The permax Package. May 26, 2004
|
|
|
- Violet Burke
- 5 years ago
- Views:
Transcription
1 The permax Package May 26, 2004 Version Author Robert J. Gray Maintainer Robert Gentleman The permax library consists of 7 functions, intended to facilitate certain basic analyses of DNA array data, especially with regard to comparing expression levels between two types of tissue. Title permax Depends R (>= 1.2) License GPL 2 R topics documented: permax permcor permsep plot.expr plot.permax rowperm summary.permax Index 11 permax 2-sample permutation t-tests for high dimensional data For high dimensional vectors of observations, computes t statistics for each attribute, and assesses significance using the permutation distribution of the maximum and minimum over all attributes. permax(data, ig1, nperm=0, logs=true, ranks=false, min.np=1, ig2, WHseed=NULL) 1
2 2 permax data Data matrix or data frame. Each case is a column, and each row is an attribute (the opposite of the standard configuration). ig1 The columns of data corresponding to group 1 nperm logs ranks min.np ig2 WHseed Value The number of random permutations to use in computing the p-values. The default is to use the entire permutation distribution, which is only feasible if the sample sizes are fairly small If logs=true (the default), then logs of the values in data are used in the statistics. If ranks=t, then within row ranks are used in place of the values in data in the t statistics. This is equivalent to using the Wilcoxon statistic. Default is ranks=f data will be subset to only rows with at least min.np values larger than min(data) in the columns in ig1 and ig2 The columns of data corresponding to group 2. The default is to include all columns not in ig1 in group 2. When both ig1 and ig2 are given, columns not in either are excluded from the tests. Initial random number seed (a vector of 3 integers). If missing, an initial seed is generated from the runif() function. Not needed if all permutations are calculated. For DNA array data, this function is designed to identify the genes which best discriminate between two tissue types. 2-sample t statistics are computed for each gene using logs (default), raw values, or ranks. Upper and lower p-values (p.upper, p.lower) are computed by comparing each statistic to the permutation distribution of the maximum and minimum (largest negative) statistic over all genes. The pind component of the output gives the p-value for the permutation distribution of each individual gene. It is strongly recommended that different seeds be used for different runs, and ideally the final seed from one run, attr(output, seed.end ), would be used as the initial seed in the next run. Output is a data.frame of class permax, with columns stat: the standardized test statistics for each row pind: individual permutation p-values (2-sided) p2: 2-sided p-value using the distribution of the max overall rows p.lower: 1-sided p-value for lower levels in group 1 p.upper: 1-sided p-value for higher levels in group 1 nml: # permutations where this row was the most significant for p.lower nmr: # permutations where this row was the most sig for p.upper m1, m2: means of groups 1 and 2 (means of logs if logs=t) s1, s2: std deviations of groups 1 and 2 (of logs if logs=t) np1,np2: # values > min(data) in groups 1 and 2 mdiff: difference of means (if logs=t the difference of geometric means) mrat: ratio of means (if logs=t ratio of geometric means) Also, if nperm>0, then output includes attributes seed.start giving the initial random number seed, and seed.end giving the value of the seed at the end. These can be accessed with the attributes() and attr() functions. summary.permax, plot.permax, permcor, permsep.
3 permcor 3 #generate make believe data set.seed(1292) ngenes < m1 <- rnorm(ngenes,4,1) m2 <- rnorm(ngenes,4,1) exp1 <- cbind(matrix(exp(rnorm(ngenes*5,m1,1)),nrow=ngenes), matrix(exp(rnorm(ngenes*10,m2,1)),nrow=ngenes)) exp1[exp1<20] <- 20 sub <- exp1>20 & exp1<150 exp1[sub] <- ifelse(runif(length(sub[sub]))<.5,20,exp1[sub]) dimnames(exp1) <- list(paste('x',format(1:ngenes,justify='l'),sep=''), paste('sample',format(1:ncol(exp1),justify='l'),sep='')) dimnames(exp1) <- list(paste('x',1:ngenes,sep=''), paste('sample',1:ncol(exp1),sep='')) exp1 <- round(exp1) uu <- permax(exp1,1:5) summary(uu,nl=5,nr=5) # 5 most extreme in each direction permcor permutation tests for correlations in high dimensional data For high dimensional vectors of observations, computes the correlation coefficient for each attribute with a specified vector of values, and assesses significance using the permutation distribution of the maximum and minimum over all attributes. permcor(data, phen, nperm=1000, logs=true, ranks=false, min.np=1, WHseed=NULL) data phen nperm logs ranks min.np WHseed Data matrix or data frame. Each case is a column, and each row is an attribute (the opposite of the standard configuration). A vector of values (the ideal phenotype pattern). The correlations of each row of data with phen will be computed. The number of random permutations to use in computing the p-values. The default is If nperm is < 0, the entire permutation distribution will be used, which is only feasible if the sample size is fairly small If logs=true (the default), then logs of the values in data are used in computing correlations (the actual values of phen are used, though). If ranks=t, then within row ranks are used in place of the values in data in the correlations. The actual values of phen are still used. Default is ranks=false. data will be subset to only rows with at least min.np values larger than min(data). Initial random number seed (a vector of 3 integers). If missing, an initial seed is generated from the runif() function. Not needed if all permutations are calculated.
4 4 permcor Value For DNA array data, this function is designed to identify the genes with the largest positive and negative correlations with the phenotype in phen. Upper and lower p-values (p.upper, p.lower) are computed by comparing each correlation to the permutation distribution of the maximum and minimum (largest negative) correlations over all genes. The pind component of the output gives the p-value for the permutation distribution of each individual gene. If phen is a vector of 1 s for the columns in group 1 and 0 s for the other columns, then the p-values from permcor() should be the same as from permax() (to within simulation precision if random permutations are used). permax() is substantially more efficient in this setting. The functions summary.permax() and plot.permax() can be used with the output of permcor(). It is strongly recommended that different seeds be used for different runs, and ideally the final seed from one run, attr(output, seed.end ), would be used as the initial seed in the next run. Output is a data.frame of class c( permcor, permax ), with columns stat: the Pearson correlation coeffcients for each row of data pind: individual permutation p-values (2-sided) p2: 2-sided p-value using the distribution of the max overall rows p.lower: 1-sided p-value for lower levels in group 1 p.upper: 1-sided p-value for higher levels in group 1 nml: # permutations where this row was the most significant for p.lower nmr: # permutations where this row was the most sig for p.upper np: # values > min(data) in each row Also, if nperm>0, then output includes attributes seed.start giving the initial random number seed, and seed.end giving the value of the seed at the end. These can be accessed with the attributes() and attr() functions. permax, summary.permax, plot.permax. set.seed(1292) ngenes < m1 <- rnorm(ngenes,4,1) m2 <- rnorm(ngenes,4,1) exp1 <- cbind(matrix(exp(rnorm(ngenes*5,m1,1)),nrow=ngenes), matrix(exp(rnorm(ngenes*10,m2,1)),nrow=ngenes)) exp1[exp1<20] <- 20 sub <- exp1>20 & exp1<150 exp1[sub] <- ifelse(runif(length(sub[sub]))<.5,20,exp1[sub]) dimnames(exp1) <- list(paste('x',format(1:ngenes,justify='l'),sep=''), paste('sample',format(1:ncol(exp1),justify='l'),sep='')) dimnames(exp1) <- list(paste('x',1:ngenes,sep=''), paste('sample',1:ncol(exp1),sep='')) exp1 <- round(exp1) #see the permax help file for the definition of exp1 u8 <- permcor(exp1,1:15) summary(u8,nr=4,nl=4) u10 <- permcor(exp1[,c(1:3,5:8)],c(1,1,1,0,0,0,0),nperm=0)
5 permsep 5 summary(u10,nl=4,nr=4) permsep Permutation analysis for complete separation Given two groups of samples of high dimensional attribute vectors (eg DNA array expression levels), determines the number of attributes which completely separate the two groups (all values in one group strictly larger than in the other), and a permutation p-value for this quantity. permsep(data, ig1, nperm=0, ig2, WHseed=NULL) data Data matrix or data frame. Each case is a column, and each row is an attribute (the opposite of the standard configuration). ig1 The columns of data corresponding to group 1 nperm ig2 WHseed The number of random permutations to use in computing the p-values. The default is to use the entire permutation distribution, which is only feasible if the sample sizes are fairly small The columns of data corresponding to group 2. The default is to include all columns not in ig1 in group 2. When both ig1 and ig2 are given, columns not in either are excluded. Initial random number seed (a vector of 3 integers). If missing, an initial seed is generated from the runif() function. Not needed if all permutations are calculated. Prints a vector giving the # genes with complete separation (all in one group larger than all in the other, the proportion of permutations with this many or more genes with complete separation (p-value) ( permutation actually means a distinct rearrangement of columns into 2 groups), the average number of genes per permutation with complete separation, and the proportion of permutations with any genes with complete separation. The value returned is a list with components ics = a vector indicating (with 1) which rows of data have complete separation, and dtcs = a vector containing the printed output. Also, if nperm>0, then the output includes attributes seed.start giving the initial random number seed, and seed.end giving the value of the seed at the end. These can be accessed with the attributes and attr functions.
