENETIC GENEALOGY JOURNEY Debbie Parker Wayne, CG, CGL Introduction to Autosomal DNA Tools Just as in the old joke about a new genealogist walking into the library and asking for the book that covers my
Populations. Arindam RoyChoudhury. Department of Biostatistics, Columbia University, New York NY 10032, U.S.A.,
 Change in Recessive Lethal Alleles Frequency in Inbred Populations arxiv:1304.2955v1 [q-bio.pe] 10 Apr 2013 Arindam RoyChoudhury Department of Biostatistics, Columbia University, New York NY 10032, U.S.A.,
Change in Recessive Lethal Alleles Frequency in Inbred Populations arxiv:1304.2955v1 [q-bio.pe] 10 Apr 2013 Arindam RoyChoudhury Department of Biostatistics, Columbia University, New York NY 10032, U.S.A.,
EUROPEAN COMMISSION Research Executive Agency Marie Curie Actions International Fellowships
 EUROPEAN COMMISSION Research Executive Agency Marie Curie Actions International Fellowships Project No: 300077 Project Acronym: RAPIDEVO Project Full Name: Rapid evolutionary responses to climate change
EUROPEAN COMMISSION Research Executive Agency Marie Curie Actions International Fellowships Project No: 300077 Project Acronym: RAPIDEVO Project Full Name: Rapid evolutionary responses to climate change
ADJUSTING POPULATION ESTIMATES FOR GENOTYPING ERROR IN NON- INVASIVE DNA-BASED MARK-RECAPTURE EXPERIMENTS
 Libraries 2007-19th Annual Conference Proceedings ADJUSTING POPULATION ESTIMATES FOR GENOTYPING ERROR IN NON- INVASIVE DNA-BASED MARK-RECAPTURE EXPERIMENTS Shannon M. Knapp Bruce A. Craig Follow this and
Libraries 2007-19th Annual Conference Proceedings ADJUSTING POPULATION ESTIMATES FOR GENOTYPING ERROR IN NON- INVASIVE DNA-BASED MARK-RECAPTURE EXPERIMENTS Shannon M. Knapp Bruce A. Craig Follow this and
2. Survey Methodology
 Analysis of Butterfly Survey Data and Methodology from San Bruno Mountain Habitat Conservation Plan (1982 2000). 2. Survey Methodology Travis Longcore University of Southern California GIS Research Laboratory
Analysis of Butterfly Survey Data and Methodology from San Bruno Mountain Habitat Conservation Plan (1982 2000). 2. Survey Methodology Travis Longcore University of Southern California GIS Research Laboratory
Genealogical trees, coalescent theory, and the analysis of genetic polymorphisms
 Genealogical trees, coalescent theory, and the analysis of genetic polymorphisms Magnus Nordborg University of Southern California The importance of history Genetic polymorphism data represent the outcome
Genealogical trees, coalescent theory, and the analysis of genetic polymorphisms Magnus Nordborg University of Southern California The importance of history Genetic polymorphism data represent the outcome
Two-point linkage analysis using the LINKAGE/FASTLINK programs
 1 Two-point linkage analysis using the LINKAGE/FASTLINK programs Copyrighted 2018 Maria Chahrour and Suzanne M. Leal These exercises will introduce the LINKAGE file format which is the standard format
1 Two-point linkage analysis using the LINKAGE/FASTLINK programs Copyrighted 2018 Maria Chahrour and Suzanne M. Leal These exercises will introduce the LINKAGE file format which is the standard format
Shuffled Complex Evolution
 Shuffled Complex Evolution Shuffled Complex Evolution An Evolutionary algorithm That performs local and global search A solution evolves locally through a memetic evolution (Local search) This local search
Shuffled Complex Evolution Shuffled Complex Evolution An Evolutionary algorithm That performs local and global search A solution evolves locally through a memetic evolution (Local search) This local search
Linkage Analysis in Merlin. Meike Bartels Kate Morley Danielle Posthuma
 Linkage Analysis in Merlin Meike Bartels Kate Morley Danielle Posthuma Software for linkage analyses Genehunter Mendel Vitesse Allegro Simwalk Loki Merlin. Mx R Lisrel MERLIN software Programs: MERLIN
Linkage Analysis in Merlin Meike Bartels Kate Morley Danielle Posthuma Software for linkage analyses Genehunter Mendel Vitesse Allegro Simwalk Loki Merlin. Mx R Lisrel MERLIN software Programs: MERLIN
Solving Assembly Line Balancing Problem using Genetic Algorithm with Heuristics- Treated Initial Population
 Solving Assembly Line Balancing Problem using Genetic Algorithm with Heuristics- Treated Initial Population 1 Kuan Eng Chong, Mohamed K. Omar, and Nooh Abu Bakar Abstract Although genetic algorithm (GA)
