Principal Component Analysis, PCA, in R
|
|
|
- Amos Pearson
- 6 years ago
- Views:
Transcription
1 enote 2 1 enote 2 Principal Component Analysis, PCA, in R
2 enote 2 INDHOLD 2 Indhold 2 Principal Component Analysis, PCA, in R Reading about PCA Example: Fisher s Iris Data Data import Basic explorative analysis PCA of Iris data Spectral data example: yarn data Exercises Reading about PCA You can use the Wehrens book, Chapter 4, pp 43-56: page/1 and/or (probably better) the Varmuza-book, chapter 3, sections : The two R-packages chemometrics and ChemometricswithR, are companions to the two books. Bro and Smilde (2014): Principal Component Analysis Analytical Methods TUTORIAL REVIEW, 6,
3 enote READING ABOUT PCA 3 Below there will be a number of important plots examplified as part of the iris-example: 1. Variance-plots ( scree-type plots) 2. Scores, loadings and biplots (main plots for interpretation of structure) 3. Explained variances for each variable 4. Validation/diagnostics plots: (a) Leverage and residuals (also called score distances and orthogonal distances (cf. the nice Figure 3.15, page 79 in the Varmuza-book) (b) The influence plot : residuals versus leverage 5. Jacknifing/bootstrapping/Crossvalidating the PCA for various purposes: (a) Deciding on number of components (b) Sensitivity/uncertainty investigation of scores and loadings. What is PCA: Developed by Karl Pearson in 1901: Pearson, K. (1901) On lines and planes of closest fit to systems of points in space. Philosophical Magazine (6) 2:
4 enote READING ABOUT PCA 4 May also be called: Singular value decomposition Karhunen-Loéve expansion Eigenvector analysis Latent vector analysis Characteristic vector analysis PCA is used for many things: Projection method Exploratory data analysis Extract information and remove noise Reduce dimensionality / Compression (Clustering) And can be described/expressed in many ways:
5 enote READING ABOUT PCA 5 Produces optimal low-dimensional plots of observations (scores) Provides an overview of the variable correlation structure (loadings) Finds linear combinations of maximal variance Orthogonal distance regression method A bilinear model for the data And can be described/expressed in many ways: X : The (centered and scaled) n p data matrix X = Observation Scores Variable Loadings + Error X = TP T + E
6 enote EXAMPLE: FISHER S IRIS DATA 6 Computations/A bit of math: X ij = A t ia p aj + e ij a=1 PCA finds X-components with maximal Y-variance: max Var(Xα) α =1 PCA is the least squares fit of the bilinear (non linear regression) model: min t,p ij PCA is the eigen decomposition of X t X PCA is the eigen decomposition of XX t (x ij A a=1 t ia p aj ) 2 PCA is the outcome of (a version of) the NIPALS algorithm 2.2 Example: Fisher s Iris Data Below there will be an exercise based on these data with some questions that PCA can be helpful in answering. Here we examplify a number of visualizations that one could do for such data including PCA-based stuff. The Fisher Iris data-set is classic, c.f.:
7 enote EXAMPLE: FISHER S IRIS DATA 7 Fisher, R.A. (1936). The use of multiple measurements in taxonomic problem. Annals of Eugenics 7: Anderson, E. (1935). The irises of the Gaspe Peninsula. Bulletin of the American Iris Society 59: 2-5. There are 150 objects, 50 Iris setosa, 50 Iris versicolor and 50 Iris virginica. The flowers of these 150 plants have been measures by a ruler. The variables are sepal length (SL), sepal width (SW), petal length (PL) and petal width PW), all in all only four variables. The original hypothesis was that I. versicolor was a hybrid of the two other species i.e. I. setosa x virginica. I. setosa is diploid; I. virginica is a tetraploid; and I. versicolor is hexaploid Data import The iris data can allready be found within R, so no import is needed: # Loading package related to Varmuza-book # (First time you need to install the package) library(chemometricswithrdata) library(chemometricswithr) data(iris) Or read the IRIS csv-data which is a copy of the file uploaded on CampusNet. Note that the Iris data given in CampusNet is slightly different from the IRIS data available. First save the data set on your computer and set the relevant working direcctory in R, e.g. by clikcing Session and choosinf Set working directory, or run the following command with the correct chosen folder path: setwd("c:/myfolderpath") And then import the data into R as follows: JCFiris=read.table("Fisher_JCF.csv",header=T,sep=";",dec=",") Note that the Iris data given by JCF is slightly different from the IRIS data available in R:
8 enote EXAMPLE: FISHER S IRIS DATA 8 summary(iris) Sepal.Width Petal.Length Petal.Width Min. :4.30 Min. :2.00 Min. :1.00 Min. :0.1 1st Qu.:5.10 1st Qu.:2.80 1st Qu.:1.60 1st Qu.:0.3 Median :5.80 Median :3.00 Median :4.35 Median :1.3 Mean :5.84 Mean :3.06 Mean :3.76 Mean :1.2 3rd Qu.:6.40 3rd Qu.:3.30 3rd Qu.:5.10 3rd Qu.:1.8 Max. :7.90 Max. :4.40 Max. :6.90 Max. :2.5 Species setosa :50 versicolor:50 virginica :50 summary(jcfiris) X PW PL SW setosa :50 Min. : 1.0 Min. :10.0 Min. :20.0 versicolor:50 1st Qu.: 3.0 1st Qu.:16.0 1st Qu.:28.0 virginica :50 Median :13.0 Median :44.0 Median :30.0 Mean :11.9 Mean :37.8 Mean :30.6 3rd Qu.:18.0 3rd Qu.:51.0 3rd Qu.:33.0 Max. :25.0 Max. :69.0 Max. :44.0 SL Min. : st Qu.: 51.0 Median : 58.0 Mean : rd Qu.: 64.0 Max. :699.0 Note the differences: The names, order and scales. AND: an outlier in the JCF-version has been changed in the R-version. Look at the first 6 observations: head(iris) Sepal.Width Petal.Length Petal.Width Species
9 enote EXAMPLE: FISHER S IRIS DATA setosa setosa setosa setosa setosa setosa head(jcfiris) X PW PL SW SL 1 setosa virginica virginica setosa virginica virginica The dimensions are the same: dim(iris) [1] dim(jcfiris) [1] Basic explorative analysis First we do some classic (univariate) explorative analysis: # 4 boxplots with color: par(mar=c(4,2,3,2),mfrow=c(2,2)) for (i in 1:4) boxplot(iris[,i] ~ iris[,5], col = 1:3, main = names(iris)[i])
10 enote EXAMPLE: FISHER S IRIS DATA 10 Sepal.Width setosa versicolor virginica setosa versicolor virginica Petal.Length Petal.Width setosa versicolor virginica setosa versicolor virginica The par(mar=c(4,2,3,2)) command controls the four margins of each individual plot in the order: bottom, left, top, right. This is helpful to make nice multi-plot pages. # Pairwise scatters: pairs(iris,col = iris$species)
11 enote EXAMPLE: FISHER S IRIS DATA Sepal.Width Petal.Length Petal.Width Species Let us, for the record, have a look at the covariance matrix: cov(iris[,1:4]) And similarly the correlation matrix: cor(iris[,1:4])
12 enote EXAMPLE: FISHER S IRIS DATA 12 Sepal.Width Petal.Length Petal.Width Sepal.Width Petal.Length Petal.Width Sepal.Width Petal.Length Petal.Width Sepal.Width Petal.Length Petal.Width PCA of Iris data First we do a basic PCA on covariances (WITHOUT Standardization - ONLY with centering): (and here using the PCA function of the ChemometricsWithR-package) irispc_without=pca(scale(iris[,1:4], scale = FALSE)) Note that the scale-function is used here to just center the four variables. # A good selection of 4 core plots: par(mar=c(4,2,3,2),mfrow=c(2,2)) scoreplot(irispc_without, col = iris$species, main = "Scores") loadingplot(irispc_without, show.names = TRUE, main = "Loadings") biplot(irispc_without, score.col = iris$species, main = "biplot") screeplot(irispc_without, type = "percentage", main = "Explained variance")