6 6 plot.expr For each gene there will be 0, 1 or 2 rearrangements with complete separation, depending on the number of unique values and the sizes of the two groups. Adding these numbers over genes and dividing by the number of rearrangements gives the average number per permutation. The value returned averages only over the rearrangements actually used, though. It is strongly recommended that different seeds be used for different runs, and ideally the final seed from one run, attr(output, seed.end ), would be used as the initial seed in the next run. permax ngenes < m1 <- rnorm(ngenes,4,1) m2 <- rnorm(ngenes,4,1) exp1 <- cbind(matrix(exp(rnorm(ngenes*5,m1,1)),nrow=ngenes), matrix(exp(rnorm(ngenes*10,m2,1)),nrow=ngenes)) exp1[exp1<20] <- 20 sub <- exp1>20 & exp1<150 exp1[sub] <- ifelse(runif(length(sub[sub]))<.5,20,exp1[sub]) dimnames(exp1) <- list(paste('x',format(1:ngenes,justify='l'),sep=''), paste('sample',format(1:ncol(exp1),justify='l'),sep='')) dimnames(exp1) <- list(paste('x',1:ngenes,sep=''), paste('sample',1:ncol(exp1),sep='')) exp1 <- round(exp1) uuu <- permsep(exp1,1:5) plot.expr Color image plot of gene expression levels Represents values in the rows of a matrix as colored rectangles in an image plot plot.expr(x, logs=true, ig1=null, ig2=null, clmn.lab=dimnames(x)[[2]], row.lab=dimnames(x)[[1]], clmn.off=null, row.off=null,...) x logs ig1 matrix or data.frame containing the values to be plotted. If logs=true, then log values are used. The columns of x for cases in group 1 (see ) ig2 The columns in group 2. By default, all the columns not in group 1.
7 plot.expr 7 clmn.lab Labels for the columns in the array. row.lab Labels for the rows in the array. clmn.off Offset for printing the column labels (<0 to put labels outside the plot). row.off Offset for printing the row labels (<0 to put labels outside the plot).... Additional arguments to image and text (see par) none Values within a row are centered and normalized to have variance 1. If ig1 is not given, then the values are centered to have mean 0. If ig1 is given, the values are centered so the means of the columns in ig1 and ig2 are equal in magnitude and opposite in direction. The plot is thus useful for comparing within rows, but differences in colors between rows have no meaning. A graphics device supporting image plots must be initialized prior to calling this function. Under Splus 3.4 for unix, the following command (without the line breaks) initializes the X window motif plot window to use 30 colors from blue (lowest levels) to yellow (highest levels) for the image plots (in this scheme a value half way between the lowest and highest values would be a medium intensity gray). motif( -xrm sgraphmotif.colorschemes : background : black; lines : yellow cyan magenta green MediumBlue red; text : white yellow cyan magenta green MediumBlue red; images : blue 30 yellow ) Side Effects An image plot is created on the current graphics device plot.permax set.seed(1292) ngenes < m1 <- rnorm(ngenes,4,1) m2 <- rnorm(ngenes,4,1) exp1 <- cbind(matrix(exp(rnorm(ngenes*5,m1,1)),nrow=ngenes), matrix(exp(rnorm(ngenes*10,m2,1)),nrow=ngenes)) exp1[exp1<20] <- 20 sub <- exp1>20 & exp1<150 exp1[sub] <- ifelse(runif(length(sub[sub]))<.5,20,exp1[sub]) dimnames(exp1) <- list(paste('x',format(1:ngenes,justify='l'),sep=''), paste('sample',format(1:ncol(exp1),justify='l'),sep='')) dimnames(exp1) <- list(paste('x',1:ngenes,sep=''), paste('sample',1:ncol(exp1),sep='')) exp1 <- round(exp1) plot.expr(exp1[1:20,])
8 8 plot.permax plot.permax Image plot of the most significant genes (attributes) from a permax analysis Given the output of permax, and the array of expression levels, creates a color image plot of the expression levels of the most significant genes plot.permax(x, data, nl=25, nr=25, logs=true, ig1=null, ig2=null, clmn.lab=dimnames(data)[[2]], row.lab=dimnames(data)[[1]], clmn.off=null, row.off=null,...) x data nl nr logs ig1 A permax object (output from permax) Matrix or data frame of expression levels used as input to permax The nl most significant genes in the lower tail will be plotted The nr most significant genes in the upper tail will be plotted If logs=true, then log values are used. The columns of data for cases in group 1 (see ) ig2 The columns in group 2. By default, all the columns not in group 1. clmn.lab row.lab clmn.off row.off Labels for the columns in the array. Labels for the rows in the array. Offset for printing the column labels (<0 to put labels outside the plot). Offset for printing the row labels (<0 to put labels outside the plot).... Additional arguments to image and text (see par) none Values within a row of data are centered and normalized to have variance 1. If ig1 is not given, then the values are centered to have mean 0. If ig1 is given, the values are centered so the means of the columns in ig1 and ig2 are equal in magnitude and opposite in direction (usually ig1 and ig2 should match the values used in the permax call). The plot is thus useful for comparing within rows, but differences in colors between rows have no meaning. The plot will give the most significant lower tail genes in the top portion (most significant at the top), and the most significant upper tail genes in the bottom portion (most significant at the bottom). This function just selects out the appropriate rows of data, and calls plot.expr(). row.names(data) or dimnames(data)[[1]] must correspond to row.names(z) for the selection to work properly. A graphics device supporting image plots must be initialized prior to calling this function. Under Splus 3.4 for unix, the following command (without the line breaks) initializes the X window motif plot window to use 30 colors from blue (lowest levels) to yellow (highest levels) for the image plots (in this scheme a value half way between the lowest and highest values would be a medium intensity gray).
9 rowperm 9 motif( -xrm sgraphmotif.colorschemes : background : black; lines : yellow cyan magenta green MediumBlue red; text : white yellow cyan magenta green MediumBlue red; images : blue 30 yellow ) Side Effects An image plot is created on the current graphics device permax, plot.expr set.seed(1292) ngenes < m1 <- rnorm(ngenes,4,1) m2 <- rnorm(ngenes,4,1) exp1 <- cbind(matrix(exp(rnorm(ngenes*5,m1,1)),nrow=ngenes), matrix(exp(rnorm(ngenes*10,m2,1)),nrow=ngenes)) exp1[exp1<20] <- 20 sub <- exp1>20 & exp1<150 exp1[sub] <- ifelse(runif(length(sub[sub]))<.5,20,exp1[sub]) dimnames(exp1) <- list(paste('x',format(1:ngenes,justify='l'),sep=''), paste('sample',format(1:ncol(exp1),justify='l'),sep='')) dimnames(exp1) <- list(paste('x',1:ngenes,sep=''), paste('sample',1:ncol(exp1),sep='')) exp1 <- round(exp1) uu <- permax(exp1,1:5) plot(uu,exp1,ig1=1:5,cex=.7) rowperm Generates random permutations within rows of a matrix Given a matrix x, returns a copy of x with a separate random permutations applied within each row. rowperm(x) x a numeric matrix A numeric matrix with each row containing the elements from the corresponding row of x permuted in a random order.
