Two Dimensional Heteronuclear Correlation Spectroscopy
|
|
|
- Caren Hudson
- 6 years ago
- Views:
Transcription
1 Two Dimensional Heteronuclear Correlation Spectroscopy Gradient HMQC William D. Wheeler, Ph.D. Department of Chemistry University of Wyoming Revised September 7, 2006
2 2 INTRODUCTION Correlation Spectroscopy Correlation experiments reveal connections between spin coupled nuclei. Spin, or scalar, coupling between nuclei is established by the appearance of cross peaks (peaks off the diagonal) in the two-dimensional spectrum. Since spin coupled nuclei are usually separated by one to three bonds, the correlation experiment, by revealing this connectivity within the compound, is often enough to establish the chemical structure. Resonances which are coupled through four and five bonds will occasionally show cross peaks also. Although most correlation experiments are designed to emphasize short range coupling, there are experiments designed to emphasize long range correlations as well. Two-dimensional spectra can be acquired in either of two modes. In the magnitude, or absolute valve mode, only the real part of the data is collected since the final spectrum is not phased. In the phase sensitive mode, both the real and imaginary parts of the data are acquired so that the spectrum can be phased. An advantage of a phase sensitive spectrum is that the peaks are much narrower and the resolution greater. A disadvantage of a phase sensitive spectrum is that twice as much data must be recorded in order to extract the phase information. Thus, it takes twice the time to record a spectrum in phase sensitive mode. Gradient HMQC The gradient Heteronuclear Multiple Quantum Correlation experiment described here shows connections between carbon atoms and the protons directly bonded to them. The experiment also furnishes information about carbon types. Quaternary carbons, for instance, have no attached protons and consequently, show no correlations. Methine carbons, can show one and only one peak. Methylene carbons can show one or two peaks depending on the chemical shifts of the two protons and methyl groups generally show a single intense peak or a single multiplet. A two dimensional spectrum usually has quite low resolution so the fine structure due to multiplets is minimal. HMQC spectra can be acquired in either magnitude or phase sensitive mode. HMQC is known as an "inverse" experiment, because the 13 C chemical shift information is encoded into the 1 H signal. The data set for the example spectrum, however, was collected on a "direct" probe. In general, direct experiments detect the X nucleus, where the coil for the X channel is the closest to the sample. Inverse experiments detect the 1 H nucleus, where the coil for the 1 H channel is the closest to the sample. In this experiment, we are detecting the 1 H nucleus where the 1 H (decoupling) coil is NOT the closest coil to the sample. The S/N ratio is reduced, but the loss is not as great as switching to "direct" detection. This makes it possible to get 13 C- 1 H correlation data in a relatively short time without having to change to an inverse probe.
3 The gradient HMQC spectrum of Gramicidin S is shown on a following page. Gramicidin S is a cyclic deca-peptide consisting of the five amino acid pairs of Leucine, Valine, Proline, Ornithine (one L and one D), Phenylalanine (one L and one D). Notice, that the peaks between 7.6 and 9.6 ppm in the proton spectrum have no correlations to the carbon spectrum. These are amine protons. The peak at 3.3 ppm is water. The solvent (99% deuterated) shows a small sharp peak at 2.5 ppm in the proton spectrum which does correlate to the very large peak at 39.5 ppm in the carbon spectrum. The lowest contour level is above the top of this peak so that it does not appear in the spectrum as shown, but it is there. Other features include: The peak for the phenyl group of Phe at {7.2,128}; the five alpha C-H s between {4.0,50} and {5.0,60} ppm and the methyl groups of Val ane Leu near {0.8,18-23}. Also evident in this spectrum are several sets of methylene groups where the two protons are non-equivalent, thus showing two peaks with different 1 H chemical shifts, but a single 13 C chemical shift. The Gramicidin S spectrum required about 10 minutes to collect. 3 References. 1) Avance Users Guide, Bruker Instruments, Chapter 16. 2) 150 and More Basic NMR Experiments, A practical Course, S. Braun, H.-O. Kalinowski, S. Berger, Wiley-VCH, 1998, pages ) Basic One- and Two-Dimensional NMR Spectroscopy, Second, enlarged edition, Horst Friebolin, VCH Publishers, 1993, pages ) Modern NMR Spectroscopy, A Guide for Chemists, Jeremy K.M. Saunders and Brian K. Hunter, Oxford University Press, 1987, pages ) Biochemistry, Second Edition, Albert L. Lehninger, Worth Publishers, Inc., 1975, pages
4 4 Inverse heteronuclear 1 H - 13 C correlation of Gramicidin S
5 5 Inverse heteronuclear 1 H - 13 C correlation of Gramicidin S
6 6 EXPERIMENTAL SETUP The concentration of the sample should be about 50 mm to obtain results similar to those described here for Gramicidin S. This is about 16 mg of a sample of molecular weight 400 in 0.8 ml of solvent. If it is not possible to obtain this concentration, then the number of scans for the 2-D experiment may need to be increased. If you halve the concentration, you will need to quadruple the number of scans in order to achieve the same signal to noise ratio. Record a 1 H spectrum. NEW (or EDC) Create a new data set for your sample. NAME name Data set name. EXPNO 1 Experiment number (must be 1). PROCNO 1 Process data number (must be 1). [SAVE] Save the data set. RPAR +proton all Read in the standard proton parameters. NS, etc. Adjust NS and other parameters as necessary. RGA, ZG Acquire some data. FT, APK, ref Fourier transform, phase and reference the spectrum. Zoom Zoom in on the region of interest. The expanded region need not contain a reference peak. [sw-sfo1] Set the sweep width and spectrometer frequency to cover the zoomed region. Reduce TD if the acquisition time (AQ) is unnecessarily large. RGA, ZG Acquire a spectrum of the zoomed region. FT, etc. Fourier transform, phase etc. Record a 13 C spectrum. (Optional) You do not need a 13 C NMR spectrum to complete the HMQC. NEW (or EDC) Create a new data set for your sample. NAME name Data set name. EXPNO 13 Experiment number (suggest 13). PROCNO 1 Process data number (must be 1). [SAVE] Save the data set. If you have already collected a 13C spectrum, skip to the Zoom section to set up a spectrum of a zoomed region, and acquire that.
7 7 RPAR +carbon all NS, etc. RGA, ZG FT, APK, ref Zoom [sw-sfo1] RGA, ZG FT, etc. Read in the standard carbon parameters. Adjust NS and other parameters as necessary. Acquire some data. Fourier transform, phase and reference the spectrum. Zoom in on the region of interest. The expanded region need not contain a reference peak. Set the sweep width and spectrometer frequency to cover the zoomed region. Reduce TD if the acquisition time (AQ) is unnecessarily large. Acquire a spectrum of the zoomed region. Fourier transform, phase etc. Set up the 13 C- 1 H correlation experiment. The following AU program sets the parameters for gradient HMQC. setup2d Pick the [GRASP-HMQC] button and then press [Setup]. You should have experiment number 1 displayed (re 1) before executing setup2d. 16 The experiment number where the HMQC spectrum will be stored. The suggested number is 16. Zero and any number from 2 to 997 is valid. [OK] Answer Delete `meta.ext` files? with [OK]. 13 The experiment number where the 13 C spectrum is stored. Enter 0 (zero) if you did not collect a 13 C spectrum. Any number from 2 to 997 is valid. [Seen] [Default] [Save] Answer Error: Routing for channel 2 is not complete. with [Seen]. Press the [Default] button in the edasp window. This sets up the signal routing for the spectrometer. Press the [Save] button to close the edasp window.
8 8 The setup2d program suggests the following file parameters. File parameter EXPNO Experiment number 1, for 1-D proton. (Required) Experiment number 13, for 1-D carbon. (Suggested) Experiment number 16, for 2-D HMQC. (Suggested) The setup2d program sets the following acquisition parameters. Use EDA to display the results. Acquisition Parameters Time domain 2 (F2) PULPROG hmqcgpqf PULse PROGram for HMQC. TD 2048 Time Domain points. NS 2 Number of Scans (integer multiple of 1) DS 16 number of Dummy Scans for 1st row. D1 1.5 sec relaxation Delay. CNST One bond C-H coupling constant, J CH P homospoil / gradient pulse D delay for homospoil / gradient recovery GPZ Z gradient 1 GPZ Z gradient 2 GPZ Z gradient 3 Time domain 1 (F1) TD 256 Time Domain points. ND0 2 Number of D0 periods per cycle. SW 120 SW of 13 C spectrum collected in experiment 13 (120 if no spectrum). O1P 60 Center frequency (in ppm) of 13 C spectrum collected in experiment 13 (60 if no spectrum).