13 enote EXAMPLE: FISHER S IRIS DATA Scores PC 1 (92.5%) PC 2 (5.3%) Loadings PC 1 (92.5%) PC 2 (5.3%) pal.width Petal.Length Petal.Width biplot PC 1 (92.5%) PC 2 (5.3%) Explained variance # PCs And now the PCA on correlations (WITH Standardization - AND with centering): irispc <- PCA(scale(iris[,1:4])) Note that the scale-function now is used to both center and standardize the four variables - he default choice of this function. par(mar=c(4,2,3,2),mfrow=c(2,2)) scoreplot(irispc, col = iris$species, main = "Scores")
14 enote EXAMPLE: FISHER S IRIS DATA 14 loadingplot(irispc, show.names = TRUE, main = "Loadings") biplot(irispc, score.col = iris$species, main = "biplot") screeplot(irispc, type = "percentage", main = "Explained variance") Scores PC 1 (73.0%) PC 2 (22.9%) Loadings PC 1 (73.0%) PC 2 (22.9%) epal.width Petal.Lengt Petal.Width biplot PC 1 (73.0%) PC 2 (22.9%) Explained variance # PCs There can be other versions of the variance plot, e.g.: par(mfrow=c(1,2)) plot(1:length(irispc$var), irispc$var, cex = 2, ylab = "variance explained",xlab = "n PC")
15 enote EXAMPLE: FISHER S IRIS DATA 15 lines(1:length(irispc$var), irispc$var) plot(1:length(irispc$var), irispc$var/sum(irispc$var), cex = 2, ylab = "(explained variance)/(total variance)",xlab = "n PC") lines(1:length(irispc$var), irispc$var/sum(irispc$var)) variance explained (explained variance)/(total variance) n PC n PC It can be useful to plot more components than just the first two: # Scores: pairs(scores(irispc), col = iris$species)
16 enote EXAMPLE: FISHER S IRIS DATA 16 PC PC 2 PC PC 4 # Loadings: par(mfrow = c(4,4), mar = c(4,4,.1,.1)) for (i in 1:4) for (j in 1:4) loadingplot(irispc, show.names = TRUE,pc=c(i,j), cex.lab=0.7)
17 enote EXAMPLE: FISHER S IRIS DATA 17 PC 1 (73.0%) pal.width Petal.Len Petal.Wid Sepal.Lengt PC 2 (22.9%) pal.width Petal.Len Petal.Wid Sepal.Lengt PC 3 (3.7%) pal.width Sepal.Lengt Petal.Len Petal.Wid PC 4 (0.5%) pal.width Petal.Wid Sepal.Lengt Petal.Len PC 1 (73.0%) PC 1 (73.0%) PC 1 (73.0%) PC 1 (73.0%) PC 1 (73.0%) epal.width Petal.Widt Petal.Len PC 2 (22.9%) epal.width PC 3 (3.7%) epal.width Petal.Len Petal.Widt PC 4 (0.5%) epal.width Petal.Widt Petal.Len PC 2 (22.9%) PC 2 (22.9%) PC 2 (22.9%) PC 2 (22.9%) PC 1 (73.0%) Petal.Length tal.width Sepal.Width Sepal.Len PC 2 (22.9%) Petal.Length tal.width Sepal.Width Sepal.Len PC 3 (3.7%) tal.width Petal.Length Sepal.Width Sepal.Len PC 4 (0.5%) tal.width Sepal.Width Petal.Length Sepal.Len PC 3 (3.7%) PC 3 (3.7%) PC 3 (3.7%) PC 3 (3.7%) PC 1 (73.0%) al.length Petal.Wi Sepal.Width PC 2 (22.9%) al.length Petal.Wi Sepal.Width PC 3 (3.7%) al.length Sepal.Width Petal.Wi PC 4 (0.5%) al.length Petal.Wi Sepal.Width PC 4 (0.5%) PC 4 (0.5%) PC 4 (0.5%) PC 4 (0.5%) A much nicer biplot can be created by the ggbiplot-package: (Now using the prcomp-function to do the PCA) ir.pca <- prcomp(iris[,1:4], center = TRUE, scale. = TRUE) library(devtools) # First time install: install_github("ggbiplot", "vqv") library(ggbiplot) g <- ggbiplot(ir.pca, obs.scale = 1, var.scale = 1,
18 enote EXAMPLE: FISHER S IRIS DATA 18 groups = iris[,5], ellipse = TRUE, circle = FALSE) print(g) Sepal.Width Petal.Length Petal.Width PC1 (73.0% explained var.) PC2 (22.9% explained var.) groups setosa versicolor virginica Generally about interpreting PCA plots: Look at variances (scree) - hope for few(2) - look for the bend Look at scores and loadings (e.g. biplot) Scores: OBSERVATION mapping preserves inter observation distances (as good as possible) Loadings: VARIABLE mapping (correlation structure)
19 enote EXAMPLE: FISHER S IRIS DATA 19 Variables in the SAME DIRECTION from (0,0) AND far away from (0,0) are highly correlated Loadings tell us on which variables the observations differ An observation to the right has high values on the variables with (large) loadings to the right An observation to the left has low values on the variables with (large) loadings to the right Look at residuals (Orthogonal distances) and leverages (score distances) (Outliers etc) Finally, let us show some of the diagnostics (residuals) plotting. For this we will use the chemometrics package: (and now the princomp function for the PCA) library(chemometrics) irispca <- princomp(iris[,1:4], cor = TRUE) # The score distances res SDist express the leverage values # The orthogonal distances express the residuals ## Plots vs object number : res <- pcadiagplot(iris[,1:4], irispca, a = 2)
20 enote EXAMPLE: FISHER S IRIS DATA Object number Score distance SD Object number Orthogonal distance OD ## Plot of the two agains each other: par(mfrow=c(1,2)) plot(res$sdist, res$odist, type = "n") text(res$sdist, res$odist, labels = row.names(iris)) ## Explained variance for each variable pcavarexpl(iris[,1:4],a=2)
21 enote EXAMPLE: FISHER S IRIS DATA 21 res$odist Explained variance Petal.Length res$sdist # Influence plot: residuals versus leverage # for different number of components: par(mfrow=c(2,2)) for (i in 1:4) { res=pcadiagplot(iris[,1:4],a=i,irispca,plot=false) plot(res$sdist,res$odist,type="n") text(res$sdist,res$odist,labels=row.names(iris)) }
22 enote EXAMPLE: FISHER S IRIS DATA 22 res$odist res$odist res$sdist res$sdist res$odist res$odist 5.0e e e res$sdist res$sdist Finally, finally let us indicate how one could do some re-sampling (similar to jacknifing ): Leaving out a certain number of the observation and plotting the loadings and/or scores for each subset data. First the loadings: # Random samples of a certain proportion of the # original number of observations are left out par(mar = c(1,1,1,1), mfrow = c(3,3)) n=length(iris[,1]) leave_out_size=0.50 for (k in 1:9){ irispc=pca(scale(iris[sample(1:n,round(n*(1-leave_out_size))),1:4])) loadingplot(irispc, show.names = TRUE, main = "Loadings")
23 enote EXAMPLE: FISHER S IRIS DATA 23 } PC 2 (22.4%) pal.width Loadings Petal.Leng Petal.Widt PC 2 (23.3%) Loadings Petal.Width etal.length Sepal.Wi PC 2 (23.9%) etal.length Petal.Width Loadings Sepal.W PC 2 (24.0%) Loadings Sepal.Wi Petal.Width etal.length PC 2 (20.8%) Loadings Sepal.Wi Petal.Width tal.length PC 2 (22.4%) Loadings pal.width Petal.Width Petal.Leng PC 2 (23.1%) Loadings etal.length Petal.Width The the scores: Sepal.Wi PC 2 (22.0%) Loadings etal.length Petal.Width Sepal.Wi PC 2 (23.4%) Loadings pal.width Petal.Leng Petal.Width par(mar = c(1,1,1,1), mfrow = c(3,3)) for (k in 1:9){ subsample <- sample(1:n,round(n*(1-leave_out_size))) irispc <- PCA(scale(iris[subsample,1:4])) scoreplot(irispc, col = iris$species[subsample], main = "Scores") }
24 enote SPECTRAL DATA EXAMPLE: YARN DATA Scores PC 2 (20.2%) Scores PC 2 (21.2%) Scores PC 2 (24.5%) Scores PC 2 (25.0%) Scores PC 2 (26.3%) Scores PC 2 (21.1%) Scores PC 2 (23.4%) Scores PC 2 (23.8%) Scores PC 2 (21.7%) The choice of showing 9 is arbitrary. Other plots of this re-sampling type could be thought of. 2.3 Spectral data example: yarn data ## Spectral data, data(yarn) # Part of chemometrics package # Try:?yarn dim(yarn$nir) ## [1]