10 10 summary.permax x <- matrix(1:12,3) x # [,1] [,2] [,3] [,4] #[1,] #[2,] #[3,] rowperm(x) # [,1] [,2] [,3] [,4] #[1,] #[2,] #[3,] summary.permax Summarizes the output of permax Finds and prints the most significant genes in the output of permax summary.permax(object, data, nl=25, nr=25,...) object data nl nr A dataframe of class permax (created by permax) The data matrix used as input to permax. If given, the rows of d corresponding to the most significant genes will also be printed. The nl most significant genes in the lower tail will be printed The nr most significant genes in the upper tail will be printed... Supplied for compatibility but not used. If d is given, it must be a data frame with row.names(d) corresponding to row.names(object), or a matrix with dimnames(d)[[1]] corresponding to row.names(object). The purpose of including d is primarily to print the rows of the original data corresponding to the most significant statistics. permax # An example is given in the permax help file
11 Index Topic hplot plot.expr, 6 plot.permax, 8 Topic htest permax, 1 permcor, 3 permsep, 5 Topic print summary.permax, 10 Topic utilities rowperm, 9 attr, 5 attributes, 5 permax, 1 permcor, 3 permsep, 5 plot.expr, 6 plot.permax, 8 print.summary.permax (summary.permax), 10 rowperm, 9 summary.permax, 10 11
Introduction to DSP ECE-S352 Fall Quarter 2000 Matlab Project 1
 Objective: Introduction to DSP ECE-S352 Fall Quarter 2000 Matlab Project 1 This Matlab Project is an extension of the basic correlation theory presented in the course. It shows a practical application
Objective: Introduction to DSP ECE-S352 Fall Quarter 2000 Matlab Project 1 This Matlab Project is an extension of the basic correlation theory presented in the course. It shows a practical application
Section 7: Using the Epilog Print Driver
 Color Mapping The Color Mapping feature is an advanced feature that must be checked to activate. Color Mapping performs two main functions: 1. It allows for multiple Speed and Power settings to be used
Color Mapping The Color Mapping feature is an advanced feature that must be checked to activate. Color Mapping performs two main functions: 1. It allows for multiple Speed and Power settings to be used
Package twilight. February 15, 2018
 Version 1.55.0 Title Estimation of local false discovery rate Package twilight February 15, 2018 Author Stefanie Scheid In a typical microarray setting with gene expression data
Version 1.55.0 Title Estimation of local false discovery rate Package twilight February 15, 2018 Author Stefanie Scheid In a typical microarray setting with gene expression data
NCSS Statistical Software
 Chapter 147 Introduction A mosaic plot is a graphical display of the cell frequencies of a contingency table in which the area of boxes of the plot are proportional to the cell frequencies of the contingency
Chapter 147 Introduction A mosaic plot is a graphical display of the cell frequencies of a contingency table in which the area of boxes of the plot are proportional to the cell frequencies of the contingency
Assessing Measurement System Variation
 Example 1 Fuel Injector Nozzle Diameters Problem A manufacturer of fuel injector nozzles has installed a new digital measuring system. Investigators want to determine how well the new system measures the
Example 1 Fuel Injector Nozzle Diameters Problem A manufacturer of fuel injector nozzles has installed a new digital measuring system. Investigators want to determine how well the new system measures the
Why Should We Care? Everyone uses plotting But most people ignore or are unaware of simple principles Default plotting tools are not always the best
 Elementary Plots Why Should We Care? Everyone uses plotting But most people ignore or are unaware of simple principles Default plotting tools are not always the best More importantly, it is easy to lie
Elementary Plots Why Should We Care? Everyone uses plotting But most people ignore or are unaware of simple principles Default plotting tools are not always the best More importantly, it is easy to lie
Introduction to ibbig
 Introduction to ibbig Aedin Culhane, Daniel Gusenleitner April 4, 2013 1 ibbig Iterative Binary Bi-clustering of Gene sets (ibbig) is a bi-clustering algorithm optimized for discovery of overlapping biclusters
Introduction to ibbig Aedin Culhane, Daniel Gusenleitner April 4, 2013 1 ibbig Iterative Binary Bi-clustering of Gene sets (ibbig) is a bi-clustering algorithm optimized for discovery of overlapping biclusters
Computer Programming ECIV 2303 Chapter 5 Two-Dimensional Plots Instructor: Dr. Talal Skaik Islamic University of Gaza Faculty of Engineering
 Computer Programming ECIV 2303 Chapter 5 Two-Dimensional Plots Instructor: Dr. Talal Skaik Islamic University of Gaza Faculty of Engineering 1 Introduction Plots are a very useful tool for presenting information.
Computer Programming ECIV 2303 Chapter 5 Two-Dimensional Plots Instructor: Dr. Talal Skaik Islamic University of Gaza Faculty of Engineering 1 Introduction Plots are a very useful tool for presenting information.
ELG3311: EXPERIMENT 2 Simulation of a Transformer Performance
 ELG33: EXPERIMENT 2 Simulation of a Transformer Performance Objective Using Matlab simulation toolbox (SIMULINK), design a model to simulate the performance of a single-phase transformer under different
ELG33: EXPERIMENT 2 Simulation of a Transformer Performance Objective Using Matlab simulation toolbox (SIMULINK), design a model to simulate the performance of a single-phase transformer under different
Ma 322: Biostatistics Solutions to Homework Assignment 1
 Ma 322: Biostatistics Solutions to Homework Assignment 1 Prof. Wickerhauser Due Friday, January 26th, 2018 Begin by obtaining access to the R software package, either by downloading a copy onto your computer
Ma 322: Biostatistics Solutions to Homework Assignment 1 Prof. Wickerhauser Due Friday, January 26th, 2018 Begin by obtaining access to the R software package, either by downloading a copy onto your computer
-f/d-b '') o, q&r{laniels, Advisor. 20rt. lmage Processing of Petrographic and SEM lmages. By James Gonsiewski. The Ohio State University
 lmage Processing of Petrographic and SEM lmages Senior Thesis Submitted in partial fulfillment of the requirements for the Bachelor of Science Degree At The Ohio State Universitv By By James Gonsiewski
lmage Processing of Petrographic and SEM lmages Senior Thesis Submitted in partial fulfillment of the requirements for the Bachelor of Science Degree At The Ohio State Universitv By By James Gonsiewski
Repeated Measures Twoway Analysis of Variance
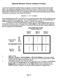 Repeated Measures Twoway Analysis of Variance A researcher was interested in whether frequency of exposure to a picture of an ugly or attractive person would influence one's liking for the photograph.
Repeated Measures Twoway Analysis of Variance A researcher was interested in whether frequency of exposure to a picture of an ugly or attractive person would influence one's liking for the photograph.