9 9 ASED EXPT [SAVE] Check that acquisition parameters are set correctly. A brief description of other parameters, not described above, is given in the pulse program at the end of this document. Save the acquisition parameters. Calculates the approximate length of time to do the experiment. This may help you to decide if you want to collect more or fewer slices or scans or points etc. The Gramicidin spectrum required about 10 minutes. DATA ACQUISITION [Spin on/off] RGA ZG TURN THE SPINNER OFF ( press the Spin On/Off button on the BOSS keyboard). Set the receiver gain. This will take a few seconds. Start the acquisition.
10 10 DATA PROCESSING The setup2d program sets the following processing parameters. Use EDP to check or modify the values. Processing Parameters Time/frequency domain 2 (F2) SI 1024 the SIze in F2 (zero fill rows). WDW QSINE Sine squared multiplication. SSB 2 90 B shifted sine squared bell. PH_mod NO No phase correction. PKNL TRUE Required with digital filter. BC_mod NO Background correct quadrature data. SF Spectrum reference frequency for 1 H. Time/frequency domain 1 (F1) SI 512 the SIze in F1 (zero fill columns). WDW QSINE Sine multiplication. (Or EM with LB 2-5) SSB 2 90 B shifted sine bell. PH_mod MC Magnitude calculation. BC_mod NO No background correction. MC2 QF Forward quadrature FFT. SF Spectrum reference frequency for 13 C. OFFSET 120 Offset (in ppm) of 13 C spectrum collected in experiment 13 (120 if no spectrum). XFB Background correct, window, zero fill, Fourier transform, phase and reference. The whole enchilada in one command. It is OK to execute this command on a partial data set, while acquisition is in progress.
11 11 X-Y Expansion the spectrum. 2-D CONTOUR DISPLAY [Limits] [PlotReg] Set the limits of the plot region. Forces XWinNMR to display the plot region. Setting the intensity scale. [DefPlot] [contours] [intensities] Sets the intensity scale for plotting to be the same as the currently displayed intensity scale. Displays the equal-intensity contours of the data. Displays ranges of intensity as a color map. PLOTTING SETTI Set the title for the 2D spectrum. The setup2d program sets the following plotting parameters. Use xwinplot to plot the spectrum. The parameter names are not visible in xwinplot, but you should get the idea. If you did not collect a carbon spectrum, you will need to either remove the F1 projection area, or create a projection to use in the F1 projection area. Plotting Parameters Projection for Frequency domain 2 PF2DU /opt/xwinnmr Disk Unit of the data set. PF2USER username Your user name. PF2NAME name Name of the data set. PF2EXP 1 Experiment number of the 1 H spectrum. PF2PROC 1 Process number of the 1 H spectrum. Projection for Frequency domain 1 PF1DU /opt/xwinnmr Disk Unit of the data set. PF1USER username Your user name. PF1NAME name Name of the data set. PF1EXP 13 Experiment number of the 13 C spectrum.
12 12 Plotting Parameters PF1PROC 1 Process number of the 13 C spectrum. Additional plotting parameters PF1CY 2.5 The value of CY for the F1 projection. This can be used to increase or decrease the height of the spectrum displayed in the F1 projection. (PF1CY = 5.0 will double the height of the 1-D spectrum). PF2CY 2.5 The value of CY for the F2 projection. Additional Processing Parameters Frequency domain F1LO bbb Set to the value for the bottom of the spectrum (ppm). F1HI ttt Set to the value for the top of the spectrum (ppm). F2LO lll Set to the value for the left side of the spectrum (ppm). F2HI rrr Set to the value for the right side of the spectrum (ppm). Additional Processing Commands Frequency domain ABS1 ABS2 Automatic Baseline Subtraction for F1 (uses F1LO, F1HI). Automatic Baseline Subtraction for F2 (uses F2LO, F2HI).
13 13 Pulse Program for gradient HMQC ;hmqcgpqf ;avance-version (02/05/31) ;HMQC ;2D H-1/X correlation via heteronuclear zero and double quantum ; coherence ;with decoupling during acquisition ;using gradient pulses for selection ;use pulseprogram 'inv4gpnd1d' for setup #include <Avance.incl> #include <Grad.incl> #include <Delay.incl> "p2=p1*2" "d0=3u" "d2=1s/(cnst2*2)" "d12=20u" "d13=4u" "DELTA1=d2-p16-d16-d13-d12" 1 ze 2 d1 do:f2 3 p1 ph1 d2 pl2:f2 UNBLKGRAD p3:f2 ph3 d0 p16:gp1 d16 p2 ph2 d13 p16:gp2 d16 d0 p3:f2 ph4 d13 p16:gp3 d16 DELTA1 d12 pl12:f2 BLKGRAD go=2 ph31 cpd2:f2 d1 do:f2 mc #0 to 2 F1QF(id0) exit
14 14 ph1=0 ph2=0 ph3=0 2 ph4= ph31= ;pl1 : f1 channel - power level for pulse (default) ;pl2 : f2 channel - power level for pulse (default) ;pl12: f2 channel - power level for CPD/BB decoupling ;p1 : f1 channel - 90 degree high power pulse ;p2 : f1 channel degree high power pulse ;p3 : f2 channel - 90 degree high power pulse ;p16: homospoil/gradient pulse ;d0 : incremented delay (2D) [3 usec] ;d1 : relaxation delay; 1-5 * T1 ;d2 : 1/(2J)XH ;d12: delay for power switching [20 usec] ;d13: short delay [4 usec] ;d16: delay for homospoil/gradient recovery ;cnst2: = J(XH) ;in0: 1/(2 * SW(X)) = DW(X) ;nd0: 2 ;NS: 1 * n ;DS: 16 ;td1: number of experiments ;FnMODE: QF ;cpd2: decoupling according to sequence defined by cpdprg2 ;pcpd2: f2 channel - 90 degree pulse for decoupling sequence ;use gradient ratio: gp 1 : gp 2 : gp 3 ; 50 : 30 : 40.1 for C-13 ; 70 : 30 : 50.1 for N-15 ;for z-only gradients: ;gpz1: 50% for C-13, 70% for N-15 ;gpz2: 30% ;gpz3: 40.1% for C-13, 50.1% for N-15 ;use gradient files: ;gpnam1: SINE.100 ;gpnam2: SINE.100 ;gpnam3: SINE.100 ;$Id: hmqcgpqf,v /06/12 09:04:42 ber Exp $
Two Dimensional Homonuclear Correlation Spectroscopy
 Two Dimensional Homonuclear Correlation Spectroscopy Gradient COSY William D. Wheeler, Ph.D. Department of Chemistry University of Wyoming April 16, 1999 Revised September 22, 1999 2 INTRODUCTION Correlation
Two Dimensional Homonuclear Correlation Spectroscopy Gradient COSY William D. Wheeler, Ph.D. Department of Chemistry University of Wyoming April 16, 1999 Revised September 22, 1999 2 INTRODUCTION Correlation
Two Dimensional Homonuclear Correlation Spectroscopy
 Two Dimensional Homonuclear Correlation Spectroscopy DQF-COSY William D. Wheeler, Ph.D. Department of Chemistry University of Wyoming September 23, 1999 2 INTRODUCTION Correlation Spectroscopy Correlation
Two Dimensional Homonuclear Correlation Spectroscopy DQF-COSY William D. Wheeler, Ph.D. Department of Chemistry University of Wyoming September 23, 1999 2 INTRODUCTION Correlation Spectroscopy Correlation
HMBC 17. Goto. Introduction AVANCE User s Guide Bruker 185
 Chapter HMBC 17 Introduction 17.1 Goto Heteronuclear Multiple Bond Correlation spectroscopy is a modified version of HMQC suitable for determining long-range 1 H- 13 C connectivity. This is useful in determining
Chapter HMBC 17 Introduction 17.1 Goto Heteronuclear Multiple Bond Correlation spectroscopy is a modified version of HMQC suitable for determining long-range 1 H- 13 C connectivity. This is useful in determining
CLIP-HSQMBC: Easy measurement of small proton-carbon coupling constants in organic molecules
 CLIP-HSQMBC: Easy measurement of small proton-carbon coupling constants in organic molecules Josep Saurí, a Teodor Parella a and Juan F. Espinosa* b Supporting information 1. Pulse sequence code (Bruker)
CLIP-HSQMBC: Easy measurement of small proton-carbon coupling constants in organic molecules Josep Saurí, a Teodor Parella a and Juan F. Espinosa* b Supporting information 1. Pulse sequence code (Bruker)
8 COSY. 8.1 Introduction. 8.2 Magnitude COSY