25 enote SPECTRAL DATA EXAMPLE: YARN DATA 25 par(mfrow = c(2, 2), mar = c(4, 4,.2,.2)) # Plotting of the 21 individual NIR spectra" max_x=max(yarn$nir) min_x=min(yarn$nir) plot(yarn$nir[1,],type="n",ylim=c(min_x,max_x)) for (i in 1:21) lines(yarn$nir[i,],col=i) # Plotting of the 21 individual NIR spectra - centered" max_x=max(scale(yarn$nir,scale=f)) min_x=min(scale(yarn$nir,scale=f)) plot(scale(yarn$nir[1,],scale=f),type="n",ylim=c(min_x,max_x)) for (i in 1:21) lines(scale(yarn$nir,scale=f)[i,],col=i) # Plotting of the 21 individual NIR spectra - centered and scaled" max_x=max(scale(yarn$nir)) min_x=min(scale(yarn$nir)) plot(scale(yarn$nir[1,]),type="n",ylim=c(min_x,max_x)) for (i in 1:21) lines(scale(yarn$nir)[i,],col=i) # Plotting of the principal variances: " yarnpc <- PCA(scale(yarn$NIR)) plot(1:length(yarnpc$var),yarnpc$var,cex=2) lines(1:length(yarnpc$var),yarnpc$var)
26 enote SPECTRAL DATA EXAMPLE: YARN DATA 26 yarn$nir[1, ] scale(yarn$nir[1, ], scale = F) Index Index scale(yarn$nir[1, ]) yarnpc$var Index :length(yarnPC$var) # Plot of y: plot(yarn$density,type="n") lines(yarn$density)
27 enote EXERCISES 27 yarn$density Index 2.4 Exercises Exercise 1 Fisher s Iris data First examine the raw data and examine whether there are obvious mistakes. After that one could use other Unscrambler features to examine the statistical properties of the objects and variable, but it in this case we go directly to PCA, as this give a very fine overview of the data, and will often show outliers immediately. Perform the PCA with leverage correction and with centering. Examine the four standard plots (score plot, loading plot, influence plot and explained variance plot).
28 enote EXERCISES 28 a) How many principal components would you need and what does the first PC (PC1) describe? b) How many percentage of the variation is described by the first two PCs? c) Can you find an outlier? It so do you have an idea why thus outlier came about? (loadings plot or scores plot)? In R: Do you see problem in the influence plot. If there is an outlier, in which other plot can you see the problem? If you see severe outliers, remove them from the data and run PCA again (and answer a, and b, again) d) Does a standardization (autoscaling) give a better model? (answer a) and b) again) e) How many PCs are needed to explain 70%, 75% and 90% of the variation in the data? f) How many PCs can you maximally get in this dataset? g) Compare the score and the loading plot, and make a biplot. Do any of the variables tell the same story? h) Are any variables more discriminative the others? Are any variables dispensable? i) Can you see the presupposed classes? Any class overlap?
29 enote EXERCISES 29 j) Does the original hypothesis seem to be OK? Exercise 2 Wine Data (To be presented by Team 1 next time) The second dataset is called VIN2: Forina, M., Armanino, C., Castino, M. and Ubigli, M. (1986). Multivariate data analysis as a discriminating method of the origin of wines. Vitis 25: Forina, M., Lanteri, S., Armanino, C., Casolino, C. and Casale, M V-PARVUS. An extendable package of programs for data exploration, classification, and correlation. ( The dataset VIN2.csv is an Excell CSV file. In this dataset there are 178 objects (Italian wines), the first 59 are Barolo wines (B1-B59), the next 71 are Grignolino wines (G60-G130) and the last 48 are Barbera wines (S131-S178). These wines have been characterized by 13 variables (chemical and physical measurements): 1. Alcohol (in %) 2. Malic acid 3. Ash 4. Alkalinity of Ash 5. Magnesium 6. Total phenols 7. Flavanoids 8. Nonflavanoid phenols 9. Proanthocyanins 10. Colour intensity 11. Colour hue 12. OD280 / OD315 of diluted wines 13. Proline (amino acid)
30 enote EXERCISES 30 The wine data can allready be found within R, so no import is needed: # Wines data: # From the JCF uploaded file: # Also slightly different from the version in the package JCFwines=read.table("VIN2.csv",header=T,sep=";",dec=",") # The wines data from the package: # The wine class information is here stored in the wine.classes object data(wines, package = "ChemometricsWithRData") head(wines) alcohol malic acid ash ash alkalinity magnesium tot. phenols [1,] [2,] [3,] [4,] [5,] [6,] flavonoids non-flav. phenols proanth col. int. col. hue OD ratio [1,] [2,] [3,] [4,] [5,] [6,] proline [1,] 1050 [2,] 1185 [3,] 1480 [4,] 735 [5,] 1450 [6,] 1290 head(jcfwines) X Wine F1 F2 F3 F4 F5 F6 F7 F8 F9 F10 F11 F12 1 S1 Barolo S2 Barolo S3 Barolo S4 Barolo
31 enote EXERCISES 31 5 S5 Barolo S6 Barolo F summary(wines) alcohol malic acid ash ash alkalinity Min. :11.0 Min. :0.74 Min. :1.36 Min. :10.6 1st Qu.:12.4 1st Qu.:1.60 1st Qu.:2.21 1st Qu.:17.2 Median :13.1 Median :1.87 Median :2.36 Median :19.5 Mean :13.0 Mean :2.34 Mean :2.37 Mean :19.5 3rd Qu.:13.7 3rd Qu.:3.10 3rd Qu.:2.56 3rd Qu.:21.5 Max. :14.8 Max. :5.80 Max. :3.23 Max. :30.0 magnesium tot. phenols flavonoids non-flav. phenols Min. : 70.0 Min. :0.98 Min. :0.34 Min. : st Qu.: st Qu.:1.74 1st Qu.:1.20 1st Qu.:0.270 Median : 98.0 Median :2.35 Median :2.13 Median :0.340 Mean : 99.6 Mean :2.29 Mean :2.02 Mean : rd Qu.: rd Qu.:2.80 3rd Qu.:2.86 3rd Qu.:0.440 Max. :162.0 Max. :3.88 Max. :5.08 Max. :0.660 proanth col. int. col. hue OD ratio Min. :0.41 Min. : 1.28 Min. :0.480 Min. :1.27 1st Qu.:1.25 1st Qu.: st Qu.: st Qu.:1.93 Median :1.55 Median : 4.68 Median :0.960 Median :2.78 Mean :1.59 Mean : 5.05 Mean :0.957 Mean :2.60 3rd Qu.:1.95 3rd Qu.: rd Qu.: rd Qu.:3.17 Max. :3.58 Max. :13.00 Max. :1.710 Max. :4.00 proline Min. : 278 1st Qu.: 500 Median : 672 Mean : 745 3rd Qu.: 985 Max. :1680 summary(jcfwines)
32 enote EXERCISES 32 X Wine F1 F2 F3 S1 : 1 Barbera:48 Min. : 3.67 Min. :0.74 Min. :1.36 S10 : 1 Barolo :59 1st Qu.: st Qu.:1.60 1st Qu.:2.21 S100 : 1 Grigno :71 Median :13.05 Median :1.86 Median :2.36 S101 : 1 Mean :12.94 Mean :2.34 Mean :2.37 S102 : 1 3rd Qu.: rd Qu.:3.08 3rd Qu.:2.56 S103 : 1 Max. :14.83 Max. :5.80 Max. :3.23 (Other):172 F4 F5 F6 F7 Min. :10.6 Min. : 70.0 Min. :0.98 Min. :0.34 1st Qu.:17.2 1st Qu.: st Qu.:1.74 1st Qu.:1.21 Median :19.5 Median : 98.0 Median :2.35 Median :2.13 Mean :19.5 Mean : 99.7 Mean :2.30 Mean :2.03 3rd Qu.:21.5 3rd Qu.: rd Qu.:2.80 3rd Qu.:2.88 Max. :30.0 Max. :162.0 Max. :3.88 Max. :5.08 F8 F9 F10 F11 Min. :0.130 Min. :0.41 Min. : 1.28 Min. : st Qu.: st Qu.:1.25 1st Qu.: st Qu.:0.782 Median :0.340 Median :1.55 Median : 4.69 Median :0.965 Mean :0.362 Mean :1.59 Mean : 5.06 Mean : rd Qu.: rd Qu.:1.95 3rd Qu.: rd Qu.:1.120 Max. :0.660 Max. :3.58 Max. :13.00 Max. :1.710 F12 F13 Min. :0.56 Min. : 278 1st Qu.:1.92 1st Qu.: 500 Median :2.78 Median : 674 Mean :2.59 Mean : 753 3rd Qu.:3.17 3rd Qu.: 989 Max. :4.00 Max. :1940 a) Examine the raw data. Are there any severe outliers you can detect? What do you think happened with the outlier, if any? b) Correct wrong data, if any (in the excel file), and use PCA again. Does the score and loading plot look significantly different now?