Solving Assembly Line Balancing Problem using Genetic Algorithm with Heuristics- Treated Initial Population 1 Kuan Eng Chong, Mohamed K. Omar, and Nooh Abu Bakar Abstract Although genetic algorithm (GA)
Forward thinking: the predictive approach
 Coalescent Theory 1 Forward thinking: the predictive approach Random variation in reproduction causes random fluctuation in allele frequencies. Can describe this process as diffusion: (Wright 1931) showed
Coalescent Theory 1 Forward thinking: the predictive approach Random variation in reproduction causes random fluctuation in allele frequencies. Can describe this process as diffusion: (Wright 1931) showed
SNP variant discovery in pedigrees using Bayesian networks. Amit R. Indap
 SNP variant discovery in pedigrees using Bayesian networks Amit R. Indap 1 1 Background Next generation sequencing technologies have reduced the cost and increased the throughput of DNA sequencing experiments
SNP variant discovery in pedigrees using Bayesian networks Amit R. Indap 1 1 Background Next generation sequencing technologies have reduced the cost and increased the throughput of DNA sequencing experiments
The number of mates of latin squares of sizes 7 and 8
 The number of mates of latin squares of sizes 7 and 8 Megan Bryant James Figler Roger Garcia Carl Mummert Yudishthisir Singh Working draft not for distribution December 17, 2012 Abstract We study the number
The number of mates of latin squares of sizes 7 and 8 Megan Bryant James Figler Roger Garcia Carl Mummert Yudishthisir Singh Working draft not for distribution December 17, 2012 Abstract We study the number
[CLIENT] SmithDNA1701 DE January 2017
![[CLIENT] SmithDNA1701 DE January 2017 [CLIENT] SmithDNA1701 DE January 2017](/thumbs/93/113058427.jpg) [CLIENT] SmithDNA1701 DE1704205 11 January 2017 DNA Discovery Plan GOAL Create a research plan to determine how the client s DNA results relate to his family tree as currently constructed. The client s
[CLIENT] SmithDNA1701 DE1704205 11 January 2017 DNA Discovery Plan GOAL Create a research plan to determine how the client s DNA results relate to his family tree as currently constructed. The client s
Reconstruction of pedigrees in clonal plant populations
 Reconstruction of pedigrees in clonal plant populations Markus Riester,a, Peter F. Stadler a,b,c,d,e, Konstantin Klemm a a Bioinformatics Group, Department of Computer Science, and Interdisciplinary Center
Reconstruction of pedigrees in clonal plant populations Markus Riester,a, Peter F. Stadler a,b,c,d,e, Konstantin Klemm a a Bioinformatics Group, Department of Computer Science, and Interdisciplinary Center
The Genetic Algorithm
 The Genetic Algorithm The Genetic Algorithm, (GA) is finding increasing applications in electromagnetics including antenna design. In this lesson we will learn about some of these techniques so you are
The Genetic Algorithm The Genetic Algorithm, (GA) is finding increasing applications in electromagnetics including antenna design. In this lesson we will learn about some of these techniques so you are
Recent effective population size estimated from segments of identity by descent in the Lithuanian population
 Anthropological Science Advance Publication Recent effective population size estimated from segments of identity by descent in the Lithuanian population Alina Urnikytė 1 *, Alma Molytė 1, Vaidutis Kučinskas
Anthropological Science Advance Publication Recent effective population size estimated from segments of identity by descent in the Lithuanian population Alina Urnikytė 1 *, Alma Molytė 1, Vaidutis Kučinskas
Assessment of alternative genotyping strategies to maximize imputation accuracy at minimal cost
 Huang et al. Genetics Selection Evolution 2012, 44:25 Genetics Selection Evolution RESEARCH Open Access Assessment of alternative genotyping strategies to maximize imputation accuracy at minimal cost Yijian
Huang et al. Genetics Selection Evolution 2012, 44:25 Genetics Selection Evolution RESEARCH Open Access Assessment of alternative genotyping strategies to maximize imputation accuracy at minimal cost Yijian
Population Adaptation for Genetic Algorithm-based Cognitive Radios