 8 COSY 8.1 Introduction COSY (COrrelation SpectroscopY) is a homonuclear 2D technique that is used to correlate the chemical shifts of 1 H nuclei which are J-coupled to one another. In this chapter, two
8 COSY 8.1 Introduction COSY (COrrelation SpectroscopY) is a homonuclear 2D technique that is used to correlate the chemical shifts of 1 H nuclei which are J-coupled to one another. In this chapter, two
Chem 203 December 15, Final Exam Part II Problem 3 of 3 (30 points)
 Name: Chem 203 December 15, 2012 Final Exam Part II Problem 3 of 3 (30 points) Select and submit TWO OUT OF THE THREE PROBLEMS FROM PART II for grading. Do not submit three problems. If you wish to unstaple
Name: Chem 203 December 15, 2012 Final Exam Part II Problem 3 of 3 (30 points) Select and submit TWO OUT OF THE THREE PROBLEMS FROM PART II for grading. Do not submit three problems. If you wish to unstaple
Chem 203 December 15, Final Exam Part II Problem 2 of 3 (30 points)
 Name: Chem 203 December 15, 2012 Final Exam Part II Problem 2 of 3 (30 points) Select and submit TWO OUT OF THE THREE PROBLEMS FROM PART II for grading. Do not submit three problems. If you wish to unstaple
Name: Chem 203 December 15, 2012 Final Exam Part II Problem 2 of 3 (30 points) Select and submit TWO OUT OF THE THREE PROBLEMS FROM PART II for grading. Do not submit three problems. If you wish to unstaple
Ultrahigh-resolution Total Correlation NMR Spectroscopy
 Ultrahigh-resolution Total Correlation NMR Spectroscopy Supporting Information Mohammadali Foroozandeh, Ralph W. Adams, Mathias Nilsson and Gareth A. Morris* All experimental spectra were recorded at a
Ultrahigh-resolution Total Correlation NMR Spectroscopy Supporting Information Mohammadali Foroozandeh, Ralph W. Adams, Mathias Nilsson and Gareth A. Morris* All experimental spectra were recorded at a
User manual Bruker DPX200 NMR spectrometer
 User manual Bruker DPX200 NMR spectrometer Insert the NMR tube in the spinner in such a way that the bottom of the tube reaches the grey disc at the bottom of the spinnerholder. Make sure that the NMR
User manual Bruker DPX200 NMR spectrometer Insert the NMR tube in the spinner in such a way that the bottom of the tube reaches the grey disc at the bottom of the spinnerholder. Make sure that the NMR
Implementing ultrafast 2D NMR experiments on a Bruker Avance Spectrometer
 Implementing ultrafast 2D NMR experiments on a Bruker Avance Spectrometer Laetitia Rouger, Benoît Charrier, Serge Akoka, Patrick Giraudeau EBSI group CEISAM laboratory http://www.sciences.univ-nantes.fr/ceisam/en_ebsi1.php
Implementing ultrafast 2D NMR experiments on a Bruker Avance Spectrometer Laetitia Rouger, Benoît Charrier, Serge Akoka, Patrick Giraudeau EBSI group CEISAM laboratory http://www.sciences.univ-nantes.fr/ceisam/en_ebsi1.php
Chem 203. Organic Spectroscopy. Midterm Examination, Part II (60 points total) Problem 4 of 4 (three out of four required, 20 points)
 NAME Chem 203 Organic Spectroscopy Midterm Examination, Part II (60 points total) Problem 4 of 4 (three out of four required, 20 points) Saturday, November 9, 2013, 9 am -??? SUBMIT THREE OF THE FOUR PROBLEMS
NAME Chem 203 Organic Spectroscopy Midterm Examination, Part II (60 points total) Problem 4 of 4 (three out of four required, 20 points) Saturday, November 9, 2013, 9 am -??? SUBMIT THREE OF THE FOUR PROBLEMS
Chem 203. Organic Spectroscopy. Midterm Examination, Part II (60 points total) Problem 1 of 4 (three out of four required, 20 points)
 NAME Chem 203 Organic Spectroscopy Midterm Examination, Part II (60 points total) Problem 1 of 4 (three out of four required, 20 points) Saturday, November 15, 2014, 9 am -??? SUBMIT THREE OF THE FOUR
NAME Chem 203 Organic Spectroscopy Midterm Examination, Part II (60 points total) Problem 1 of 4 (three out of four required, 20 points) Saturday, November 15, 2014, 9 am -??? SUBMIT THREE OF THE FOUR
Chem 203. Organic Spectroscopy. Midterm Examination, Part II (60 points total) Problem 4 of 4 (three out of four required, 20 points)
 NAME Chem 203 Organic Spectroscopy Midterm Examination, Part II (60 points total) Problem 4 of 4 (three out of four required, 20 points) Saturday, November 15, 2014, 9 am -??? SUBMIT THREE OF THE FOUR
NAME Chem 203 Organic Spectroscopy Midterm Examination, Part II (60 points total) Problem 4 of 4 (three out of four required, 20 points) Saturday, November 15, 2014, 9 am -??? SUBMIT THREE OF THE FOUR
400 MHz spectrometer user manual
 400 MHz spectrometer user manual january 2017 Sandrine Denis-Quanquin 1. THE NMR SPECTROMETER... 3 2. MANUAL MODE / AUTOMATION... 4 2.1 SAMPLE CHANGER... 4 2.2 MANUAL MODE... 4 2.3 AUTOMATION... 4 3. PRELIMINARY
400 MHz spectrometer user manual january 2017 Sandrine Denis-Quanquin 1. THE NMR SPECTROMETER... 3 2. MANUAL MODE / AUTOMATION... 4 2.1 SAMPLE CHANGER... 4 2.2 MANUAL MODE... 4 2.3 AUTOMATION... 4 3. PRELIMINARY
NMR Spectrometer Operation: xwinnmr
 NMR Spectrometer Operation: xwinnmr Dr. Robert Peterson Facility Manager NMR Technology Center UCLA-DOE Institute for Genomics and Proteomics UCLA Dept. of Chemistry and Biochemistry Overview This is a
NMR Spectrometer Operation: xwinnmr Dr. Robert Peterson Facility Manager NMR Technology Center UCLA-DOE Institute for Genomics and Proteomics UCLA Dept. of Chemistry and Biochemistry Overview This is a
Relaxation-encoded NMR experiments for mixture analysis: REST and beer. Electronic Supporting Information
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2017 Relaxation-encoded NMR experiments for mixture analysis: REST and beer Electronic Supporting Information
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2017 Relaxation-encoded NMR experiments for mixture analysis: REST and beer Electronic Supporting Information
Step by step procedure for NMR data acquisition
 Step by step procedure for NMR data acquisition Spectrometers The UTHSCSA 500, 600, and 700 MHz spectrometers are each equipped with 4 independent RF channels and are each operated by a Red Hat Linux workstation
Step by step procedure for NMR data acquisition Spectrometers The UTHSCSA 500, 600, and 700 MHz spectrometers are each equipped with 4 independent RF channels and are each operated by a Red Hat Linux workstation
Program. short intro the hardware locking and shimming 1D proton setup presaturation spectra
 Program short intro the hardware locking and shimming 1D proton setup presaturation spectra 13 C spectra, DEPT processing of 1D file transfer and backup 2D homonuclear + 2D processing 2D heteronuclear
Program short intro the hardware locking and shimming 1D proton setup presaturation spectra 13 C spectra, DEPT processing of 1D file transfer and backup 2D homonuclear + 2D processing 2D heteronuclear
KJM D-SELECTIVE NMR Experiments on the AVI-600 and AVII-600. Version 1.0. Topspin 3.5 Windows 7 Topspin 1.3 Windows XP
 KJM 9250 1D-SELECTIVE NMR Experiments on the AVI-600 and AVII-600 Version 1.0 Topspin 3.5 Windows 7 Topspin 1.3 Windows XP Professor Emeritus Alistair Lawrence Wilkins, University of Waikato, New Zealand.
KJM 9250 1D-SELECTIVE NMR Experiments on the AVI-600 and AVII-600 Version 1.0 Topspin 3.5 Windows 7 Topspin 1.3 Windows XP Professor Emeritus Alistair Lawrence Wilkins, University of Waikato, New Zealand.
H Micro-Imaging. Tuning and Matching. i. Open any 1H data set and type wobb.
 - 1-1 H Micro-Imaging The NMR-specific properties of the objects are visualized as multidimensional images. Translational motion can be observed and spectroscopic information can be spatially resolved.