Counting and Probability Math 2320
 Counting and Probability Math 2320 For a finite set A, the number of elements of A is denoted by A. We have two important rules for counting. 1. Union rule: Let A and B be two finite sets. Then A B = A
Counting and Probability Math 2320 For a finite set A, the number of elements of A is denoted by A. We have two important rules for counting. 1. Union rule: Let A and B be two finite sets. Then A B = A
Assessing Measurement System Variation
 Assessing Measurement System Variation Example 1: Fuel Injector Nozzle Diameters Problem A manufacturer of fuel injector nozzles installs a new digital measuring system. Investigators want to determine
Assessing Measurement System Variation Example 1: Fuel Injector Nozzle Diameters Problem A manufacturer of fuel injector nozzles installs a new digital measuring system. Investigators want to determine
Why Should We Care? More importantly, it is easy to lie or deceive people with bad plots
 Elementary Plots Why Should We Care? Everyone uses plotting But most people ignore or are unaware of simple principles Default plotting tools (or default settings) are not always the best More importantly,
Elementary Plots Why Should We Care? Everyone uses plotting But most people ignore or are unaware of simple principles Default plotting tools (or default settings) are not always the best More importantly,
Quantitative Analysis of Tone Value Reproduction Limits
 Robert Chung* and Ping-hsu Chen* Keywords: Standard, Tonality, Highlight, Shadow, E* ab Abstract ISO 12647-2 (2004) defines tone value reproduction limits requirement as, half-tone dot patterns within
Robert Chung* and Ping-hsu Chen* Keywords: Standard, Tonality, Highlight, Shadow, E* ab Abstract ISO 12647-2 (2004) defines tone value reproduction limits requirement as, half-tone dot patterns within
Introduction to ibbig
 Introduction to ibbig Aedin Culhane, Daniel Gusenleitner June 13, 2018 1 ibbig Iterative Binary Bi-clustering of Gene sets (ibbig) is a bi-clustering algorithm optimized for discovery of overlapping biclusters
Introduction to ibbig Aedin Culhane, Daniel Gusenleitner June 13, 2018 1 ibbig Iterative Binary Bi-clustering of Gene sets (ibbig) is a bi-clustering algorithm optimized for discovery of overlapping biclusters
Basics of Probability
 Basics of Probability Dublin R May 30, 2013 1 Overview Overview Basics of Probability (some definitions, the prob package) Dice Rolls and the Birthday Distribution ( histograms ) Gambler s Ruin ( plotting
Basics of Probability Dublin R May 30, 2013 1 Overview Overview Basics of Probability (some definitions, the prob package) Dice Rolls and the Birthday Distribution ( histograms ) Gambler s Ruin ( plotting
Package MLP. April 14, 2013
 Package MLP April 14, 2013 Maintainer Tobias Verbeke License GPL-3 Title MLP Type Package Author Nandini Raghavan, Tobias Verbeke, An De Bondt with contributions by Javier
Package MLP April 14, 2013 Maintainer Tobias Verbeke License GPL-3 Title MLP Type Package Author Nandini Raghavan, Tobias Verbeke, An De Bondt with contributions by Javier
Chapter 3 WORLDWIDE PATENTING ACTIVITY
 Chapter 3 WORLDWIDE PATENTING ACTIVITY Patent activity is recognized throughout the world as an indicator of innovation. This chapter examines worldwide patent activities in terms of patent applications
Chapter 3 WORLDWIDE PATENTING ACTIVITY Patent activity is recognized throughout the world as an indicator of innovation. This chapter examines worldwide patent activities in terms of patent applications
!"#$%&'("&)*("*+,)-(#'.*/$'-0%$1$"&-!!!"#$%&'(!"!!"#$%"&&'()*+*!
 !"#$%&'("&)*("*+,)-(#'.*/$'-0%$1$"&-!!!"#$%&'(!"!!"#$%"&&'()*+*! In this Module, we will consider dice. Although people have been gambling with dice and related apparatus since at least 3500 BCE, amazingly
!"#$%&'("&)*("*+,)-(#'.*/$'-0%$1$"&-!!!"#$%&'(!"!!"#$%"&&'()*+*! In this Module, we will consider dice. Although people have been gambling with dice and related apparatus since at least 3500 BCE, amazingly
Department of Statistics and Operations Research Undergraduate Programmes
 Department of Statistics and Operations Research Undergraduate Programmes OPERATIONS RESEARCH YEAR LEVEL 2 INTRODUCTION TO LINEAR PROGRAMMING SSOA021 Linear Programming Model: Formulation of an LP model;
Department of Statistics and Operations Research Undergraduate Programmes OPERATIONS RESEARCH YEAR LEVEL 2 INTRODUCTION TO LINEAR PROGRAMMING SSOA021 Linear Programming Model: Formulation of an LP model;
Acceptance Charts. Sample StatFolio: acceptance chart.sgp
 Acceptance Charts Summary The Acceptance Charts procedure creates control charts with modified control limits based on both the standard deviation of the process and on specification limits for the variable
Acceptance Charts Summary The Acceptance Charts procedure creates control charts with modified control limits based on both the standard deviation of the process and on specification limits for the variable
Correlation and Regression
 Correlation and Regression Shepard and Feng (1972) presented participants with an unfolded cube and asked them to mentally refold the cube with the shaded square on the bottom to determine if the two arrows
Correlation and Regression Shepard and Feng (1972) presented participants with an unfolded cube and asked them to mentally refold the cube with the shaded square on the bottom to determine if the two arrows
EFFECTS OF PHASE AND AMPLITUDE ERRORS ON QAM SYSTEMS WITH ERROR- CONTROL CODING AND SOFT DECISION DECODING
 Clemson University TigerPrints All Theses Theses 8-2009 EFFECTS OF PHASE AND AMPLITUDE ERRORS ON QAM SYSTEMS WITH ERROR- CONTROL CODING AND SOFT DECISION DECODING Jason Ellis Clemson University, jellis@clemson.edu
Clemson University TigerPrints All Theses Theses 8-2009 EFFECTS OF PHASE AND AMPLITUDE ERRORS ON QAM SYSTEMS WITH ERROR- CONTROL CODING AND SOFT DECISION DECODING Jason Ellis Clemson University, jellis@clemson.edu
Ansoft Designer Tutorial ECE 584 October, 2004
 Ansoft Designer Tutorial ECE 584 October, 2004 This tutorial will serve as an introduction to the Ansoft Designer Microwave CAD package by stepping through a simple design problem. Please note that there
Ansoft Designer Tutorial ECE 584 October, 2004 This tutorial will serve as an introduction to the Ansoft Designer Microwave CAD package by stepping through a simple design problem. Please note that there
28th Seismic Research Review: Ground-Based Nuclear Explosion Monitoring Technologies
 8th Seismic Research Review: Ground-Based Nuclear Explosion Monitoring Technologies A LOWER BOUND ON THE STANDARD ERROR OF AN AMPLITUDE-BASED REGIONAL DISCRIMINANT D. N. Anderson 1, W. R. Walter, D. K.
8th Seismic Research Review: Ground-Based Nuclear Explosion Monitoring Technologies A LOWER BOUND ON THE STANDARD ERROR OF AN AMPLITUDE-BASED REGIONAL DISCRIMINANT D. N. Anderson 1, W. R. Walter, D. K.
Statistics 101: Section L Laboratory 10
 Statistics 101: Section L Laboratory 10 This lab looks at the sampling distribution of the sample proportion pˆ and probabilities associated with sampling from a population with a categorical variable.
Statistics 101: Section L Laboratory 10 This lab looks at the sampling distribution of the sample proportion pˆ and probabilities associated with sampling from a population with a categorical variable.
Toolwear Charts. Sample StatFolio: toolwear chart.sgp. Sample Data: STATGRAPHICS Rev. 9/16/2013
 Toolwear Charts Summary... 1 Data Input... 2 Toolwear Chart... 5 Analysis Summary... 6 Analysis Options... 7 MR(2)/R/S Chart... 8 Toolwear Chart Report... 10 Runs Tests... 10 Tolerance Chart... 11 Save
Toolwear Charts Summary... 1 Data Input... 2 Toolwear Chart... 5 Analysis Summary... 6 Analysis Options... 7 MR(2)/R/S Chart... 8 Toolwear Chart Report... 10 Runs Tests... 10 Tolerance Chart... 11 Save
Grades 6 8 Innoventure Components That Meet Common Core Mathematics Standards
 Grades 6 8 Innoventure Components That Meet Common Core Mathematics Standards Strand Ratios and Relationships The Number System Expressions and Equations Anchor Standard Understand ratio concepts and use
Grades 6 8 Innoventure Components That Meet Common Core Mathematics Standards Strand Ratios and Relationships The Number System Expressions and Equations Anchor Standard Understand ratio concepts and use
Summary... 1 Sample Data... 2 Data Input... 3 C Chart... 4 C Chart Report... 6 Analysis Summary... 7 Analysis Options... 8 Save Results...
 C Chart Summary... 1 Sample Data... 2 Data Input... 3 C Chart... 4 C Chart Report... 6 Analysis Summary... 7 Analysis Options... 8 Save Results... 9 Summary The C Chart procedure creates a control chart
C Chart Summary... 1 Sample Data... 2 Data Input... 3 C Chart... 4 C Chart Report... 6 Analysis Summary... 7 Analysis Options... 8 Save Results... 9 Summary The C Chart procedure creates a control chart
Computer Graphics: Graphics Output Primitives Primitives Attributes
 Computer Graphics: Graphics Output Primitives Primitives Attributes By: A. H. Abdul Hafez Abdul.hafez@hku.edu.tr, 1 Outlines 1. OpenGL state variables 2. RGB color components 1. direct color storage 2.