33 enote EXERCISES 33 c) Try PCA without standardization: Which variables are important here and why? d) Try PCA with standardization. Which variables are important here, and would you recommend removing any of them from the data set? Which variables are especially important for the Barbera wines? e) Suppose that alcohol % and proanthocyanins were especially healthy which wine would you recommend? f) Use some re-sampling/jack-knifing methods to test for significance of the variable - are all the variables stable?
Intro to R for Epidemiologists
 Lab 3 (1/29/15) Intro to R for Epidemiologists Many of these questions go beyond the information provided in the lecture. Therefore, you may need to use R help files and the internet to search for answers.
Lab 3 (1/29/15) Intro to R for Epidemiologists Many of these questions go beyond the information provided in the lecture. Therefore, you may need to use R help files and the internet to search for answers.
Session 124TS, A Practical Guide to Machine Learning for Actuaries. Presenters: Dave M. Liner, FSA, MAAA, CERA
 Session 124TS, A Practical Guide to Machine Learning for Actuaries Presenters: Dave M. Liner, FSA, MAAA, CERA SOA Antitrust Disclaimer SOA Presentation Disclaimer A practical guide to machine learning
Session 124TS, A Practical Guide to Machine Learning for Actuaries Presenters: Dave M. Liner, FSA, MAAA, CERA SOA Antitrust Disclaimer SOA Presentation Disclaimer A practical guide to machine learning
Efficient Target Detection from Hyperspectral Images Based On Removal of Signal Independent and Signal Dependent Noise
 IOSR Journal of Electronics and Communication Engineering (IOSR-JECE) e-issn: 2278-2834,p- ISSN: 2278-8735.Volume 9, Issue 6, Ver. III (Nov - Dec. 2014), PP 45-49 Efficient Target Detection from Hyperspectral
IOSR Journal of Electronics and Communication Engineering (IOSR-JECE) e-issn: 2278-2834,p- ISSN: 2278-8735.Volume 9, Issue 6, Ver. III (Nov - Dec. 2014), PP 45-49 Efficient Target Detection from Hyperspectral
Basic Concepts of the R Language
 Basic Concepts of the R Language L. Torgo ltorgo@dcc.fc.up.pt Departamento de Ciência de Computadores Faculdade de Ciências / Universidade do Porto Oct, 2014 Basic Interaction Basic interaction with the
Basic Concepts of the R Language L. Torgo ltorgo@dcc.fc.up.pt Departamento de Ciência de Computadores Faculdade de Ciências / Universidade do Porto Oct, 2014 Basic Interaction Basic interaction with the
The techniques with ERDAS IMAGINE include:
 The techniques with ERDAS IMAGINE include: 1. Data correction - radiometric and geometric correction 2. Radiometric enhancement - enhancing images based on the values of individual pixels 3. Spatial enhancement
The techniques with ERDAS IMAGINE include: 1. Data correction - radiometric and geometric correction 2. Radiometric enhancement - enhancing images based on the values of individual pixels 3. Spatial enhancement
Remote Sensing 4113 Lab 08: Filtering and Principal Components Mar. 28, 2018
 Remote Sensing 4113 Lab 08: Filtering and Principal Components Mar. 28, 2018 In this lab we will explore Filtering and Principal Components analysis. We will again use the Aster data of the Como Bluffs
Remote Sensing 4113 Lab 08: Filtering and Principal Components Mar. 28, 2018 In this lab we will explore Filtering and Principal Components analysis. We will again use the Aster data of the Como Bluffs
Chapter 1 Exercises 1
 Chapter 1 Exercises 1 Data Analysis & Graphics Using R, 2 nd edn Solutions to Selected Exercises (December 15, 2006) Preliminaries > library(daag) Exercise 1 The following table gives the size of the floor
Chapter 1 Exercises 1 Data Analysis & Graphics Using R, 2 nd edn Solutions to Selected Exercises (December 15, 2006) Preliminaries > library(daag) Exercise 1 The following table gives the size of the floor
Enhancement of Speech Signal Based on Improved Minima Controlled Recursive Averaging and Independent Component Analysis
 Enhancement of Speech Signal Based on Improved Minima Controlled Recursive Averaging and Independent Component Analysis Mohini Avatade & S.L. Sahare Electronics & Telecommunication Department, Cummins
Enhancement of Speech Signal Based on Improved Minima Controlled Recursive Averaging and Independent Component Analysis Mohini Avatade & S.L. Sahare Electronics & Telecommunication Department, Cummins
arxiv: v1 [eess.sp] 20 Dec 2017
![arxiv: v1 [eess.sp] 20 Dec 2017 arxiv: v1 [eess.sp] 20 Dec 2017](/thumbs/87/96619770.jpg) GSM CommSense-based through-the-wall sensing Abhishek Bhatta Electrical Engineering Department Amit Kumar Mishra Electrical Engineering Department arxiv:1712.08574v1 [eess.sp] 20 Dec 2017 University of
GSM CommSense-based through-the-wall sensing Abhishek Bhatta Electrical Engineering Department Amit Kumar Mishra Electrical Engineering Department arxiv:1712.08574v1 [eess.sp] 20 Dec 2017 University of
Hyperspectral image processing and analysis
 Hyperspectral image processing and analysis Lecture 12 www.utsa.edu/lrsg/teaching/ees5083/l12-hyper.ppt Multi- vs. Hyper- Hyper-: Narrow bands ( 20 nm in resolution or FWHM) and continuous measurements.
Hyperspectral image processing and analysis Lecture 12 www.utsa.edu/lrsg/teaching/ees5083/l12-hyper.ppt Multi- vs. Hyper- Hyper-: Narrow bands ( 20 nm in resolution or FWHM) and continuous measurements.