- 1-1 H Micro-Imaging The NMR-specific properties of the objects are visualized as multidimensional images. Translational motion can be observed and spectroscopic information can be spatially resolved.
2D heteronuclear correlation experiments
 2D heteronuclear correlation experiments Assistant Professor Kenneth Kongstad Bioanalytical Chemistry and Metabolomics Research Group Section for Natural Products and Peptides Department of Drug Design
2D heteronuclear correlation experiments Assistant Professor Kenneth Kongstad Bioanalytical Chemistry and Metabolomics Research Group Section for Natural Products and Peptides Department of Drug Design
KJM D-SELECTIVE NMR Experiments on the AVIIIHD-800. Version 1.0. Topspin 3.5 Windows 7
 KJM 9250 1D-SELECTIVE NMR Experiments on the AVIIIHD-800 Version 1.0 Topspin 3.5 Windows 7 Professor Emeritus Alistair Lawrence Wilkins, University of Waikato, New Zealand. January 2018 1D-SELECTIVE NMR
KJM 9250 1D-SELECTIVE NMR Experiments on the AVIIIHD-800 Version 1.0 Topspin 3.5 Windows 7 Professor Emeritus Alistair Lawrence Wilkins, University of Waikato, New Zealand. January 2018 1D-SELECTIVE NMR
SUPPORTING INFORMATION
 Eur. J. Org. Chem. 2008 WILEY-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2008 ISSN 1434 193X SUPPORTING INFORMATION Title: Structural Elucidation with NMR Spectroscopy: Practical Strategies for Organic
Eur. J. Org. Chem. 2008 WILEY-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2008 ISSN 1434 193X SUPPORTING INFORMATION Title: Structural Elucidation with NMR Spectroscopy: Practical Strategies for Organic
NMR Hardware 06/06/2017. Outline. Instrumentation: Magnet. Increasing magnetic field increases Sensitivity, by power of 3/2 Dispersion, linearly
 NMR Hardware Outline Magnet Lock Shims Gradient Probe Signal generation and transmitters Receiver and digitizer Variable temperature system Solids hardware Instrumentation: Magnet Often the most impressive
NMR Hardware Outline Magnet Lock Shims Gradient Probe Signal generation and transmitters Receiver and digitizer Variable temperature system Solids hardware Instrumentation: Magnet Often the most impressive
Fast Methods for Small Molecules
 Fast Methods for Small Molecules Technical Overview Throughput is a key concern in many NMR laboratories, and using faster methods is one way to increase it. Traditionally, multidimensional NMR requires
Fast Methods for Small Molecules Technical Overview Throughput is a key concern in many NMR laboratories, and using faster methods is one way to increase it. Traditionally, multidimensional NMR requires
Laboratory Experiments for Nuclear Magnetic Resonance Spectroscopy May 6, 2004
 CONTENTS 1 Contents Laboratory Experiments for Nuclear Magnetic Resonance Spectroscopy May 6, 2004 1 Introduction 3 2 Safety 3 2.1 High Magnetic Fields......................................... 3 2.1.1
CONTENTS 1 Contents Laboratory Experiments for Nuclear Magnetic Resonance Spectroscopy May 6, 2004 1 Introduction 3 2 Safety 3 2.1 High Magnetic Fields......................................... 3 2.1.1
Exercise at the Spectrometer
 Exercise at the Spectrometer General The script is written for Bruker spectrometers. The color code means, red for all commands and green for parameters in the TopSpin commando line. The script gives additional
Exercise at the Spectrometer General The script is written for Bruker spectrometers. The color code means, red for all commands and green for parameters in the TopSpin commando line. The script gives additional
Arenium Acid-Catalyzed Deuteration of Aromatic Hydrocarbons
 Supporting Information for Arenium Acid-Catalyzed euteration of Aromatic Hydrocarbons Simon uttwyler 1, Anna Butterfield 2, and Jay S. Siegel 2 * 1 epartment of Chemistry, Yale University, 350 Edwards
Supporting Information for Arenium Acid-Catalyzed euteration of Aromatic Hydrocarbons Simon uttwyler 1, Anna Butterfield 2, and Jay S. Siegel 2 * 1 epartment of Chemistry, Yale University, 350 Edwards
Open acqi window if the button has been lost. autolocking routine, alock= y for autolocking, alock= n for typical manual locking
 Glossary of Common NMR Commands and Terms aa acqi ai alock aph array at points (np) axis='p' axis= pd BPsvf bc bs cd directory abort acquisition, hard stop Open acqi window if the button has been lost
Glossary of Common NMR Commands and Terms aa acqi ai alock aph array at points (np) axis='p' axis= pd BPsvf bc bs cd directory abort acquisition, hard stop Open acqi window if the button has been lost
γ-trifluoromethyl proline: Evaluation as a structural substitute of proline for solid state 19 F-NMR peptide studies
 Electronic Supplementary Material (ESI) for Organic & Biomolecular Chemistry. This journal is The Royal Society of Chemistry 2015 γ-trifluoromethyl proline: Evaluation as a structural substitute of proline
Electronic Supplementary Material (ESI) for Organic & Biomolecular Chemistry. This journal is The Royal Society of Chemistry 2015 γ-trifluoromethyl proline: Evaluation as a structural substitute of proline
MT-540. D-Ribavirin, [ 3 H]- Lot A DG
![MT-540. D-Ribavirin, [ 3 H]- Lot A DG MT-540. D-Ribavirin, [ 3 H]- Lot A DG](/thumbs/85/92603848.jpg) MT-54 D-Ribavirin, [ 3 H]- Lot 165-145-225-A-2835-DG A) All chromatograms were run using the HPLC method described on the Product Data Sheet. Concentrations and volumes: Standard solution concentration
MT-54 D-Ribavirin, [ 3 H]- Lot 165-145-225-A-2835-DG A) All chromatograms were run using the HPLC method described on the Product Data Sheet. Concentrations and volumes: Standard solution concentration
1D Transient NOE on the Bruker DRX-500 and DRX-600
 1D Transient NOE on the Bruker DRX-500 and DRX-600 Reference: Stott, K., Stonehouse, J., Keeler, T.L. and Shaka, A.J., J. Amer. Chem. Soc. 1995, 117 (14), pp. 4199-4200. At thermal equilibrium in a strong
1D Transient NOE on the Bruker DRX-500 and DRX-600 Reference: Stott, K., Stonehouse, J., Keeler, T.L. and Shaka, A.J., J. Amer. Chem. Soc. 1995, 117 (14), pp. 4199-4200. At thermal equilibrium in a strong
MT Gemcitabine, [5-3 H(N)]- Lot A NK
![MT Gemcitabine, [5-3 H(N)]- Lot A NK MT Gemcitabine, [5-3 H(N)]- Lot A NK](/thumbs/93/112954466.jpg) MT-1572 Gemcitabine, [5-3 H(N)]- Lot 194-14-234-A-2514-NK A) All chromatograms were run using the HPLC method described on the Product Data Sheet. Concentrations and volumes: Standard solution concentration
MT-1572 Gemcitabine, [5-3 H(N)]- Lot 194-14-234-A-2514-NK A) All chromatograms were run using the HPLC method described on the Product Data Sheet. Concentrations and volumes: Standard solution concentration
Development of an Enantioselective Route towards the. Lycopodium Alkaloids: Total Synthesis of Lycopodine.
 Development of an Enantioselective Route towards the Lycopodium Alkaloids: Total Synthesis of Lycopodine. Hua Yang and Rich G. Carter* Department of Chemistry, Oregon State University, Corvallis, OR 97331.
Development of an Enantioselective Route towards the Lycopodium Alkaloids: Total Synthesis of Lycopodine. Hua Yang and Rich G. Carter* Department of Chemistry, Oregon State University, Corvallis, OR 97331.
PINMRF. Varian 300 MHz NMR Spectrometers User Guide for Advanced 1D and Basic 2D NMR Experiments
 PINMRF Varian 300 MHz NMR Spectrometers User Guide for Advanced 1D and Basic 2D NMR Experiments INCLUDING: Inova-300-1 w/ 5mm 4-nucleus probe 365 WTHR Inova-300-2 w/ 5mm 4-nucleus probe 4100 BRWN Table
PINMRF Varian 300 MHz NMR Spectrometers User Guide for Advanced 1D and Basic 2D NMR Experiments INCLUDING: Inova-300-1 w/ 5mm 4-nucleus probe 365 WTHR Inova-300-2 w/ 5mm 4-nucleus probe 4100 BRWN Table
KJM Version 1.0. Topspin 3.5 Windows 7 Topspin 1.3 Windows XP
 KJM 9250 1 H NMR spectra on the AVI-600 and AVII-600 Version 1.0 Topspin 3.5 Windows 7 Topspin 1.3 Windows XP Professor Emeritus Alistair Lawrence Wilkins, University of Waikato, New Zealand. January 2018
KJM 9250 1 H NMR spectra on the AVI-600 and AVII-600 Version 1.0 Topspin 3.5 Windows 7 Topspin 1.3 Windows XP Professor Emeritus Alistair Lawrence Wilkins, University of Waikato, New Zealand. January 2018
NMR Spectroscopy with Radio Frequency Gradients.