 Population Adaptation for Genetic Algorithm-based Cognitive Radios Timothy R. Newman, Rakesh Rajbanshi, Alexander M. Wyglinski, Joseph B. Evans, and Gary J. Minden Information Technology and Telecommunications
Population Adaptation for Genetic Algorithm-based Cognitive Radios Timothy R. Newman, Rakesh Rajbanshi, Alexander M. Wyglinski, Joseph B. Evans, and Gary J. Minden Information Technology and Telecommunications
PEDIGREE ANALYSIS OF FLORIDA MANGO CULTIVARS
 Proc. Fla. State Hort. Soc. 118:192-197. 2005. PEDIGREE ANALYSIS OF FLORIDA MANGO CULTIVARS CECILE T. OLANO, 1 RAYMOND J. SCHNELL, 1 * WILBER E. QUINTANILLA 1 AND RICHARD J. CAMPBELL 2 1 National Germplasm
Proc. Fla. State Hort. Soc. 118:192-197. 2005. PEDIGREE ANALYSIS OF FLORIDA MANGO CULTIVARS CECILE T. OLANO, 1 RAYMOND J. SCHNELL, 1 * WILBER E. QUINTANILLA 1 AND RICHARD J. CAMPBELL 2 1 National Germplasm
Viral epidemiology and the Coalescent
 Viral epidemiology and the Coalescent Philippe Lemey and Marc A. Suchard Department of Microbiology and Immunology K.U. Leuven, and Departments of Biomathematics and Human Genetics David Geffen School
Viral epidemiology and the Coalescent Philippe Lemey and Marc A. Suchard Department of Microbiology and Immunology K.U. Leuven, and Departments of Biomathematics and Human Genetics David Geffen School
Evaluating genetic traceability methods for captive bred marine fish and their applications in fisheries management and wildlife forensics
 The following supplements accompany the article Evaluating genetic traceability methods for captive bred marine fish and their applications in fisheries management and wildlife forensics Jonas Bylemans*,
The following supplements accompany the article Evaluating genetic traceability methods for captive bred marine fish and their applications in fisheries management and wildlife forensics Jonas Bylemans*,
BI515 - Population Genetics
 BI515 - Population Genetics Fall 2014 Michael Sorenson msoren@bu.edu Office hours (BRB529): M, Th, F 4-5PM or by appt. (send e-mail) My research: Avian behavior, systematics, population genetics, and molecular
BI515 - Population Genetics Fall 2014 Michael Sorenson msoren@bu.edu Office hours (BRB529): M, Th, F 4-5PM or by appt. (send e-mail) My research: Avian behavior, systematics, population genetics, and molecular
Sea Duck Joint Venture Annual Project Summary for Endorsed Projects FY08 (October 1, 2007 to September 30, 2008)
 Sea Duck Joint Venture Annual Project Summary for Endorsed Projects FY08 (October 1, 2007 to September 30, 2008) Project Title: SDJV#16, Ducks Unlimited Canada s Common Eider Initiative (year five of a
Sea Duck Joint Venture Annual Project Summary for Endorsed Projects FY08 (October 1, 2007 to September 30, 2008) Project Title: SDJV#16, Ducks Unlimited Canada s Common Eider Initiative (year five of a
Puzzling Pedigrees. Essential Question: How can pedigrees be used to study the inheritance of human traits?
 Name: Puzzling Pedigrees Essential Question: How can pedigrees be used to study the inheritance of human traits? Studying inheritance in humans is more difficult than studying inheritance in fruit flies
Name: Puzzling Pedigrees Essential Question: How can pedigrees be used to study the inheritance of human traits? Studying inheritance in humans is more difficult than studying inheritance in fruit flies
Package pedantics. R topics documented: April 18, Type Package
 Type Package Package pedantics April 18, 2018 Title Functions to Facilitate Power and Sensitivity Analyses for Genetic Studies of Natural Populations Version 1.7 Date 2018-04-18 Depends R (>= 2.4.0), MasterBayes,
Type Package Package pedantics April 18, 2018 Title Functions to Facilitate Power and Sensitivity Analyses for Genetic Studies of Natural Populations Version 1.7 Date 2018-04-18 Depends R (>= 2.4.0), MasterBayes,
BIOL 502 Population Genetics Spring 2017
 BIOL 502 Population Genetics Spring 2017 Week 8 Inbreeding Arun Sethuraman California State University San Marcos Table of contents 1. Inbreeding Coefficient 2. Mating Systems 3. Consanguinity and Inbreeding
BIOL 502 Population Genetics Spring 2017 Week 8 Inbreeding Arun Sethuraman California State University San Marcos Table of contents 1. Inbreeding Coefficient 2. Mating Systems 3. Consanguinity and Inbreeding
Pedigrees How do scientists trace hereditary diseases through a family history?