Removal of ocular artifacts from EEG signals using adaptive threshold PCA and Wavelet transforms
 Available online at www.interscience.in Removal of ocular artifacts from s using adaptive threshold PCA and Wavelet transforms P. Ashok Babu 1, K.V.S.V.R.Prasad 2 1 Narsimha Reddy Engineering College,
Available online at www.interscience.in Removal of ocular artifacts from s using adaptive threshold PCA and Wavelet transforms P. Ashok Babu 1, K.V.S.V.R.Prasad 2 1 Narsimha Reddy Engineering College,
MULTISPECTRAL IMAGE PROCESSING I
 TM1 TM2 337 TM3 TM4 TM5 TM6 Dr. Robert A. Schowengerdt TM7 Landsat Thematic Mapper (TM) multispectral images of desert and agriculture near Yuma, Arizona MULTISPECTRAL IMAGE PROCESSING I SENSORS Multispectral
TM1 TM2 337 TM3 TM4 TM5 TM6 Dr. Robert A. Schowengerdt TM7 Landsat Thematic Mapper (TM) multispectral images of desert and agriculture near Yuma, Arizona MULTISPECTRAL IMAGE PROCESSING I SENSORS Multispectral
LASER server: ancestry tracing with genotypes or sequence reads
 LASER server: ancestry tracing with genotypes or sequence reads The LASER method Supplementary Data For each ancestry reference panel of N individuals, LASER applies principal components analysis (PCA)
LASER server: ancestry tracing with genotypes or sequence reads The LASER method Supplementary Data For each ancestry reference panel of N individuals, LASER applies principal components analysis (PCA)
Description cabiplot caprojection Remarks and examples References Also see
 Title stata.com ca postestimation plots Postestimation plots for ca and camat cabiplot caprojection Remarks and examples References Also see The following postestimation commands are of special interest
Title stata.com ca postestimation plots Postestimation plots for ca and camat cabiplot caprojection Remarks and examples References Also see The following postestimation commands are of special interest
How can it be right when it feels so wrong? Outliers, diagnostics, non-constant variance
 How can it be right when it feels so wrong? Outliers, diagnostics, non-constant variance D. Alex Hughes November 19, 2014 D. Alex Hughes Problems? November 19, 2014 1 / 61 1 Outliers Generally Residual
How can it be right when it feels so wrong? Outliers, diagnostics, non-constant variance D. Alex Hughes November 19, 2014 D. Alex Hughes Problems? November 19, 2014 1 / 61 1 Outliers Generally Residual
Department of Statistics and Operations Research Undergraduate Programmes
 Department of Statistics and Operations Research Undergraduate Programmes OPERATIONS RESEARCH YEAR LEVEL 2 INTRODUCTION TO LINEAR PROGRAMMING SSOA021 Linear Programming Model: Formulation of an LP model;
Department of Statistics and Operations Research Undergraduate Programmes OPERATIONS RESEARCH YEAR LEVEL 2 INTRODUCTION TO LINEAR PROGRAMMING SSOA021 Linear Programming Model: Formulation of an LP model;
Big Data Framework for Synchrophasor Data Analysis
 Big Data Framework for Synchrophasor Data Analysis Pavel Etingov, Jason Hou, Huiying Ren, Heng Wang, Troy Zuroske, and Dimitri Zarzhitsky Pacific Northwest National Laboratory North American Synchrophasor
Big Data Framework for Synchrophasor Data Analysis Pavel Etingov, Jason Hou, Huiying Ren, Heng Wang, Troy Zuroske, and Dimitri Zarzhitsky Pacific Northwest National Laboratory North American Synchrophasor
Why Should We Care? Everyone uses plotting But most people ignore or are unaware of simple principles Default plotting tools are not always the best
 Elementary Plots Why Should We Care? Everyone uses plotting But most people ignore or are unaware of simple principles Default plotting tools are not always the best More importantly, it is easy to lie
Elementary Plots Why Should We Care? Everyone uses plotting But most people ignore or are unaware of simple principles Default plotting tools are not always the best More importantly, it is easy to lie
SSB Debate: Model-based Inference vs. Machine Learning
 SSB Debate: Model-based nference vs. Machine Learning June 3, 2018 SSB 2018 June 3, 2018 1 / 20 Machine learning in the biological sciences SSB 2018 June 3, 2018 2 / 20 Machine learning in the biological
SSB Debate: Model-based nference vs. Machine Learning June 3, 2018 SSB 2018 June 3, 2018 1 / 20 Machine learning in the biological sciences SSB 2018 June 3, 2018 2 / 20 Machine learning in the biological
Topics for today. Why not use R for graphics? Why use R for graphics? Introduction to R Graphics: U i R t t fi. Using R to create figures
 Topics for today Introduction to R Graphics: U i R t t fi Using R to create figures BaRC Hot Topics October 2011 George Bell, Ph.D. http://iona.wi.mit.edu/bio/education/r2011/ Getting started with R Drawing
Topics for today Introduction to R Graphics: U i R t t fi Using R to create figures BaRC Hot Topics October 2011 George Bell, Ph.D. http://iona.wi.mit.edu/bio/education/r2011/ Getting started with R Drawing
Image analysis. CS/CME/BioE/Biophys/BMI 279 Oct. 31 and Nov. 2, 2017 Ron Dror
 Image analysis CS/CME/BioE/Biophys/BMI 279 Oct. 31 and Nov. 2, 2017 Ron Dror 1 Outline Images in molecular and cellular biology Reducing image noise Mean and Gaussian filters Frequency domain interpretation
Image analysis CS/CME/BioE/Biophys/BMI 279 Oct. 31 and Nov. 2, 2017 Ron Dror 1 Outline Images in molecular and cellular biology Reducing image noise Mean and Gaussian filters Frequency domain interpretation
Why Should We Care? More importantly, it is easy to lie or deceive people with bad plots
 Elementary Plots Why Should We Care? Everyone uses plotting But most people ignore or are unaware of simple principles Default plotting tools (or default settings) are not always the best More importantly,
Elementary Plots Why Should We Care? Everyone uses plotting But most people ignore or are unaware of simple principles Default plotting tools (or default settings) are not always the best More importantly,
Review. In an experiment, there is one variable that is of primary interest. There are several other factors, which may affect the measured result.
 Review Observational study vs experiment Experimental designs In an experiment, there is one variable that is of primary interest. There are several other factors, which may affect the measured result.
Review Observational study vs experiment Experimental designs In an experiment, there is one variable that is of primary interest. There are several other factors, which may affect the measured result.
Multiresolution Analysis of Connectivity
 Multiresolution Analysis of Connectivity Atul Sajjanhar 1, Guojun Lu 2, Dengsheng Zhang 2, Tian Qi 3 1 School of Information Technology Deakin University 221 Burwood Highway Burwood, VIC 3125 Australia
Multiresolution Analysis of Connectivity Atul Sajjanhar 1, Guojun Lu 2, Dengsheng Zhang 2, Tian Qi 3 1 School of Information Technology Deakin University 221 Burwood Highway Burwood, VIC 3125 Australia
A new edited k-nearest neighbor rule in the pattern classi"cation problem
 Pattern Recognition 33 (2000) 521}528 A new edited -nearest neighbor rule in the pattern classi"cation problem Kazuo Hattori*, Masahito Taahashi Department of Electrical Engineering and Electronics, Toyohashi
Pattern Recognition 33 (2000) 521}528 A new edited -nearest neighbor rule in the pattern classi"cation problem Kazuo Hattori*, Masahito Taahashi Department of Electrical Engineering and Electronics, Toyohashi
(3 pts) 1. Which statements are usually true of a left-skewed distribution? (circle all that are correct)
 STAT 451 - Practice Exam I Name (print): Section: This is a practice exam - it s a representative sample of problems that may appear on the exam and also substantially longer than the in-class exam. It
STAT 451 - Practice Exam I Name (print): Section: This is a practice exam - it s a representative sample of problems that may appear on the exam and also substantially longer than the in-class exam. It
Assessing Measurement System Variation
 Example 1 Fuel Injector Nozzle Diameters Problem A manufacturer of fuel injector nozzles has installed a new digital measuring system. Investigators want to determine how well the new system measures the
Example 1 Fuel Injector Nozzle Diameters Problem A manufacturer of fuel injector nozzles has installed a new digital measuring system. Investigators want to determine how well the new system measures the
Texture characterization in DIRSIG