 RF GRASP TM NMR Spectroscopy with Radio Frequency Gradients. BRUKER Werner E. Maas Bruker Instruments, Inc. 19 Fortune Drive Billerica, MA 01821 USA version 1.2 February, 1996 Copyright 1996 Bruker Instruments,
RF GRASP TM NMR Spectroscopy with Radio Frequency Gradients. BRUKER Werner E. Maas Bruker Instruments, Inc. 19 Fortune Drive Billerica, MA 01821 USA version 1.2 February, 1996 Copyright 1996 Bruker Instruments,
Instruction for Operating the Bruker Avance III 800 MHz NMR Spectrometers in UTMB
 Instruction for Operating the Bruker Avance III 800 MHz NMR Spectrometers in UTMB Written by Tianzhi Wang, date: February 8, 2013. No food, no drink in NMR room and no internet in NMR host computer except
Instruction for Operating the Bruker Avance III 800 MHz NMR Spectrometers in UTMB Written by Tianzhi Wang, date: February 8, 2013. No food, no drink in NMR room and no internet in NMR host computer except
Eclipse+ NMR Training Guide
 Eclipse+ NMR Training Guide Version 4.3.4 ECLIPSE+ NMR Training Guide Revision 20050617 Copyright 2005 by JEOL USA, Inc. Analytical Instruments Division 11 Dearborn Road Peabody, MA 01960 (978) 535-5900
Eclipse+ NMR Training Guide Version 4.3.4 ECLIPSE+ NMR Training Guide Revision 20050617 Copyright 2005 by JEOL USA, Inc. Analytical Instruments Division 11 Dearborn Road Peabody, MA 01960 (978) 535-5900
NMR Basics. Lecture 2
 NMR Basics Lecture 2 Continuous wave (CW) vs. FT NMR There are two ways of tuning a piano: - key by key and recording each sound (or frequency). - or, kind of brutal, is to hit with a sledgehammer and
NMR Basics Lecture 2 Continuous wave (CW) vs. FT NMR There are two ways of tuning a piano: - key by key and recording each sound (or frequency). - or, kind of brutal, is to hit with a sledgehammer and
Acceptance Solid State NMR Test Procedure. for Avance NMR Systems
 Acceptance Solid State NMR Test Procedure for Avance NMR Systems Manual P/N B92999 Acceptance Solid State NMR Test Procedures for AVANCE systems Index: 04 Number: B92999 Page: 1 (42) 1. Purpose 2. Area
Acceptance Solid State NMR Test Procedure for Avance NMR Systems Manual P/N B92999 Acceptance Solid State NMR Test Procedures for AVANCE systems Index: 04 Number: B92999 Page: 1 (42) 1. Purpose 2. Area
10. Phase Cycling and Pulsed Field Gradients Introduction to Phase Cycling - Quadrature images
 10. Phase Cycling and Pulsed Field Gradients 10.1 Introduction to Phase Cycling - Quadrature images The selection of coherence transfer pathways (CTP) by phase cycling or PFGs is the tool that allows the
10. Phase Cycling and Pulsed Field Gradients 10.1 Introduction to Phase Cycling - Quadrature images The selection of coherence transfer pathways (CTP) by phase cycling or PFGs is the tool that allows the
Student Name: Date Completed: Supervisor:
 2 NMR Training for the 600 MHz NMR with Chempack INOVA 600 Tests and Assignment Certification Student Name: 600-Test #1: The student will be given a written test administered by Dr. Lee. This test will
2 NMR Training for the 600 MHz NMR with Chempack INOVA 600 Tests and Assignment Certification Student Name: 600-Test #1: The student will be given a written test administered by Dr. Lee. This test will
NMR spectrometer usage at the BioNMR facility ETH Zürich
 NMR spectrometer usage at the BioNMR facility ETH Zürich Accounts 2 Safety precautions: strong magnetic fields 2 Parts of an NMR spectrometer 3 NMR data storage 3 Start topspin software 3 Initial steps
NMR spectrometer usage at the BioNMR facility ETH Zürich Accounts 2 Safety precautions: strong magnetic fields 2 Parts of an NMR spectrometer 3 NMR data storage 3 Start topspin software 3 Initial steps
Your first NMR measurement
 Your first NMR measurement Introduction Select 10mM water in D2O as NMR sample. The NMR spectrum of such sample consists of only two signals: the water signal and the peak of the reference (TSP). Follow
Your first NMR measurement Introduction Select 10mM water in D2O as NMR sample. The NMR spectrum of such sample consists of only two signals: the water signal and the peak of the reference (TSP). Follow
TopSpin Guide Book. Basic NMR Experiments User Manual. Innovation with Integrity. Version 002 NMR
 TopSpin Guide Book Basic NMR Experiments User Manual Version 002 Innovation with Integrity NMR Copyright by Bruker Corporation All rights reserved. No part of this publication may be reproduced, stored
TopSpin Guide Book Basic NMR Experiments User Manual Version 002 Innovation with Integrity NMR Copyright by Bruker Corporation All rights reserved. No part of this publication may be reproduced, stored
Instructions for 1 H-, 13 C-, 19 F-, and 31 P-Spectra on the Varian Mercury-Vx-300
 1 Instructions for 1 H-, 13 C-, 19 F-, and 31 P-Spectra on the Varian Mercury-Vx-300 Please note: Under no circumstances move the magnet or the automatic sampler table. Do not attempt to use the auto sampler
1 Instructions for 1 H-, 13 C-, 19 F-, and 31 P-Spectra on the Varian Mercury-Vx-300 Please note: Under no circumstances move the magnet or the automatic sampler table. Do not attempt to use the auto sampler
Principios Básicos de RMN en sólidos destinado a usuarios. Gustavo Monti. Fa.M.A.F. Universidad Nacional de Córdoba Argentina
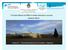 Principios Básicos de RMN en sólidos destinado a usuarios Gustavo Monti Fa.M.A.F. Universidad Nacional de Córdoba Argentina magnet 1 2 4 5 6 computer 3 Block diagrama of a traditional NMR spectrometer.
Principios Básicos de RMN en sólidos destinado a usuarios Gustavo Monti Fa.M.A.F. Universidad Nacional de Córdoba Argentina magnet 1 2 4 5 6 computer 3 Block diagrama of a traditional NMR spectrometer.
Arrayed Acquisition in VNMR and on the Gemini 1
 Arrayed Acquisition in VNMR and on the Gemini 1 I. Arrays in VNMR Vnmr allows you to quickly define experiments in which a series of spectra can be obtained as a function of any NMR parameter. For example,
Arrayed Acquisition in VNMR and on the Gemini 1 I. Arrays in VNMR Vnmr allows you to quickly define experiments in which a series of spectra can be obtained as a function of any NMR parameter. For example,
If the magnetic field is larger, more energy is required to excite a given nucleus.