Computer Graphics: Graphics Output Primitives Primitives Attributes By: A. H. Abdul Hafez Abdul.hafez@hku.edu.tr, 1 Outlines 1. OpenGL state variables 2. RGB color components 1. direct color storage 2.
Package pedigreemm. R topics documented: February 20, 2015
 Version 0.3-3 Date 2013-09-27 Title Pedigree-based mixed-effects models Author Douglas Bates and Ana Ines Vazquez, Package pedigreemm February 20, 2015 Maintainer Ana Ines Vazquez
Version 0.3-3 Date 2013-09-27 Title Pedigree-based mixed-effects models Author Douglas Bates and Ana Ines Vazquez, Package pedigreemm February 20, 2015 Maintainer Ana Ines Vazquez
Remote Sensing 4113 Lab 08: Filtering and Principal Components Mar. 28, 2018
 Remote Sensing 4113 Lab 08: Filtering and Principal Components Mar. 28, 2018 In this lab we will explore Filtering and Principal Components analysis. We will again use the Aster data of the Como Bluffs
Remote Sensing 4113 Lab 08: Filtering and Principal Components Mar. 28, 2018 In this lab we will explore Filtering and Principal Components analysis. We will again use the Aster data of the Como Bluffs
Computers and Imaging
 Computers and Imaging Telecommunications 1 P. Mathys Two Different Methods Vector or object-oriented graphics. Images are generated by mathematical descriptions of line (vector) segments. Bitmap or raster
Computers and Imaging Telecommunications 1 P. Mathys Two Different Methods Vector or object-oriented graphics. Images are generated by mathematical descriptions of line (vector) segments. Bitmap or raster
Solutions 2: Probability and Counting
 Massachusetts Institute of Technology MITES 18 Physics III Solutions : Probability and Counting Due Tuesday July 3 at 11:59PM under Fernando Rendon s door Preface: The basic methods of probability and
Massachusetts Institute of Technology MITES 18 Physics III Solutions : Probability and Counting Due Tuesday July 3 at 11:59PM under Fernando Rendon s door Preface: The basic methods of probability and
Design of Parallel Algorithms. Communication Algorithms
 + Design of Parallel Algorithms Communication Algorithms + Topic Overview n One-to-All Broadcast and All-to-One Reduction n All-to-All Broadcast and Reduction n All-Reduce and Prefix-Sum Operations n Scatter
+ Design of Parallel Algorithms Communication Algorithms + Topic Overview n One-to-All Broadcast and All-to-One Reduction n All-to-All Broadcast and Reduction n All-Reduce and Prefix-Sum Operations n Scatter
Package RVtests. R topics documented: February 19, 2015
 Type Package Title Rare Variant Tests Version 1.2 Date 2013-05-27 Author, and C. M. Greenwood Package RVtests February 19, 2015 Maintainer Depends R (>= 2.12.1), glmnet,
Type Package Title Rare Variant Tests Version 1.2 Date 2013-05-27 Author, and C. M. Greenwood Package RVtests February 19, 2015 Maintainer Depends R (>= 2.12.1), glmnet,
Image Extraction using Image Mining Technique
 IOSR Journal of Engineering (IOSRJEN) e-issn: 2250-3021, p-issn: 2278-8719 Vol. 3, Issue 9 (September. 2013), V2 PP 36-42 Image Extraction using Image Mining Technique Prof. Samir Kumar Bandyopadhyay,
IOSR Journal of Engineering (IOSRJEN) e-issn: 2250-3021, p-issn: 2278-8719 Vol. 3, Issue 9 (September. 2013), V2 PP 36-42 Image Extraction using Image Mining Technique Prof. Samir Kumar Bandyopadhyay,
4.5.1 Mirroring Gain/Offset Registers GPIO CMV Snapshot Control... 14
 Thank you for choosing the MityCAM-C8000 from Critical Link. The MityCAM-C8000 MityViewer Quick Start Guide will guide you through the software installation process and the steps to acquire your first
Thank you for choosing the MityCAM-C8000 from Critical Link. The MityCAM-C8000 MityViewer Quick Start Guide will guide you through the software installation process and the steps to acquire your first
Drawing Bode Plots (The Last Bode Plot You Will Ever Make) Charles Nippert
 Drawing Bode Plots (The Last Bode Plot You Will Ever Make) Charles Nippert This set of notes describes how to prepare a Bode plot using Mathcad. Follow these instructions to draw Bode plot for any transfer
Drawing Bode Plots (The Last Bode Plot You Will Ever Make) Charles Nippert This set of notes describes how to prepare a Bode plot using Mathcad. Follow these instructions to draw Bode plot for any transfer
Brief Introduction to Vision and Images
 Brief Introduction to Vision and Images Charles S. Tritt, Ph.D. January 24, 2012 Version 1.1 Structure of the Retina There is only one kind of rod. Rods are very sensitive and used mainly in dim light.
Brief Introduction to Vision and Images Charles S. Tritt, Ph.D. January 24, 2012 Version 1.1 Structure of the Retina There is only one kind of rod. Rods are very sensitive and used mainly in dim light.
Section 6.4. Sampling Distributions and Estimators
 Section 6.4 Sampling Distributions and Estimators IDEA Ch 5 and part of Ch 6 worked with population. Now we are going to work with statistics. Sample Statistics to estimate population parameters. To make
Section 6.4 Sampling Distributions and Estimators IDEA Ch 5 and part of Ch 6 worked with population. Now we are going to work with statistics. Sample Statistics to estimate population parameters. To make
EE 435/535: Error Correcting Codes Project 1, Fall 2009: Extended Hamming Code. 1 Introduction. 2 Extended Hamming Code: Encoding. 1.
 EE 435/535: Error Correcting Codes Project 1, Fall 2009: Extended Hamming Code Project #1 is due on Tuesday, October 6, 2009, in class. You may turn the project report in early. Late projects are accepted
EE 435/535: Error Correcting Codes Project 1, Fall 2009: Extended Hamming Code Project #1 is due on Tuesday, October 6, 2009, in class. You may turn the project report in early. Late projects are accepted
Digital Image Processing. Lecture # 6 Corner Detection & Color Processing
 Digital Image Processing Lecture # 6 Corner Detection & Color Processing 1 Corners Corners (interest points) Unlike edges, corners (patches of pixels surrounding the corner) do not necessarily correspond
Digital Image Processing Lecture # 6 Corner Detection & Color Processing 1 Corners Corners (interest points) Unlike edges, corners (patches of pixels surrounding the corner) do not necessarily correspond
Stitching MetroPro Application
 OMP-0375F Stitching MetroPro Application Stitch.app This booklet is a quick reference; it assumes that you are familiar with MetroPro and the instrument. Information on MetroPro is provided in Getting
OMP-0375F Stitching MetroPro Application Stitch.app This booklet is a quick reference; it assumes that you are familiar with MetroPro and the instrument. Information on MetroPro is provided in Getting
In this talk I will be talking about improving the accuracy of S phase estimation from cytometric data containing DNA content. A new method of interpo
 In this talk I will be talking about improving the accuracy of S phase estimation from cytometric data containing DNA content. A new method of interpolation, parabolic splines (PS), for Probability State
In this talk I will be talking about improving the accuracy of S phase estimation from cytometric data containing DNA content. A new method of interpolation, parabolic splines (PS), for Probability State
The Bead. beadarray: : An R Package for Illumina BeadArrays. Bead Preparation and Array Production. Beads in Wells. Mark Dunning -
 beadarray: : An R Package for Illumina BeadArrays Mark Dunning - md392@cam.ac.uk PhD Student - Computational Biology Group, Department of Oncology - University of Cambridge Address The Bead Probe 23 b
beadarray: : An R Package for Illumina BeadArrays Mark Dunning - md392@cam.ac.uk PhD Student - Computational Biology Group, Department of Oncology - University of Cambridge Address The Bead Probe 23 b
L2. Image processing in MATLAB
 L2. Image processing in MATLAB 1. Introduction MATLAB environment offers an easy way to prototype applications that are based on complex mathematical computations. This annex presents some basic image
L2. Image processing in MATLAB 1. Introduction MATLAB environment offers an easy way to prototype applications that are based on complex mathematical computations. This annex presents some basic image
4/9/2015. Simple Graphics and Image Processing. Simple Graphics. Overview of Turtle Graphics (continued) Overview of Turtle Graphics
 Simple Graphics and Image Processing The Plan For Today Website Updates Intro to Python Quiz Corrections Missing Assignments Graphics and Images Simple Graphics Turtle Graphics Image Processing Assignment
Simple Graphics and Image Processing The Plan For Today Website Updates Intro to Python Quiz Corrections Missing Assignments Graphics and Images Simple Graphics Turtle Graphics Image Processing Assignment
Digital Image Processing. Lecture # 8 Color Processing
 Digital Image Processing Lecture # 8 Color Processing 1 COLOR IMAGE PROCESSING COLOR IMAGE PROCESSING Color Importance Color is an excellent descriptor Suitable for object Identification and Extraction
Digital Image Processing Lecture # 8 Color Processing 1 COLOR IMAGE PROCESSING COLOR IMAGE PROCESSING Color Importance Color is an excellent descriptor Suitable for object Identification and Extraction
9.1. Probability and Statistics
 9. Probability and Statistics Measured signals exhibit deterministic (predictable) and random (unpredictable) behavior. The deterministic behavior is often governed by a differential equation, while the
9. Probability and Statistics Measured signals exhibit deterministic (predictable) and random (unpredictable) behavior. The deterministic behavior is often governed by a differential equation, while the
Unit Nine Precalculus Practice Test Probability & Statistics. Name: Period: Date: NON-CALCULATOR SECTION
 Name: Period: Date: NON-CALCULATOR SECTION Vocabulary: Define each word and give an example. 1. discrete mathematics 2. dependent outcomes 3. series Short Answer: 4. Describe when to use a combination.