 Why? Pedigrees How do scientists trace hereditary diseases through a family history? Imagine you want to learn about an inherited genetic trait present in your family. How would you find out the chances
Why? Pedigrees How do scientists trace hereditary diseases through a family history? Imagine you want to learn about an inherited genetic trait present in your family. How would you find out the chances
ISI Web of Knowledge Page 1 (Articles ) [ 1 ]
![ISI Web of Knowledge Page 1 (Articles ) [ 1 ] ISI Web of Knowledge Page 1 (Articles ) [ 1 ]](/thumbs/72/67568562.jpg) ISI Web of Knowledge Page 1 (Articles 1 -- 32) [ 1 ] Record 1 of 32 Title Checklist of the birds of Aruba, Curacao and Bonaire, South Caribbean Author(s) Prins, TG; Reuter, JH; Debrot, AO; Wattel, J; Nijman,
ISI Web of Knowledge Page 1 (Articles 1 -- 32) [ 1 ] Record 1 of 32 Title Checklist of the birds of Aruba, Curacao and Bonaire, South Caribbean Author(s) Prins, TG; Reuter, JH; Debrot, AO; Wattel, J; Nijman,
Growing the Family Tree: The Power of DNA in Reconstructing Family Relationships
 Growing the Family Tree: The Power of DNA in Reconstructing Family Relationships Luke A. D. Hutchison Natalie M. Myres Scott R. Woodward Sorenson Molecular Genealogy Foundation (www.smgf.org) 2511 South
Growing the Family Tree: The Power of DNA in Reconstructing Family Relationships Luke A. D. Hutchison Natalie M. Myres Scott R. Woodward Sorenson Molecular Genealogy Foundation (www.smgf.org) 2511 South
Population Management User,s Manual
 Population Management 2000 User,s Manual PM2000 version 1.163 14 July 2002 Robert C. Lacy Chicago Zoological Society Jonathan D. Ballou National Zoological Park Smithsonian Institution Software developed
Population Management 2000 User,s Manual PM2000 version 1.163 14 July 2002 Robert C. Lacy Chicago Zoological Society Jonathan D. Ballou National Zoological Park Smithsonian Institution Software developed
Multiple Male Feeders at Nests of the Veery
 Multiple Male Feeders at Nests of the Veery Author(s): Matthew R. Halley and Christopher M. Heckscher Source: The Wilson Journal of Ornithology, 124(2):396-399. Published By: The Wilson Ornithological
Multiple Male Feeders at Nests of the Veery Author(s): Matthew R. Halley and Christopher M. Heckscher Source: The Wilson Journal of Ornithology, 124(2):396-399. Published By: The Wilson Ornithological
Report on Research Conducted in Partial Support by the Nuttal Ornithological Club November 2010
 Report on Research Conducted in Partial Support by the Nuttal Ornithological Club November 2010 Grant issued to: New England Institute for Landscape Ecology, 266 Prospect Hill Road, Canaan, New Hampshire
Report on Research Conducted in Partial Support by the Nuttal Ornithological Club November 2010 Grant issued to: New England Institute for Landscape Ecology, 266 Prospect Hill Road, Canaan, New Hampshire
Bottlenecks reduce genetic variation Genetic Drift
 Bottlenecks reduce genetic variation Genetic Drift Northern Elephant Seals were reduced to ~30 individuals in the 1800s. Rare alleles are likely to be lost during a bottleneck Two important determinants
Bottlenecks reduce genetic variation Genetic Drift Northern Elephant Seals were reduced to ~30 individuals in the 1800s. Rare alleles are likely to be lost during a bottleneck Two important determinants
arxiv: v1 [cs.ai] 13 Dec 2014
![arxiv: v1 [cs.ai] 13 Dec 2014 arxiv: v1 [cs.ai] 13 Dec 2014](/thumbs/96/129238166.jpg) Combinatorial Structure of the Deterministic Seriation Method with Multiple Subset Solutions Mark E. Madsen Department of Anthropology, Box 353100, University of Washington, Seattle WA, 98195 USA arxiv:1412.6060v1
Combinatorial Structure of the Deterministic Seriation Method with Multiple Subset Solutions Mark E. Madsen Department of Anthropology, Box 353100, University of Washington, Seattle WA, 98195 USA arxiv:1412.6060v1
Parentage analysis. Every person receives a unique set of genetic information from their parents - half from Mom and half from Dad
 Parentage analysis Similar techniques as those used in human parentage testing! With 99.99% probability, you ARE the father Every person receives a unique set of genetic information from their parents