 1 2 If an NMR-active nucleus such as 1 H or 13 C is put into a magnet field, then it will come into resonance if it is irradiated with rf at the correct frequency. The correct frequency depends mainly
1 2 If an NMR-active nucleus such as 1 H or 13 C is put into a magnet field, then it will come into resonance if it is irradiated with rf at the correct frequency. The correct frequency depends mainly
Efficient Palladium-catalyzed Coupling Reactions of Aryl Bromides and Chlorides with Phenols
 Efficient Palladium-catalyzed Coupling Reactions of Aryl Bromides and Chlorides with Phenols Tongjie Hu, a Thomas Schulz, b Christian Torborg, b Xiaorong Chen, a Jun Wang, a Matthias Beller b* and Jun
Efficient Palladium-catalyzed Coupling Reactions of Aryl Bromides and Chlorides with Phenols Tongjie Hu, a Thomas Schulz, b Christian Torborg, b Xiaorong Chen, a Jun Wang, a Matthias Beller b* and Jun
An NMR Caveman s Guide to Quickly Acquiring Spectroscopic Data By Brian Sparling
 An NMR Caveman s Guide to Quickly Acquiring Spectroscopic Data By Brian Sparling Disclaimer: this guide is meant to be a quick, routine means of obtaining characterization data for unknown compounds in
An NMR Caveman s Guide to Quickly Acquiring Spectroscopic Data By Brian Sparling Disclaimer: this guide is meant to be a quick, routine means of obtaining characterization data for unknown compounds in
Workshop on Rapid Scan EPR. University of Denver EPR Center and Bruker BioSpin July 28, 2013
 Workshop on Rapid Scan EPR University of Denver EPR Center and Bruker BioSpin July 28, 2013 Direct detection Direct detected magnetic resonance that is, without modulation and phase-sensitive detection
Workshop on Rapid Scan EPR University of Denver EPR Center and Bruker BioSpin July 28, 2013 Direct detection Direct detected magnetic resonance that is, without modulation and phase-sensitive detection
Gradients. Effects of B0 gradients on transverse magnetisation Similar to figure 10 of Sattler review Progr. NMR 34 (1999), 93
 Gradients 1. What are gradients? Modern high-resolution NMR probes contain -besides the RF coils - additional coils that can be fed a DC current. The coils are built so that a pulse (~1 ms long) of DC
Gradients 1. What are gradients? Modern high-resolution NMR probes contain -besides the RF coils - additional coils that can be fed a DC current. The coils are built so that a pulse (~1 ms long) of DC
NMR FACILITY NEWSLETTER
 NMR FACILITY NEWSLETTER Department of Chemistry and Biochemistry Matt Revington-Facility Coordinator mrevingt@uwindsor.ca Ext 3997 Workshop Announcement : Advanced Topics in NMR There will be an Advanced
NMR FACILITY NEWSLETTER Department of Chemistry and Biochemistry Matt Revington-Facility Coordinator mrevingt@uwindsor.ca Ext 3997 Workshop Announcement : Advanced Topics in NMR There will be an Advanced
PULSED/CW NUCLEAR MAGNETIC RESONANCE
 PULSED/CW NUCLEAR MAGNETIC RESONANCE The Second Generation of TeachSpin s Classic Explore NMR for both Hydrogen (at 21 MHz) and Fluorine Nuclei Magnetic Field Stabilized to 1 part in 2 million Homogenize
PULSED/CW NUCLEAR MAGNETIC RESONANCE The Second Generation of TeachSpin s Classic Explore NMR for both Hydrogen (at 21 MHz) and Fluorine Nuclei Magnetic Field Stabilized to 1 part in 2 million Homogenize
Reference Manual SPECTRUM. Signal Processing for Experimental Chemistry Teaching and Research / University of Maryland
 Reference Manual SPECTRUM Signal Processing for Experimental Chemistry Teaching and Research / University of Maryland Version 1.1, Dec, 1990. 1988, 1989 T. C. O Haver The File Menu New Generates synthetic
Reference Manual SPECTRUM Signal Processing for Experimental Chemistry Teaching and Research / University of Maryland Version 1.1, Dec, 1990. 1988, 1989 T. C. O Haver The File Menu New Generates synthetic
Chapter 1. 1 The NMR Spectrometer. 1.1 Components of an NMR Spectrometer The Magnet
 Chapter 1 1 The NMR Spectrometer 1.1 Components of an NMR Spectrometer 1.1.1 The Magnet In most current NMR spectrometers the magnetic field is generated by a superconducting magnet (Fig. 1.1). The first
Chapter 1 1 The NMR Spectrometer 1.1 Components of an NMR Spectrometer 1.1.1 The Magnet In most current NMR spectrometers the magnetic field is generated by a superconducting magnet (Fig. 1.1). The first
2015 Spin echoes and projection imaging
 1. Spin Echoes 1.1 Find f0, transmit amplitudes, and shim settings In order to acquire spin echoes, we first need to find the appropriate scanner settings using the FID GUI. This was all done last week,
1. Spin Echoes 1.1 Find f0, transmit amplitudes, and shim settings In order to acquire spin echoes, we first need to find the appropriate scanner settings using the FID GUI. This was all done last week,
Setting up a Multi sine impedance measurement
 Setting up a Multi sine impedance measurement Case study: how do I setup a Multi Sine impedance measurement? 1 Single sine vs Multi sine Traditional electrochemical impedance spectroscopy measurements
Setting up a Multi sine impedance measurement Case study: how do I setup a Multi Sine impedance measurement? 1 Single sine vs Multi sine Traditional electrochemical impedance spectroscopy measurements
GUIDELINES FOR THE REPRESENTATION OF PULSE SEQUENCES FOR SOLUTION-STATE NUCLEAR MAGNETIC RESONANCE SPECTROMETRY
 Pure Appl. Chem., Vol. 73, No. 11, pp. 1749 1764, 2001. 2001 IUPAC INTERNATIONAL UNION OF PURE AND APPLIED CHEMISTRY COMMITTEE ON PRINTED AND ELECTRONIC PUBLICATIONS WORKING PARTY ON SPECTROSCOPIC DATA
Pure Appl. Chem., Vol. 73, No. 11, pp. 1749 1764, 2001. 2001 IUPAC INTERNATIONAL UNION OF PURE AND APPLIED CHEMISTRY COMMITTEE ON PRINTED AND ELECTRONIC PUBLICATIONS WORKING PARTY ON SPECTROSCOPIC DATA
Chapter 11 Coherence Editing: Pulse-field Gradients and Phase Cycling
 Chapter 11 Coherence Editing: Pulse-field Gradients and Phase Cycling Coherence editing is used to remove unwanted signals from NMR spectra. For example, in the double quantum filtered COSY experiment,
Chapter 11 Coherence Editing: Pulse-field Gradients and Phase Cycling Coherence editing is used to remove unwanted signals from NMR spectra. For example, in the double quantum filtered COSY experiment,
Supplementary Figure 1. Scanning Electron Microscopy images of the pristine electrodes. (a) negative electrode and (b) positive electrode.
 a b Supplementary Figure 1. Scanning Electron Microscopy images of the pristine electrodes. (a) negative electrode and (b) positive electrode. Images were performed using a FEI/Philips XL4 microscope with
a b Supplementary Figure 1. Scanning Electron Microscopy images of the pristine electrodes. (a) negative electrode and (b) positive electrode. Images were performed using a FEI/Philips XL4 microscope with
= knd 1/ 2 m 2 / 3 t 1/ 6 c
 DNA Sequencing with Sinusoidal Voltammetry Brazill, S. A., P. H. Kim, et al. (2001). "Capillary Gel Electrophoresis with Sinusoidal Voltammetric Detection: A Strategy To Allow Four-"Color" DNA Sequencing."
DNA Sequencing with Sinusoidal Voltammetry Brazill, S. A., P. H. Kim, et al. (2001). "Capillary Gel Electrophoresis with Sinusoidal Voltammetric Detection: A Strategy To Allow Four-"Color" DNA Sequencing."
STEM Spectrum Imaging Tutorial
 STEM Spectrum Imaging Tutorial Gatan, Inc. 5933 Coronado Lane, Pleasanton, CA 94588 Tel: (925) 463-0200 Fax: (925) 463-0204 April 2001 Contents 1 Introduction 1.1 What is Spectrum Imaging? 2 Hardware 3
STEM Spectrum Imaging Tutorial Gatan, Inc. 5933 Coronado Lane, Pleasanton, CA 94588 Tel: (925) 463-0200 Fax: (925) 463-0204 April 2001 Contents 1 Introduction 1.1 What is Spectrum Imaging? 2 Hardware 3
Avance GRASP. Installation/User Manual BRUKER. Version
 Avance GRASP Installation/User Manual Version 002 BRUKER The information in this manual may be altered without notice. BRUKER accepts no responsibility for actions taken as a result of use of this manual.
Avance GRASP Installation/User Manual Version 002 BRUKER The information in this manual may be altered without notice. BRUKER accepts no responsibility for actions taken as a result of use of this manual.