Name: Period: Date: NON-CALCULATOR SECTION Vocabulary: Define each word and give an example. 1. discrete mathematics 2. dependent outcomes 3. series Short Answer: 4. Describe when to use a combination.
GREATER CLARK COUNTY SCHOOLS PACING GUIDE. Algebra I MATHEMATICS G R E A T E R C L A R K C O U N T Y S C H O O L S
 GREATER CLARK COUNTY SCHOOLS PACING GUIDE Algebra I MATHEMATICS 2014-2015 G R E A T E R C L A R K C O U N T Y S C H O O L S ANNUAL PACING GUIDE Quarter/Learning Check Days (Approx) Q1/LC1 11 Concept/Skill
GREATER CLARK COUNTY SCHOOLS PACING GUIDE Algebra I MATHEMATICS 2014-2015 G R E A T E R C L A R K C O U N T Y S C H O O L S ANNUAL PACING GUIDE Quarter/Learning Check Days (Approx) Q1/LC1 11 Concept/Skill
Combinatorics and Intuitive Probability
 Chapter Combinatorics and Intuitive Probability The simplest probabilistic scenario is perhaps one where the set of possible outcomes is finite and these outcomes are all equally likely. A subset of the
Chapter Combinatorics and Intuitive Probability The simplest probabilistic scenario is perhaps one where the set of possible outcomes is finite and these outcomes are all equally likely. A subset of the
CSCD 409 Scientific Programming. Module 6: Plotting (Chpt 5)
 CSCD 409 Scientific Programming Module 6: Plotting (Chpt 5) 2008-2012, Prentice Hall, Paul Schimpf All rights reserved. No portion of this presentation may be reproduced, in whole or in part, in any form
CSCD 409 Scientific Programming Module 6: Plotting (Chpt 5) 2008-2012, Prentice Hall, Paul Schimpf All rights reserved. No portion of this presentation may be reproduced, in whole or in part, in any form
MATLAB Image Processing Toolbox
 MATLAB Image Processing Toolbox Copyright: Mathworks 1998. The following is taken from the Matlab Image Processing Toolbox users guide. A complete online manual is availabe in the PDF form (about 5MB).
MATLAB Image Processing Toolbox Copyright: Mathworks 1998. The following is taken from the Matlab Image Processing Toolbox users guide. A complete online manual is availabe in the PDF form (about 5MB).
This week we will work with your Landsat images and classify them using supervised classification.
 GEPL 4500/5500 Lab 4: Supervised Classification: Part I: Selecting Training Sets Due: 4/6/04 This week we will work with your Landsat images and classify them using supervised classification. There are
GEPL 4500/5500 Lab 4: Supervised Classification: Part I: Selecting Training Sets Due: 4/6/04 This week we will work with your Landsat images and classify them using supervised classification. There are
ImagesPlus Basic Interface Operation
 ImagesPlus Basic Interface Operation The basic interface operation menu options are located on the File, View, Open Images, Open Operators, and Help main menus. File Menu New The New command creates a
ImagesPlus Basic Interface Operation The basic interface operation menu options are located on the File, View, Open Images, Open Operators, and Help main menus. File Menu New The New command creates a
1 Running the Program
 GNUbik Copyright c 1998,2003 John Darrington 2004 John Darrington, Dale Mellor Permission is granted to make and distribute verbatim copies of this manual provided the copyright notice and this permission
GNUbik Copyright c 1998,2003 John Darrington 2004 John Darrington, Dale Mellor Permission is granted to make and distribute verbatim copies of this manual provided the copyright notice and this permission
Class #16: Experiment Matlab and Data Analysis
 Class #16: Experiment Matlab and Data Analysis Purpose: The objective of this experiment is to add to our Matlab skill set so that data can be easily plotted and analyzed with simple tools. Background:
Class #16: Experiment Matlab and Data Analysis Purpose: The objective of this experiment is to add to our Matlab skill set so that data can be easily plotted and analyzed with simple tools. Background:
Princeton ELE 201, Spring 2014 Laboratory No. 2 Shazam
 Princeton ELE 201, Spring 2014 Laboratory No. 2 Shazam 1 Background In this lab we will begin to code a Shazam-like program to identify a short clip of music using a database of songs. The basic procedure
Princeton ELE 201, Spring 2014 Laboratory No. 2 Shazam 1 Background In this lab we will begin to code a Shazam-like program to identify a short clip of music using a database of songs. The basic procedure
EXPERIMENT 9 Problem Solving: First-order Transient Circuits
 EXPERIMENT 9 Problem Solving: First-order Transient Circuits I. Introduction In transient analyses, we determine voltages and currents as functions of time. Typically, the time dependence is demonstrated
EXPERIMENT 9 Problem Solving: First-order Transient Circuits I. Introduction In transient analyses, we determine voltages and currents as functions of time. Typically, the time dependence is demonstrated
Heads Up! A c t i v i t y 5. The Problem. Name Date
 . Name Date A c t i v i t y 5 Heads Up! In this activity, you will study some important concepts in a branch of mathematics known as probability. You are using probability when you say things like: It
. Name Date A c t i v i t y 5 Heads Up! In this activity, you will study some important concepts in a branch of mathematics known as probability. You are using probability when you say things like: It
Most typical tests can also be done as permutation tests. For example: Two sample tests (e.g., t-test, MWU test)
 Permutation tests: Permutation tests are a large group of statistical procedures. Most typical tests can also be done as permutation tests. For example: Two sample tests (e.g., t-test, MWU test) Three
Permutation tests: Permutation tests are a large group of statistical procedures. Most typical tests can also be done as permutation tests. For example: Two sample tests (e.g., t-test, MWU test) Three
Exam Time. Final Exam Review. TR class Monday December 9 12:30 2:30. These review slides and earlier ones found linked to on BlackBoard
 Final Exam Review These review slides and earlier ones found linked to on BlackBoard Bring a photo ID card: Rocket Card, Driver's License Exam Time TR class Monday December 9 12:30 2:30 Held in the regular
Final Exam Review These review slides and earlier ones found linked to on BlackBoard Bring a photo ID card: Rocket Card, Driver's License Exam Time TR class Monday December 9 12:30 2:30 Held in the regular
Package beadarrayfilter
 Type Package Package beadarrayfilter Title Bead filtering for Illumina bead arrays Version 1.1.0 Date 2013-02-04 February 19, 2015 Author Anyiawung Chiara Forcheh, Geert Verbeke, Adetayo Kasim, Dan Lin,
Type Package Package beadarrayfilter Title Bead filtering for Illumina bead arrays Version 1.1.0 Date 2013-02-04 February 19, 2015 Author Anyiawung Chiara Forcheh, Geert Verbeke, Adetayo Kasim, Dan Lin,
3) Start ImageJ, install CM Engine as a macro (instructions here:
 Instructions for CM Engine use 1) Download CM Engine from SourceForge (http://cm- engine.sourceforge.net/) or from the Rothstein Lab website (http://www.rothsteinlab.com/cm- engine.zip ). 2) Download ImageJ