Parentage analysis Similar techniques as those used in human parentage testing! With 99.99% probability, you ARE the father Every person receives a unique set of genetic information from their parents
Characterization of the global Brown Swiss cattle population structure
 Swedish University of Agricultural Sciences Faculty of Veterinary Medicine and Animal Science Characterization of the global Brown Swiss cattle population structure Worede Zinabu Gebremariam Examensarbete
Swedish University of Agricultural Sciences Faculty of Veterinary Medicine and Animal Science Characterization of the global Brown Swiss cattle population structure Worede Zinabu Gebremariam Examensarbete
MS.LS2.A: Interdependent Relationships in Ecosystems. MS.LS2.C: Ecosystem Dynamics, Functioning, and Resilience. MS.LS4.D: Biodiversity and Humans
 Disciplinary Core Idea MS.LS2.A: Interdependent Relationships in Ecosystems Similarly, predatory interactions may reduce the number of organisms or eliminate whole populations of organisms. Mutually beneficial
Disciplinary Core Idea MS.LS2.A: Interdependent Relationships in Ecosystems Similarly, predatory interactions may reduce the number of organisms or eliminate whole populations of organisms. Mutually beneficial
Behavioral Adaptations for Survival 1. Co-evolution of predator and prey ( evolutionary arms races )
 Behavioral Adaptations for Survival 1 Co-evolution of predator and prey ( evolutionary arms races ) Outline Mobbing Behavior What is an adaptation? The Comparative Method Divergent and convergent evolution
Behavioral Adaptations for Survival 1 Co-evolution of predator and prey ( evolutionary arms races ) Outline Mobbing Behavior What is an adaptation? The Comparative Method Divergent and convergent evolution
Genomic Variation of Inbreeding and Ancestry in the Remaining Two Isle Royale Wolves
 Journal of Heredity, 17, 1 16 doi:1.19/jhered/esw8 Original Article Advance Access publication December 1, 16 Original Article Genomic Variation of Inbreeding and Ancestry in the Remaining Two Isle Royale
Journal of Heredity, 17, 1 16 doi:1.19/jhered/esw8 Original Article Advance Access publication December 1, 16 Original Article Genomic Variation of Inbreeding and Ancestry in the Remaining Two Isle Royale
GENOMIC REARRANGEMENT ALGORITHMS
 GENOMIC REARRANGEMENT ALGORITHMS KAREN LOSTRITTO Abstract. In this paper, I discuss genomic rearrangement. Specifically, I describe the formal representation of these genomic rearrangements as well as
GENOMIC REARRANGEMENT ALGORITHMS KAREN LOSTRITTO Abstract. In this paper, I discuss genomic rearrangement. Specifically, I describe the formal representation of these genomic rearrangements as well as
Mehrdad Amirghasemi a* Reza Zamani a
 The roles of evolutionary computation, fitness landscape, constructive methods and local searches in the development of adaptive systems for infrastructure planning Mehrdad Amirghasemi a* Reza Zamani a
The roles of evolutionary computation, fitness landscape, constructive methods and local searches in the development of adaptive systems for infrastructure planning Mehrdad Amirghasemi a* Reza Zamani a
Non-Paternity: Implications and Resolution
 Non-Paternity: Implications and Resolution Michelle Beckwith PTC Labs 2006 AABB HITA Meeting October 8, 2006 Considerations when identifying victims using relatives Identification requires knowledge of
Non-Paternity: Implications and Resolution Michelle Beckwith PTC Labs 2006 AABB HITA Meeting October 8, 2006 Considerations when identifying victims using relatives Identification requires knowledge of
State of the Estuary Report 2015
 1 State of the Estuary Report 2015 Summary PROCESSES Feeding Chicks, Brandt s Cormorant Prepared by Nadav Nur Point Blue Conservation Science State of the Estuary 2015: Processes Brandt s Cormorant Reproductive
1 State of the Estuary Report 2015 Summary PROCESSES Feeding Chicks, Brandt s Cormorant Prepared by Nadav Nur Point Blue Conservation Science State of the Estuary 2015: Processes Brandt s Cormorant Reproductive