TUTORIAL PROGRAM FID (Windows 95 Version)
 TUTORIAL PROGRAM FID (Windows 95 Version) FID was written to help beginners understand the features of the pulse NMR experiment. For a given set of input parameters, which include frequencies, intensities,
TUTORIAL PROGRAM FID (Windows 95 Version) FID was written to help beginners understand the features of the pulse NMR experiment. For a given set of input parameters, which include frequencies, intensities,
Supplementary Information
 Supplementary Information CP HISQC: a better version of HSQC experiment for intrinsically disordered proteins under physiological conditions. Tairan Yuwen a & Nikolai R. Skrynnikov a,b * (a) Department
Supplementary Information CP HISQC: a better version of HSQC experiment for intrinsically disordered proteins under physiological conditions. Tairan Yuwen a & Nikolai R. Skrynnikov a,b * (a) Department
To a slurry of 2,2 -dilithiobiphenyl bis TMEDA adduct (16) (27.0 g, 67.8 mmol) in diethyl
 Page S1 Contents of the supporting information:?? Experimental procedure for 19.?? Characterization of 27 (including 1 H-, 13 C-, DEPT, 1 H- 1 H COSY, 1 H- 13 C correlation spectra) and X-Ray data for
Page S1 Contents of the supporting information:?? Experimental procedure for 19.?? Characterization of 27 (including 1 H-, 13 C-, DEPT, 1 H- 1 H COSY, 1 H- 13 C correlation spectra) and X-Ray data for
H 2 O and fat imaging
 H 2 O and fat imaging Xu Feng Outline Introduction benefit from the separation of water and fat imaging Chemical Shift definition of chemical shift origin of chemical shift equations of chemical shift
H 2 O and fat imaging Xu Feng Outline Introduction benefit from the separation of water and fat imaging Chemical Shift definition of chemical shift origin of chemical shift equations of chemical shift
Temps can range -130 to +120 C. See instructions below (part I). Such data can also be acquired on Athena and the AVANCE-360.
 Flourine-19 Experiments on Varian Spectrometers updated: 2010 May 12 The difficulty with 19 F experiments involving decoupling (or 2D heterocorrelation) is the close proximity of the 19 F resonant frequency
Flourine-19 Experiments on Varian Spectrometers updated: 2010 May 12 The difficulty with 19 F experiments involving decoupling (or 2D heterocorrelation) is the close proximity of the 19 F resonant frequency
Experiment 2: Electronic Enhancement of S/N and Boxcar Filtering
 Experiment 2: Electronic Enhancement of S/N and Boxcar Filtering Synopsis: A simple waveform generator will apply a triangular voltage ramp through an R/C circuit. A storage digital oscilloscope, or an
Experiment 2: Electronic Enhancement of S/N and Boxcar Filtering Synopsis: A simple waveform generator will apply a triangular voltage ramp through an R/C circuit. A storage digital oscilloscope, or an
COMMUNICATIONS Volume-Selective Multipulse Spin-Echo Spectroscopy
 JOURNAL OF MAGNETC RESONANCE 72,379-384 (1987) COMMUNCATONS Volume-Selective Multipulse Spin-Echo Spectroscopy R. KMMCH* AND D. HOEPFEL? *Universitri t Urn, Sektion Kernresonanzspektroskopie, D-7900 Urn,
JOURNAL OF MAGNETC RESONANCE 72,379-384 (1987) COMMUNCATONS Volume-Selective Multipulse Spin-Echo Spectroscopy R. KMMCH* AND D. HOEPFEL? *Universitri t Urn, Sektion Kernresonanzspektroskopie, D-7900 Urn,
OPTIMIZATION AND PRACTICAL IMPLEMENTATION OF ULTRAFAST 2D NMR EXPERIMENTS
 Quim. Nova, Vol. 36, No. 4, 577-581, 2013 OPTIMIZATION AND PRACTICAL IMPLEMENTATION OF ULTRAFAST 2D NMR EXPERIMENTS Luiz H. K. Queiroz Júnior* Departamento de Química, Universidade Federal de São Carlos,
Quim. Nova, Vol. 36, No. 4, 577-581, 2013 OPTIMIZATION AND PRACTICAL IMPLEMENTATION OF ULTRAFAST 2D NMR EXPERIMENTS Luiz H. K. Queiroz Júnior* Departamento de Química, Universidade Federal de São Carlos,
The Agilent OneNMR Probe
 The Agilent OneNMR Probe Technical Overview Introduction The Agilent OneNMR probe represents a new class of NMR probes. This technology is the most signifi cant advance in solution-state probes in over
The Agilent OneNMR Probe Technical Overview Introduction The Agilent OneNMR probe represents a new class of NMR probes. This technology is the most signifi cant advance in solution-state probes in over
Supporting Information
 Supporting Information Loss of the phenolic hydroxyl group and aromaticity from the side chain of anti-proliferative 0-methyl-aplog-, a simplified analog of aplysiatoxin, enhances its tumor-promoting and
Supporting Information Loss of the phenolic hydroxyl group and aromaticity from the side chain of anti-proliferative 0-methyl-aplog-, a simplified analog of aplysiatoxin, enhances its tumor-promoting and
SpinCore RadioProcessor LabVIEW Extensions
 NMR Interface User's Manual SpinCore Technologies, Inc. http:// Congratulations and thank you for choosing a design from SpinCore Technologies, Inc. We appreciate your business! At SpinCore we try to fully
NMR Interface User's Manual SpinCore Technologies, Inc. http:// Congratulations and thank you for choosing a design from SpinCore Technologies, Inc. We appreciate your business! At SpinCore we try to fully
Evaluation of Adiabatic Frequency-Modulated Schemes for Broadband Decoupling in Isotropic Liquids
 JOURNAL OF MAGNETIC RESONANCE, Series A 119, 129 133 (1996) ARTICLE NO. 0062 Evaluation of Adiabatic Frequency-Modulated Schemes for Broadband Decoupling in Isotropic Liquids RIQIANG FU AND GEOFFREY BODENHAUSEN*
JOURNAL OF MAGNETIC RESONANCE, Series A 119, 129 133 (1996) ARTICLE NO. 0062 Evaluation of Adiabatic Frequency-Modulated Schemes for Broadband Decoupling in Isotropic Liquids RIQIANG FU AND GEOFFREY BODENHAUSEN*
1 Introduction. 2 The basic principles of NMR
 1 Introduction Since 1977 when the first clinical MRI scanner was patented nuclear magnetic resonance imaging is increasingly being used for medical diagnosis and in scientific research and application
1 Introduction Since 1977 when the first clinical MRI scanner was patented nuclear magnetic resonance imaging is increasingly being used for medical diagnosis and in scientific research and application
Supporting Information
 Highly diastereoselective cyclopropanation of -methylstyrene catalyzed by a C 2 -symmetrical chiral iron porphyrin complex Daniela Intrieri, Stéphane Le Gac, Alessandro Caselli, Eric Rose, Bernard Boitrel,
Highly diastereoselective cyclopropanation of -methylstyrene catalyzed by a C 2 -symmetrical chiral iron porphyrin complex Daniela Intrieri, Stéphane Le Gac, Alessandro Caselli, Eric Rose, Bernard Boitrel,
Supplementary Figure 1 Overview of samples used for the structure determination of the Pf 26mer Box C/D RNA. a, A,C lab -RNA, b, A,G lab -RNA, c, A,U
 Supplementary Figure 1 Overview of samples used for the structure determination of the Pf 26mer Box C/D RNA. a, A,C lab -RNA, b, A,G lab -RNA, c, A,U lab -RNA, d, C,U lab -RNA, e, G,C lab -RNA, f, G,U
Supplementary Figure 1 Overview of samples used for the structure determination of the Pf 26mer Box C/D RNA. a, A,C lab -RNA, b, A,G lab -RNA, c, A,U lab -RNA, d, C,U lab -RNA, e, G,C lab -RNA, f, G,U
Panasonic, 2 Channel FFT Analyzer VS-3321A. DC to 200kHz,512K word memory,and 2sets of FDD
 Panasonic, 2 Channel FFT Analyzer VS-3321A DC to 200kHz,512K word memory,and 2sets of FDD New generation 2CH FFT Anal General The FFT analyzer is a realtime signal analyzer using the Fast Fourier Transform
Panasonic, 2 Channel FFT Analyzer VS-3321A DC to 200kHz,512K word memory,and 2sets of FDD New generation 2CH FFT Anal General The FFT analyzer is a realtime signal analyzer using the Fast Fourier Transform
ENGR 210 Lab 12: Sampling and Aliasing
 ENGR 21 Lab 12: Sampling and Aliasing In the previous lab you examined how A/D converters actually work. In this lab we will consider some of the consequences of how fast you sample and of the signal processing
ENGR 21 Lab 12: Sampling and Aliasing In the previous lab you examined how A/D converters actually work. In this lab we will consider some of the consequences of how fast you sample and of the signal processing
An EPR Primer 2. Basic EPR Theory 2.1. Introduction to Spectroscopy 2.1.1