 Rochester Institute of Technology RIT Scholar Works Theses Thesis/Dissertation Collections 2001 Texture characterization in DIRSIG Christy Burtner Follow this and additional works at: http://scholarworks.rit.edu/theses
Rochester Institute of Technology RIT Scholar Works Theses Thesis/Dissertation Collections 2001 Texture characterization in DIRSIG Christy Burtner Follow this and additional works at: http://scholarworks.rit.edu/theses
GE 113 REMOTE SENSING
 GE 113 REMOTE SENSING Topic 8. Image Classification and Accuracy Assessment Lecturer: Engr. Jojene R. Santillan jrsantillan@carsu.edu.ph Division of Geodetic Engineering College of Engineering and Information
GE 113 REMOTE SENSING Topic 8. Image Classification and Accuracy Assessment Lecturer: Engr. Jojene R. Santillan jrsantillan@carsu.edu.ph Division of Geodetic Engineering College of Engineering and Information
Efficient Signal Identification using the Spectral Correlation Function and Pattern Recognition
 Efficient Signal Identification using the Spectral Correlation Function and Pattern Recognition Theodore Trebaol, Jeffrey Dunn, and Daniel D. Stancil Acknowledgement: J. Peha, M. Sirbu, P. Steenkiste Outline
Efficient Signal Identification using the Spectral Correlation Function and Pattern Recognition Theodore Trebaol, Jeffrey Dunn, and Daniel D. Stancil Acknowledgement: J. Peha, M. Sirbu, P. Steenkiste Outline
Spring 2017 Math 54 Test #2 Name:
 Spring 2017 Math 54 Test #2 Name: You may use a TI calculator and formula sheets from the textbook. Show your work neatly and systematically for full credit. Total points: 101 1. (6) Suppose P(E) = 0.37
Spring 2017 Math 54 Test #2 Name: You may use a TI calculator and formula sheets from the textbook. Show your work neatly and systematically for full credit. Total points: 101 1. (6) Suppose P(E) = 0.37
Package plotpc. September 27, Index 10. Plot principal component loadings
 Version 1.0.4 Package plotpc September 27, 2015 Title Plot Principal Component Histograms Around a Scatter Plot Author Stephen Milborrow Maintainer Stephen Milborrow Depends grid Description
Version 1.0.4 Package plotpc September 27, 2015 Title Plot Principal Component Histograms Around a Scatter Plot Author Stephen Milborrow Maintainer Stephen Milborrow Depends grid Description
RECENT developments have seen lot of power system
 Auto Detection of Power System Events Using Wide Area Frequency Measurements Gopal Gajjar and S. A. Soman Dept. of Electrical Engineering, Indian Institute of Technology Bombay, India 476 Email: gopalgajjar@ieee.org
Auto Detection of Power System Events Using Wide Area Frequency Measurements Gopal Gajjar and S. A. Soman Dept. of Electrical Engineering, Indian Institute of Technology Bombay, India 476 Email: gopalgajjar@ieee.org
Assessing Measurement System Variation
 Assessing Measurement System Variation Example 1: Fuel Injector Nozzle Diameters Problem A manufacturer of fuel injector nozzles installs a new digital measuring system. Investigators want to determine
Assessing Measurement System Variation Example 1: Fuel Injector Nozzle Diameters Problem A manufacturer of fuel injector nozzles installs a new digital measuring system. Investigators want to determine
Dependence in Classification of Aluminium Waste
 Journal of Physics: Conference Series PAPER OPEN ACCESS Dependence in Classification of Aluminium Waste To cite this article: Y Resti 05 J. Phys.: Conf. Ser. 6 005 Recent citations - A probability approach
Journal of Physics: Conference Series PAPER OPEN ACCESS Dependence in Classification of Aluminium Waste To cite this article: Y Resti 05 J. Phys.: Conf. Ser. 6 005 Recent citations - A probability approach
Image analysis. CS/CME/BIOPHYS/BMI 279 Fall 2015 Ron Dror
 Image analysis CS/CME/BIOPHYS/BMI 279 Fall 2015 Ron Dror A two- dimensional image can be described as a function of two variables f(x,y). For a grayscale image, the value of f(x,y) specifies the brightness
Image analysis CS/CME/BIOPHYS/BMI 279 Fall 2015 Ron Dror A two- dimensional image can be described as a function of two variables f(x,y). For a grayscale image, the value of f(x,y) specifies the brightness
Contents. List of Figures List of Tables. Structure of the Book How to Use this Book Online Resources Acknowledgements
 Contents List of Figures List of Tables Preface Notation Structure of the Book How to Use this Book Online Resources Acknowledgements Notational Conventions Notational Conventions for Probabilities xiii
Contents List of Figures List of Tables Preface Notation Structure of the Book How to Use this Book Online Resources Acknowledgements Notational Conventions Notational Conventions for Probabilities xiii
INTERACTIVE DATA VISUALIZATION WITH BOKEH. Interactive Data Visualization with Bokeh
 INTERACTIVE DATA VISUALIZATION WITH BOKEH Interactive Data Visualization with Bokeh What is Bokeh? Interactive visualization, controls, and tools Versatile and high-level graphics High-level statistical
INTERACTIVE DATA VISUALIZATION WITH BOKEH Interactive Data Visualization with Bokeh What is Bokeh? Interactive visualization, controls, and tools Versatile and high-level graphics High-level statistical
4 Exploration. 4.1 Data exploration using R tools
 4 Exploration The statistical background of all methods discussed in this chapter can be found Analysing Ecological Data by Zuur, Ieno and Smith (2007). Here, we only discuss how to apply the methods in
4 Exploration The statistical background of all methods discussed in this chapter can be found Analysing Ecological Data by Zuur, Ieno and Smith (2007). Here, we only discuss how to apply the methods in
Voice Activity Detection
 Voice Activity Detection Speech Processing Tom Bäckström Aalto University October 2015 Introduction Voice activity detection (VAD) (or speech activity detection, or speech detection) refers to a class
Voice Activity Detection Speech Processing Tom Bäckström Aalto University October 2015 Introduction Voice activity detection (VAD) (or speech activity detection, or speech detection) refers to a class
Remote Sensing Instruction Laboratory
 Laboratory Session 217513 Geographic Information System and Remote Sensing - 1 - Remote Sensing Instruction Laboratory Assist.Prof.Dr. Weerakaset Suanpaga Department of Civil Engineering, Faculty of Engineering
Laboratory Session 217513 Geographic Information System and Remote Sensing - 1 - Remote Sensing Instruction Laboratory Assist.Prof.Dr. Weerakaset Suanpaga Department of Civil Engineering, Faculty of Engineering
Learning Dota 2 Team Compositions
 Learning Dota 2 Team Compositions Atish Agarwala atisha@stanford.edu Michael Pearce pearcemt@stanford.edu Abstract Dota 2 is a multiplayer online game in which two teams of five players control heroes
Learning Dota 2 Team Compositions Atish Agarwala atisha@stanford.edu Michael Pearce pearcemt@stanford.edu Abstract Dota 2 is a multiplayer online game in which two teams of five players control heroes
SPEECH ENHANCEMENT WITH SIGNAL SUBSPACE FILTER BASED ON PERCEPTUAL POST FILTERING
 SPEECH ENHANCEMENT WITH SIGNAL SUBSPACE FILTER BASED ON PERCEPTUAL POST FILTERING K.Ramalakshmi Assistant Professor, Dept of CSE Sri Ramakrishna Institute of Technology, Coimbatore R.N.Devendra Kumar Assistant
SPEECH ENHANCEMENT WITH SIGNAL SUBSPACE FILTER BASED ON PERCEPTUAL POST FILTERING K.Ramalakshmi Assistant Professor, Dept of CSE Sri Ramakrishna Institute of Technology, Coimbatore R.N.Devendra Kumar Assistant
Image interpretation and analysis
 Image interpretation and analysis Grundlagen Fernerkundung, Geo 123.1, FS 2014 Lecture 7a Rogier de Jong Michael Schaepman Why are snow, foam, and clouds white? Why are snow, foam, and clouds white? Today
Image interpretation and analysis Grundlagen Fernerkundung, Geo 123.1, FS 2014 Lecture 7a Rogier de Jong Michael Schaepman Why are snow, foam, and clouds white? Why are snow, foam, and clouds white? Today
Discussion of The power of monitoring: how to make the most of a contaminated multivariate sample
 Stat Methods Appl https://doi.org/.7/s-7-- COMMENT Discussion of The power of monitoring: how to make the most of a contaminated multivariate sample Domenico Perrotta Francesca Torti Accepted: December
Stat Methods Appl https://doi.org/.7/s-7-- COMMENT Discussion of The power of monitoring: how to make the most of a contaminated multivariate sample Domenico Perrotta Francesca Torti Accepted: December