Instructions for CM Engine use 1) Download CM Engine from SourceForge (http://cm- engine.sourceforge.net/) or from the Rothstein Lab website (http://www.rothsteinlab.com/cm- engine.zip ). 2) Download ImageJ
Figure 1: Energy Distributions for light
 Lecture 4: Colour The physical description of colour Colour vision is a very complicated biological and psychological phenomenon. It can be described in many different ways, including by physics, by subjective
Lecture 4: Colour The physical description of colour Colour vision is a very complicated biological and psychological phenomenon. It can be described in many different ways, including by physics, by subjective
Matlab for CS6320 Beginners
 Matlab for CS6320 Beginners Basics: Starting Matlab o CADE Lab remote access o Student version on your own computer Change the Current Folder to the directory where your programs, images, etc. will be
Matlab for CS6320 Beginners Basics: Starting Matlab o CADE Lab remote access o Student version on your own computer Change the Current Folder to the directory where your programs, images, etc. will be
Describing Data Visually. Describing Data Visually. Describing Data Visually 9/28/12. Applied Statistics in Business & Economics, 4 th edition
 A PowerPoint Presentation Package to Accompany Applied Statistics in Business & Economics, 4 th edition David P. Doane and Lori E. Seward Prepared by Lloyd R. Jaisingh Describing Data Visually Chapter
A PowerPoint Presentation Package to Accompany Applied Statistics in Business & Economics, 4 th edition David P. Doane and Lori E. Seward Prepared by Lloyd R. Jaisingh Describing Data Visually Chapter
Core Learning Standards for Mathematics Grade 6
 Core Learning Standards for Mathematics Grade 6 Write and evaluate numerical expressions involving whole-number exponents. Write, read, and evaluate expressions; identify parts of an expression using mathematical
Core Learning Standards for Mathematics Grade 6 Write and evaluate numerical expressions involving whole-number exponents. Write, read, and evaluate expressions; identify parts of an expression using mathematical
PASS Sample Size Software. These options specify the characteristics of the lines, labels, and tick marks along the X and Y axes.
 Chapter 940 Introduction This section describes the options that are available for the appearance of a scatter plot. A set of all these options can be stored as a template file which can be retrieved later.
Chapter 940 Introduction This section describes the options that are available for the appearance of a scatter plot. A set of all these options can be stored as a template file which can be retrieved later.
AgilEye Manual Version 2.0 February 28, 2007
 AgilEye Manual Version 2.0 February 28, 2007 1717 Louisiana NE Suite 202 Albuquerque, NM 87110 (505) 268-4742 support@agiloptics.com 2 (505) 268-4742 v. 2.0 February 07, 2007 3 Introduction AgilEye Wavefront
AgilEye Manual Version 2.0 February 28, 2007 1717 Louisiana NE Suite 202 Albuquerque, NM 87110 (505) 268-4742 support@agiloptics.com 2 (505) 268-4742 v. 2.0 February 07, 2007 3 Introduction AgilEye Wavefront
Bias errors in PIV: the pixel locking effect revisited.
 Bias errors in PIV: the pixel locking effect revisited. E.F.J. Overmars 1, N.G.W. Warncke, C. Poelma and J. Westerweel 1: Laboratory for Aero & Hydrodynamics, University of Technology, Delft, The Netherlands,
Bias errors in PIV: the pixel locking effect revisited. E.F.J. Overmars 1, N.G.W. Warncke, C. Poelma and J. Westerweel 1: Laboratory for Aero & Hydrodynamics, University of Technology, Delft, The Netherlands,
ACTIVITY 6.7 Selecting and Rearranging Things
 ACTIVITY 6.7 SELECTING AND REARRANGING THINGS 757 OBJECTIVES ACTIVITY 6.7 Selecting and Rearranging Things 1. Determine the number of permutations. 2. Determine the number of combinations. 3. Recognize
ACTIVITY 6.7 SELECTING AND REARRANGING THINGS 757 OBJECTIVES ACTIVITY 6.7 Selecting and Rearranging Things 1. Determine the number of permutations. 2. Determine the number of combinations. 3. Recognize
MATLAB - Lecture # 5
 MATLAB - Lecture # 5 Two Dimensional Plots / Chapter 5 Topics Covered: 1. Plotting basic 2-D plots. The plot command. The fplot command. Plotting multiple graphs in the same plot. MAKING X-Y PLOTS 105
MATLAB - Lecture # 5 Two Dimensional Plots / Chapter 5 Topics Covered: 1. Plotting basic 2-D plots. The plot command. The fplot command. Plotting multiple graphs in the same plot. MAKING X-Y PLOTS 105
(Catalog Number 1746 NR4) Product Data
 (Catalog Number 1746 NR4) Product Data The 1746 NR4 / RTD sensor combination is easy to install and provides greater output (ohms/ C or ohms/ F), accuracy, linearity and repeatability with temperature,
(Catalog Number 1746 NR4) Product Data The 1746 NR4 / RTD sensor combination is easy to install and provides greater output (ohms/ C or ohms/ F), accuracy, linearity and repeatability with temperature,
Two-dimensional Plots
 Two-dimensional Plots ELEC 206 Prof. Siripong Potisuk 1 The Plot Command The simplest command for 2-D plotting Syntax: >> plot(x,y) The arguments x and y are vectors (1-D arrays) which must be of the same
Two-dimensional Plots ELEC 206 Prof. Siripong Potisuk 1 The Plot Command The simplest command for 2-D plotting Syntax: >> plot(x,y) The arguments x and y are vectors (1-D arrays) which must be of the same
STAB22 section 2.4. Figure 2: Data set 2. Figure 1: Data set 1
 STAB22 section 2.4 2.73 The four correlations are all 0.816, and all four regressions are ŷ = 3 + 0.5x. (b) can be answered by drawing fitted line plots in the four cases. See Figures 1, 2, 3 and 4. Figure
STAB22 section 2.4 2.73 The four correlations are all 0.816, and all four regressions are ŷ = 3 + 0.5x. (b) can be answered by drawing fitted line plots in the four cases. See Figures 1, 2, 3 and 4. Figure
Package timeseq. July 17, 2017
 Type Package Package timeseq July 17, 2017 Title Detecting Differentially Expressed Genes in Time Course RNA-Seq Data Version 1.0.3 Date 2017-7-17 Author Fan Gao, Xiaoxiao Sun Maintainer Fan Gao
Type Package Package timeseq July 17, 2017 Title Detecting Differentially Expressed Genes in Time Course RNA-Seq Data Version 1.0.3 Date 2017-7-17 Author Fan Gao, Xiaoxiao Sun Maintainer Fan Gao
WORLDWIDE PATENTING ACTIVITY
 WORLDWIDE PATENTING ACTIVITY IP5 Statistics Report 2011 Patent activity is recognized throughout the world as a measure of innovation. This chapter examines worldwide patent activities in terms of patent
WORLDWIDE PATENTING ACTIVITY IP5 Statistics Report 2011 Patent activity is recognized throughout the world as a measure of innovation. This chapter examines worldwide patent activities in terms of patent
Hyperspectral image processing and analysis
 Hyperspectral image processing and analysis Lecture 12 www.utsa.edu/lrsg/teaching/ees5083/l12-hyper.ppt Multi- vs. Hyper- Hyper-: Narrow bands ( 20 nm in resolution or FWHM) and continuous measurements.
Hyperspectral image processing and analysis Lecture 12 www.utsa.edu/lrsg/teaching/ees5083/l12-hyper.ppt Multi- vs. Hyper- Hyper-: Narrow bands ( 20 nm in resolution or FWHM) and continuous measurements.