 An EPR Primer 2 This chapter is an introduction to the basic theory and practice of EPR spectroscopy. It gives you sufficient background to understand the following chapters. In addition, we strongly encourage
An EPR Primer 2 This chapter is an introduction to the basic theory and practice of EPR spectroscopy. It gives you sufficient background to understand the following chapters. In addition, we strongly encourage
7 Equipments. Spectrometers
 7 Equipments Spectrometers There are three spectrometers located in the NMR laboratory: Varian UNITYplus 500 MHz (NMR500), Varian UNITYplus 500 MHz (Nightmare) and Varian INOVA 600 MHz. All spectrometers
7 Equipments Spectrometers There are three spectrometers located in the NMR laboratory: Varian UNITYplus 500 MHz (NMR500), Varian UNITYplus 500 MHz (Nightmare) and Varian INOVA 600 MHz. All spectrometers
LLS - Introduction to Equipment
 Published on Advanced Lab (http://experimentationlab.berkeley.edu) Home > LLS - Introduction to Equipment LLS - Introduction to Equipment All pages in this lab 1. Low Light Signal Measurements [1] 2. Introduction
Published on Advanced Lab (http://experimentationlab.berkeley.edu) Home > LLS - Introduction to Equipment LLS - Introduction to Equipment All pages in this lab 1. Low Light Signal Measurements [1] 2. Introduction
6.S02 MRI Lab Acquire MR signals. 2.1 Free Induction decay (FID)
 6.S02 MRI Lab 1 2. Acquire MR signals Connecting to the scanner Connect to VMware on the Lab Macs. Download and extract the following zip file in the MRI Lab dropbox folder: https://www.dropbox.com/s/ga8ga4a0sxwe62e/mit_download.zip
6.S02 MRI Lab 1 2. Acquire MR signals Connecting to the scanner Connect to VMware on the Lab Macs. Download and extract the following zip file in the MRI Lab dropbox folder: https://www.dropbox.com/s/ga8ga4a0sxwe62e/mit_download.zip
Analysis of Data Chemistry 838
 Chemistry 838 Thomas V. Atkinson, Ph.D. Senior Academic Specialist Department of Chemistry Michigan State University East Lansing, MI 4884 TABLE OF CONTENTS TABLE OF CONTENTS...1 TABLE OF TABLES...1 TABLE
Chemistry 838 Thomas V. Atkinson, Ph.D. Senior Academic Specialist Department of Chemistry Michigan State University East Lansing, MI 4884 TABLE OF CONTENTS TABLE OF CONTENTS...1 TABLE OF TABLES...1 TABLE
Instrumental Considerations
 Instrumental Considerations Many of the limits of detection that are reported are for the instrument and not for the complete method. This may be because the instrument is the one thing that the analyst
Instrumental Considerations Many of the limits of detection that are reported are for the instrument and not for the complete method. This may be because the instrument is the one thing that the analyst
M R I Physics Course. Jerry Allison Ph.D., Chris Wright B.S., Tom Lavin B.S., Nathan Yanasak Ph.D. Department of Radiology Medical College of Georgia
 M R I Physics Course Jerry Allison Ph.D., Chris Wright B.S., Tom Lavin B.S., Nathan Yanasak Ph.D. Department of Radiology Medical College of Georgia M R I Physics Course Magnetic Resonance Imaging Spatial
M R I Physics Course Jerry Allison Ph.D., Chris Wright B.S., Tom Lavin B.S., Nathan Yanasak Ph.D. Department of Radiology Medical College of Georgia M R I Physics Course Magnetic Resonance Imaging Spatial
PHY3902 PHY3904. Nuclear magnetic resonance Laboratory Protocol
 PHY3902 PHY3904 Nuclear magnetic resonance Laboratory Protocol PHY3902 PHY3904 Nuclear magnetic resonance Laboratory Protocol GETTING STARTED You might be tempted now to put a sample in the probe and try
PHY3902 PHY3904 Nuclear magnetic resonance Laboratory Protocol PHY3902 PHY3904 Nuclear magnetic resonance Laboratory Protocol GETTING STARTED You might be tempted now to put a sample in the probe and try
B12. SIMPSON Exercises
 B12 SIMPSON Exercises Thomas Vosegaard SIMPSON may be downloaded from http://www.bionmr.chem.au.dk/download/b12. SIMPSON Exercises Everything typed in Courier refers to commands to be typed on the keyboard
B12 SIMPSON Exercises Thomas Vosegaard SIMPSON may be downloaded from http://www.bionmr.chem.au.dk/download/b12. SIMPSON Exercises Everything typed in Courier refers to commands to be typed on the keyboard
Fundamentals of NMR MRI Workshop Dayananda Sagar Institutes 29 May 2014 K.V.RAMANATHAN NMR Research Centre Indian Institute of Science Bangalore
 Fundamentals of NMR MRI Workshop Dayananda Sagar Institutes 29 May 2014 K.V.RAMANATHAN NMR Research Centre Indian Institute of Science Bangalore NMR It is an Ubiquitous Technique Physics Chemistry Structural
Fundamentals of NMR MRI Workshop Dayananda Sagar Institutes 29 May 2014 K.V.RAMANATHAN NMR Research Centre Indian Institute of Science Bangalore NMR It is an Ubiquitous Technique Physics Chemistry Structural
HETERONUCLEAR IMAGING. Topics to be Discussed:
 HETERONUCLEAR IMAGING BioE-594 Advanced MRI By:- Rajitha Mullapudi 04/06/2006 Topics to be Discussed: What is heteronuclear imaging. Comparing the hardware of MRI and heteronuclear imaging. Clinical applications
HETERONUCLEAR IMAGING BioE-594 Advanced MRI By:- Rajitha Mullapudi 04/06/2006 Topics to be Discussed: What is heteronuclear imaging. Comparing the hardware of MRI and heteronuclear imaging. Clinical applications
The Multidimensional Filter Diagonalization Method
 Journal of Magnetic Resonance 144, 357 366 (2000) doi:10.1006/jmre.2000.2066, available online at http://www.idealibrary.com on The Multidimensional Filter Diagonalization Method II. Application to 2D
Journal of Magnetic Resonance 144, 357 366 (2000) doi:10.1006/jmre.2000.2066, available online at http://www.idealibrary.com on The Multidimensional Filter Diagonalization Method II. Application to 2D
Qualion NMR Process\Lab NMR (Nuclei Magnetic Resonance) Analyzer Model MASH, Style D
 Qualion NMR Process\Lab NMR (Nuclei Magnetic Resonance) Analyzer Model MASH, Style D User Guide UM SW50407.05 Table of Contents Table of Contents Table of Contents...1 Preface...6 1. SYSTEM OVERVIEW...
Qualion NMR Process\Lab NMR (Nuclei Magnetic Resonance) Analyzer Model MASH, Style D User Guide UM SW50407.05 Table of Contents Table of Contents Table of Contents...1 Preface...6 1. SYSTEM OVERVIEW...
List of Commands and Parameters
 List of Commands and Parameters For information about a command or parameter, in VNMR top window, type: man('parameter_name') the information will display in bottom text window Below are a list of commands
List of Commands and Parameters For information about a command or parameter, in VNMR top window, type: man('parameter_name') the information will display in bottom text window Below are a list of commands
The Scientist and Engineer's Guide to Digital Signal Processing By Steven W. Smith, Ph.D.
 The Scientist and Engineer's Guide to Digital Signal Processing By Steven W. Smith, Ph.D. Home The Book by Chapters About the Book Steven W. Smith Blog Contact Book Search Download this chapter in PDF
The Scientist and Engineer's Guide to Digital Signal Processing By Steven W. Smith, Ph.D. Home The Book by Chapters About the Book Steven W. Smith Blog Contact Book Search Download this chapter in PDF
Seized Drugs Operational Guidelines for the Thermo FTIR Comparative and Analytical Division
 Operational Guidelines for the Thermo FTIR Comparative and Analytical Division THERMO FOURIER TRANSFORM INFRARED (FTIR) SPECTROMETER Instrument Nicolet 4700 Series FTIR spectrometer (Serial Number AFZ0400253)
Operational Guidelines for the Thermo FTIR Comparative and Analytical Division THERMO FOURIER TRANSFORM INFRARED (FTIR) SPECTROMETER Instrument Nicolet 4700 Series FTIR spectrometer (Serial Number AFZ0400253)
Instruction manual for T3DS software. Tool for THz Time-Domain Spectroscopy. Release 4.0
 Instruction manual for T3DS software Release 4.0 Table of contents 0. Setup... 3 1. Start-up... 5 2. Input parameters and delay line control... 6 3. Slow scan measurement... 8 4. Fast scan measurement...
Instruction manual for T3DS software Release 4.0 Table of contents 0. Setup... 3 1. Start-up... 5 2. Input parameters and delay line control... 6 3. Slow scan measurement... 8 4. Fast scan measurement...