Hyperspectral Image Data
 CEE 615: Digital Image Processing Lab 11: Hyperspectral Noise p. 1 Hyperspectral Image Data Files needed for this exercise (all are standard ENVI files): Images: cup95eff.int &.hdr Spectral Library: jpl1.sli
CEE 615: Digital Image Processing Lab 11: Hyperspectral Noise p. 1 Hyperspectral Image Data Files needed for this exercise (all are standard ENVI files): Images: cup95eff.int &.hdr Spectral Library: jpl1.sli
Biology 559R: Introduction to Phylogenetic Comparative Methods Topics for this week (Feb 3 & 5):
 Biology 559R: Introduction to Phylogenetic Comparative Methods Topics for this week (Feb 3 & 5): Chronogram estimation: Penalized Likelihood Approach BEAST Presentations of your projects 1 The Anatomy
Biology 559R: Introduction to Phylogenetic Comparative Methods Topics for this week (Feb 3 & 5): Chronogram estimation: Penalized Likelihood Approach BEAST Presentations of your projects 1 The Anatomy
Geostatistical estimation applied to highly skewed data. Dr. Isobel Clark, Geostokos Limited, Alloa, Scotland
 "Geostatistical estimation applied to highly skewed data", Joint Statistical Meetings, Dallas, Texas, August 1999 Geostatistical estimation applied to highly skewed data Dr. Isobel Clark, Geostokos Limited,
"Geostatistical estimation applied to highly skewed data", Joint Statistical Meetings, Dallas, Texas, August 1999 Geostatistical estimation applied to highly skewed data Dr. Isobel Clark, Geostokos Limited,
3 Selecting the standard map and area of interest
 Anomalies, EOF/PCA with WAM Mati Kahru 2005-2009 1 Anomalies, EOF/PC analysis with WAM 1 Introduction Calculating anomalies is a powerful method of change detection in time series. Empirical Orthogonal
Anomalies, EOF/PCA with WAM Mati Kahru 2005-2009 1 Anomalies, EOF/PC analysis with WAM 1 Introduction Calculating anomalies is a powerful method of change detection in time series. Empirical Orthogonal
Simplifying the Art of Terahertz Measurements
 Simplifying the Art of Terahertz Measurements Achieving metrology-level accuracy with a manual probe system With significant expansion of emerging THz applications, such as non-invasive spectroscopy, security
Simplifying the Art of Terahertz Measurements Achieving metrology-level accuracy with a manual probe system With significant expansion of emerging THz applications, such as non-invasive spectroscopy, security
UNIVERSITETET FOR MILJØ- OG BIOVITSKAP
 UNIVERSITETET FOR MILJØ- OG BIOVITSKAP 1 Photo: Ingunn Nævdal http://www.nsg.no/ind ex.cfm?id= 53192 MILK QUALITY BREEDING VALUE PREDICTION BASED ON FTIR SPECTRA Tormod ÅDNØY, Theo ME MEUWISSEN, Binyamin
UNIVERSITETET FOR MILJØ- OG BIOVITSKAP 1 Photo: Ingunn Nævdal http://www.nsg.no/ind ex.cfm?id= 53192 MILK QUALITY BREEDING VALUE PREDICTION BASED ON FTIR SPECTRA Tormod ÅDNØY, Theo ME MEUWISSEN, Binyamin
Hand & Upper Body Based Hybrid Gesture Recognition
 Hand & Upper Body Based Hybrid Gesture Prerna Sharma #1, Naman Sharma *2 # Research Scholor, G. B. P. U. A. & T. Pantnagar, India * Ideal Institue of Technology, Ghaziabad, India Abstract Communication
Hand & Upper Body Based Hybrid Gesture Prerna Sharma #1, Naman Sharma *2 # Research Scholor, G. B. P. U. A. & T. Pantnagar, India * Ideal Institue of Technology, Ghaziabad, India Abstract Communication
Lab 8. Signal Analysis Using Matlab Simulink
 E E 2 7 5 Lab June 30, 2006 Lab 8. Signal Analysis Using Matlab Simulink Introduction The Matlab Simulink software allows you to model digital signals, examine power spectra of digital signals, represent
E E 2 7 5 Lab June 30, 2006 Lab 8. Signal Analysis Using Matlab Simulink Introduction The Matlab Simulink software allows you to model digital signals, examine power spectra of digital signals, represent
Drum Transcription Based on Independent Subspace Analysis
 Report for EE 391 Special Studies and Reports for Electrical Engineering Drum Transcription Based on Independent Subspace Analysis Yinyi Guo Center for Computer Research in Music and Acoustics, Stanford,
Report for EE 391 Special Studies and Reports for Electrical Engineering Drum Transcription Based on Independent Subspace Analysis Yinyi Guo Center for Computer Research in Music and Acoustics, Stanford,
Feature analysis of EEG signals using SOM
 1 Portál pre odborné publikovanie ISSN 1338-0087 Feature analysis of EEG signals using SOM Gráfová Lucie Elektrotechnika, Medicína 21.02.2011 The most common use of EEG includes the monitoring and diagnosis
1 Portál pre odborné publikovanie ISSN 1338-0087 Feature analysis of EEG signals using SOM Gráfová Lucie Elektrotechnika, Medicína 21.02.2011 The most common use of EEG includes the monitoring and diagnosis
ENVI Classic Tutorial: Spectral Angle Mapper (SAM) and Spectral Information Divergence (SID) Classification 2
 ENVI Classic Tutorial: Spectral Angle Mapper (SAM) and Spectral Information Divergence (SID) Classification Spectral Angle Mapper (SAM) and Spectral Information Divergence (SID) Classification 2 Files
ENVI Classic Tutorial: Spectral Angle Mapper (SAM) and Spectral Information Divergence (SID) Classification Spectral Angle Mapper (SAM) and Spectral Information Divergence (SID) Classification 2 Files
R Short Course Session 3
 R Short Course Session 3 Daniel Zhao, PhD Sixia Chen, PhD Department of Biostatistics and Epidemiology College of Public Health, OUHSC 11/6/2015 Scatter plot QQ plot Histogram Curve Bar chart Pie chart
R Short Course Session 3 Daniel Zhao, PhD Sixia Chen, PhD Department of Biostatistics and Epidemiology College of Public Health, OUHSC 11/6/2015 Scatter plot QQ plot Histogram Curve Bar chart Pie chart
Introduction to ibbig
 Introduction to ibbig Aedin Culhane, Daniel Gusenleitner April 4, 2013 1 ibbig Iterative Binary Bi-clustering of Gene sets (ibbig) is a bi-clustering algorithm optimized for discovery of overlapping biclusters
Introduction to ibbig Aedin Culhane, Daniel Gusenleitner April 4, 2013 1 ibbig Iterative Binary Bi-clustering of Gene sets (ibbig) is a bi-clustering algorithm optimized for discovery of overlapping biclusters
The study of human populations involves working not PART 2. Cemetery Investigation: An Exercise in Simple Statistics POPULATIONS
 PART 2 POPULATIONS Cemetery Investigation: An Exercise in Simple Statistics 4 When you have completed this exercise, you will be able to: 1. Work effectively with data that must be organized in a useful
PART 2 POPULATIONS Cemetery Investigation: An Exercise in Simple Statistics 4 When you have completed this exercise, you will be able to: 1. Work effectively with data that must be organized in a useful
Super-Resolution of Multispectral Images
 IJSRD - International Journal for Scientific Research & Development Vol. 1, Issue 3, 2013 ISSN (online): 2321-0613 Super-Resolution of Images Mr. Dhaval Shingala 1 Ms. Rashmi Agrawal 2 1 PG Student, Computer
IJSRD - International Journal for Scientific Research & Development Vol. 1, Issue 3, 2013 ISSN (online): 2321-0613 Super-Resolution of Images Mr. Dhaval Shingala 1 Ms. Rashmi Agrawal 2 1 PG Student, Computer
COMPARATIVE PERFORMANCE ANALYSIS OF HAND GESTURE RECOGNITION TECHNIQUES
 International Journal of Advanced Research in Engineering and Technology (IJARET) Volume 9, Issue 3, May - June 2018, pp. 177 185, Article ID: IJARET_09_03_023 Available online at http://www.iaeme.com/ijaret/issues.asp?jtype=ijaret&vtype=9&itype=3
International Journal of Advanced Research in Engineering and Technology (IJARET) Volume 9, Issue 3, May - June 2018, pp. 177 185, Article ID: IJARET_09_03_023 Available online at http://www.iaeme.com/ijaret/issues.asp?jtype=ijaret&vtype=9&itype=3
HYPERSPECTRAL IMAGE DATA MINING FOR BAND SELECTION IN AGRICULTURAL APPLICATIONS
 HYPERSPECTRAL IMAGE DATA MINING FOR BAND SELECTION IN AGRICULTURAL APPLICATIONS S. G. Bajwa, P. Bajcsy, P. Groves, L. F. Tian ABSTRACT. Hyperspectral remote sensing produces large volumes of data, quite
HYPERSPECTRAL IMAGE DATA MINING FOR BAND SELECTION IN AGRICULTURAL APPLICATIONS S. G. Bajwa, P. Bajcsy, P. Groves, L. F. Tian ABSTRACT. Hyperspectral remote sensing produces large volumes of data, quite
Unit Nine Precalculus Practice Test Probability & Statistics. Name: Period: Date: NON-CALCULATOR SECTION
 Name: Period: Date: NON-CALCULATOR SECTION Vocabulary: Define each word and give an example. 1. discrete mathematics 2. dependent outcomes 3. series Short Answer: 4. Describe when to use a combination.