A Kinect-based 3D hand-gesture interface for 3D databases
 A Kinect-based 3D hand-gesture interface for 3D databases Abstract. The use of natural interfaces improves significantly aspects related to human-computer interaction and consequently the productivity
A Kinect-based 3D hand-gesture interface for 3D databases Abstract. The use of natural interfaces improves significantly aspects related to human-computer interaction and consequently the productivity
Package Anaquin. January 12, 2019
 Type Package Title Statistical analysis of sequins Version 2.6.1 Date 2017-08-08 Author Ted Wong Package Anaquin January 12, 2019 Maintainer Ted Wong The project is intended to support
Type Package Title Statistical analysis of sequins Version 2.6.1 Date 2017-08-08 Author Ted Wong Package Anaquin January 12, 2019 Maintainer Ted Wong The project is intended to support
 CMPS 12A Introduction to Programming Programming Assignment 5 In this assignment you will write a Java program that finds all solutions to the n-queens problem, for. Begin by reading the Wikipedia article
CMPS 12A Introduction to Programming Programming Assignment 5 In this assignment you will write a Java program that finds all solutions to the n-queens problem, for. Begin by reading the Wikipedia article
MAS336 Computational Problem Solving. Problem 3: Eight Queens
 MAS336 Computational Problem Solving Problem 3: Eight Queens Introduction Francis J. Wright, 2007 Topics: arrays, recursion, plotting, symmetry The problem is to find all the distinct ways of choosing
MAS336 Computational Problem Solving Problem 3: Eight Queens Introduction Francis J. Wright, 2007 Topics: arrays, recursion, plotting, symmetry The problem is to find all the distinct ways of choosing
17. Symmetries. Thus, the example above corresponds to the matrix: We shall now look at how permutations relate to trees.
 7 Symmetries 7 Permutations A permutation of a set is a reordering of its elements Another way to look at it is as a function Φ that takes as its argument a set of natural numbers of the form {, 2,, n}
7 Symmetries 7 Permutations A permutation of a set is a reordering of its elements Another way to look at it is as a function Φ that takes as its argument a set of natural numbers of the form {, 2,, n}
Measurement Systems Analysis
 11 Measurement Systems Analysis Measurement Systems Analysis Overview, 11-2, 11-4 Gage Run Chart, 11-23 Gage Linearity and Accuracy Study, 11-27 MINITAB User s Guide 2 11-1 Chapter 11 Measurement Systems
11 Measurement Systems Analysis Measurement Systems Analysis Overview, 11-2, 11-4 Gage Run Chart, 11-23 Gage Linearity and Accuracy Study, 11-27 MINITAB User s Guide 2 11-1 Chapter 11 Measurement Systems
Evaluation of image quality of the compression schemes JPEG & JPEG 2000 using a Modular Colour Image Difference Model.
 Evaluation of image quality of the compression schemes JPEG & JPEG 2000 using a Modular Colour Image Difference Model. Mary Orfanidou, Liz Allen and Dr Sophie Triantaphillidou, University of Westminster,
Evaluation of image quality of the compression schemes JPEG & JPEG 2000 using a Modular Colour Image Difference Model. Mary Orfanidou, Liz Allen and Dr Sophie Triantaphillidou, University of Westminster,
CS534 Introduction to Computer Vision. Linear Filters. Ahmed Elgammal Dept. of Computer Science Rutgers University
 CS534 Introduction to Computer Vision Linear Filters Ahmed Elgammal Dept. of Computer Science Rutgers University Outlines What are Filters Linear Filters Convolution operation Properties of Linear Filters
CS534 Introduction to Computer Vision Linear Filters Ahmed Elgammal Dept. of Computer Science Rutgers University Outlines What are Filters Linear Filters Convolution operation Properties of Linear Filters
GE U111 HTT&TL, Lab 1: The Speed of Sound in Air, Acoustic Distance Measurement & Basic Concepts in MATLAB
 GE U111 HTT&TL, Lab 1: The Speed of Sound in Air, Acoustic Distance Measurement & Basic Concepts in MATLAB Contents 1 Preview: Programming & Experiments Goals 2 2 Homework Assignment 3 3 Measuring The
GE U111 HTT&TL, Lab 1: The Speed of Sound in Air, Acoustic Distance Measurement & Basic Concepts in MATLAB Contents 1 Preview: Programming & Experiments Goals 2 2 Homework Assignment 3 3 Measuring The
(i) Understanding the basic concepts of signal modeling, correlation, maximum likelihood estimation, least squares and iterative numerical methods
 Tools and Applications Chapter Intended Learning Outcomes: (i) Understanding the basic concepts of signal modeling, correlation, maximum likelihood estimation, least squares and iterative numerical methods
Tools and Applications Chapter Intended Learning Outcomes: (i) Understanding the basic concepts of signal modeling, correlation, maximum likelihood estimation, least squares and iterative numerical methods
GLOSSARY. a * (b * c) = (a * b) * c. A property of operations. An operation * is called associative if:
 Associativity A property of operations. An operation * is called associative if: a * (b * c) = (a * b) * c for every possible a, b, and c. Axiom For Greek geometry, an axiom was a 'self-evident truth'.
Associativity A property of operations. An operation * is called associative if: a * (b * c) = (a * b) * c for every possible a, b, and c. Axiom For Greek geometry, an axiom was a 'self-evident truth'.
6. Multivariate EDA. ACE 492 SA - Spatial Analysis Fall 2003
 1 Objectives 6. Multivariate EDA ACE 492 SA - Spatial Analysis Fall 2003 c 2003 by Luc Anselin, All Rights Reserved This lab covers some basic approaches to carry out EDA with a focus on discovering multivariate
1 Objectives 6. Multivariate EDA ACE 492 SA - Spatial Analysis Fall 2003 c 2003 by Luc Anselin, All Rights Reserved This lab covers some basic approaches to carry out EDA with a focus on discovering multivariate
Keysight Technologies Making Accurate Intermodulation Distortion Measurements with the PNA-X Network Analyzer, 10 MHz to 26.5 GHz
 Keysight Technologies Making Accurate Intermodulation Distortion Measurements with the PNA-X Network Analyzer, 10 MHz to 26.5 GHz Application Note Overview This application note describes accuracy considerations
Keysight Technologies Making Accurate Intermodulation Distortion Measurements with the PNA-X Network Analyzer, 10 MHz to 26.5 GHz Application Note Overview This application note describes accuracy considerations
Autodesk Moldflow Insight AMI Shrink Analysis Results
 Autodesk Moldflow Insight 2012 AMI Shrink Analysis Results Revision 1, 23 March 2012. This document contains Autodesk and third-party software license agreements/notices and/or additional terms and conditions
Autodesk Moldflow Insight 2012 AMI Shrink Analysis Results Revision 1, 23 March 2012. This document contains Autodesk and third-party software license agreements/notices and/or additional terms and conditions
Same Area, Different Perimeter; Same Perimeter, Different Area
 S E S S I O N 2. 5 A Same Area, Different Perimeter; Same Perimeter, Different Area Math Focus Points Using tiles to find the area and perimeter of a rectangle Understanding that rectangles can have the
S E S S I O N 2. 5 A Same Area, Different Perimeter; Same Perimeter, Different Area Math Focus Points Using tiles to find the area and perimeter of a rectangle Understanding that rectangles can have the
Analyzing Data Properties using Statistical Sampling Techniques
 Analyzing Data Properties using Statistical Sampling Techniques Illustrated on Scientific File Formats and Compression Features Julian M. Kunkel kunkel@dkrz.de 2016-06-21 Outline 1 Introduction 2 Exploring
Analyzing Data Properties using Statistical Sampling Techniques Illustrated on Scientific File Formats and Compression Features Julian M. Kunkel kunkel@dkrz.de 2016-06-21 Outline 1 Introduction 2 Exploring
Package EILA. February 19, Index 6. The CEU-CHD-YRI admixed simulation data
 Type Package Title Efficient Inference of Local Ancestry Version 0.1-2 Date 2013-09-09 Package EILA February 19, 2015 Author James J. Yang, Jia Li, Anne Buu, and L. Keoki Williams Maintainer James J. Yang
Type Package Title Efficient Inference of Local Ancestry Version 0.1-2 Date 2013-09-09 Package EILA February 19, 2015 Author James J. Yang, Jia Li, Anne Buu, and L. Keoki Williams Maintainer James J. Yang