Name: Period: Date: NON-CALCULATOR SECTION Vocabulary: Define each word and give an example. 1. discrete mathematics 2. dependent outcomes 3. series Short Answer: 4. Describe when to use a combination.
Variance and Anomaly Analysis with WIM/WAM Mati Kahru
 Variance and Anomaly Analysis with WIM/WAM Mati Kahru 2008 1 Variance and Anomaly Analysis with WIM/WAM 1 Introduction Analysis of temporal variance of image data provides important clues on the functioning
Variance and Anomaly Analysis with WIM/WAM Mati Kahru 2008 1 Variance and Anomaly Analysis with WIM/WAM 1 Introduction Analysis of temporal variance of image data provides important clues on the functioning
Repeated Measures Twoway Analysis of Variance
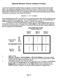 Repeated Measures Twoway Analysis of Variance A researcher was interested in whether frequency of exposure to a picture of an ugly or attractive person would influence one's liking for the photograph.
Repeated Measures Twoway Analysis of Variance A researcher was interested in whether frequency of exposure to a picture of an ugly or attractive person would influence one's liking for the photograph.
EXST 7037 Multivariate Analysis Factor Analysis (SASy version) Page 1
 EXST 7037 Multivariate Analysis Factor Analysis (SASy version) Page 1 1 *** CH05SD ***; 2 *****************************************************************************; 3 *** The Second International Math
EXST 7037 Multivariate Analysis Factor Analysis (SASy version) Page 1 1 *** CH05SD ***; 2 *****************************************************************************; 3 *** The Second International Math
Colour image watermarking in real life
 Colour image watermarking in real life Konstantin Krasavin University of Joensuu, Finland ABSTRACT: In this report we present our work for colour image watermarking in different domains. First we consider
Colour image watermarking in real life Konstantin Krasavin University of Joensuu, Finland ABSTRACT: In this report we present our work for colour image watermarking in different domains. First we consider
Learning Some Simple Plotting Features of R 15
 Learning Some Simple Plotting Features of R 15 This independent exercise will help you learn how R plotting functions work. This activity focuses on how you might use graphics to help you interpret large
Learning Some Simple Plotting Features of R 15 This independent exercise will help you learn how R plotting functions work. This activity focuses on how you might use graphics to help you interpret large
Marie-France OUDIN, Denys CHAUME INTRODUCTION
 XIV International congress of th~! S.I.P. -Hambourg 1980- AN A1.Q FOR IHE HITERrRETATIDN Marie-France OUDIN, Denys CHAUME Scientific Center I.B.M. France / 36 Avenue Raymond Poincare -PARIS 75016- ++-f+++++
XIV International congress of th~! S.I.P. -Hambourg 1980- AN A1.Q FOR IHE HITERrRETATIDN Marie-France OUDIN, Denys CHAUME Scientific Center I.B.M. France / 36 Avenue Raymond Poincare -PARIS 75016- ++-f+++++
A Closed Form for False Location Injection under Time Difference of Arrival
 A Closed Form for False Location Injection under Time Difference of Arrival Lauren M. Huie Mark L. Fowler lauren.huie@rl.af.mil mfowler@binghamton.edu Air Force Research Laboratory, Rome, N Department
A Closed Form for False Location Injection under Time Difference of Arrival Lauren M. Huie Mark L. Fowler lauren.huie@rl.af.mil mfowler@binghamton.edu Air Force Research Laboratory, Rome, N Department
CHEMOMETRICS IN SPECTROSCOPY Part 27: Linearity in Calibration
 This column was originally published in Spectroscopy, 13(6), p. 19-21 (1998) CHEMOMETRICS IN SPECTROSCOPY Part 27: Linearity in Calibration by Howard Mark and Jerome Workman Those who know us know that
This column was originally published in Spectroscopy, 13(6), p. 19-21 (1998) CHEMOMETRICS IN SPECTROSCOPY Part 27: Linearity in Calibration by Howard Mark and Jerome Workman Those who know us know that
Global Journal of Engineering Science and Research Management
 A KERNEL BASED APPROACH: USING MOVIE SCRIPT FOR ASSESSING BOX OFFICE PERFORMANCE Mr.K.R. Dabhade *1 Ms. S.S. Ponde 2 *1 Computer Science Department. D.I.E.M.S. 2 Asst. Prof. Computer Science Department,
A KERNEL BASED APPROACH: USING MOVIE SCRIPT FOR ASSESSING BOX OFFICE PERFORMANCE Mr.K.R. Dabhade *1 Ms. S.S. Ponde 2 *1 Computer Science Department. D.I.E.M.S. 2 Asst. Prof. Computer Science Department,
Steps involved in microarray analysis after the experiments
 Steps involved in microarray analysis after the experiments Scanning slides to create images Conversion of images to numerical data Processing of raw numerical data Further analysis Clustering Integration
Steps involved in microarray analysis after the experiments Scanning slides to create images Conversion of images to numerical data Processing of raw numerical data Further analysis Clustering Integration
Introduction to ibbig
 Introduction to ibbig Aedin Culhane, Daniel Gusenleitner June 13, 2018 1 ibbig Iterative Binary Bi-clustering of Gene sets (ibbig) is a bi-clustering algorithm optimized for discovery of overlapping biclusters
Introduction to ibbig Aedin Culhane, Daniel Gusenleitner June 13, 2018 1 ibbig Iterative Binary Bi-clustering of Gene sets (ibbig) is a bi-clustering algorithm optimized for discovery of overlapping biclusters
Color appearance in image displays
 Rochester Institute of Technology RIT Scholar Works Presentations and other scholarship 1-18-25 Color appearance in image displays Mark Fairchild Follow this and additional works at: http://scholarworks.rit.edu/other
Rochester Institute of Technology RIT Scholar Works Presentations and other scholarship 1-18-25 Color appearance in image displays Mark Fairchild Follow this and additional works at: http://scholarworks.rit.edu/other
Data 1 Assessment Calculator allowed for all questions
 Foundation Higher Data Assessment Calculator allowed for all questions MATHSWATCH All questions Time for the test: 45 minutes Name: MATHSWATCH ANSWERS Grade Title of clip Marks Score Percentage Clip 4
Foundation Higher Data Assessment Calculator allowed for all questions MATHSWATCH All questions Time for the test: 45 minutes Name: MATHSWATCH ANSWERS Grade Title of clip Marks Score Percentage Clip 4
1990 Census Measures. Fast Track Project Technical Report Patrick S. Malone ( ; 9-May-00
 1990 Census Measures Fast Track Project Technical Report Patrick S. Malone (919-668-6910; malone@alumni.duke.edu) 9-May-00 Table of Contents I. Scale Description II. Report Sample III. Scaling IV. Differences
1990 Census Measures Fast Track Project Technical Report Patrick S. Malone (919-668-6910; malone@alumni.duke.edu) 9-May-00 Table of Contents I. Scale Description II. Report Sample III. Scaling IV. Differences
Recommender Systems TIETS43 Collaborative Filtering
 + Recommender Systems TIETS43 Collaborative Filtering Fall 2017 Kostas Stefanidis kostas.stefanidis@uta.fi https://coursepages.uta.fi/tiets43/ selection Amazon generates 35% of their sales through recommendations
+ Recommender Systems TIETS43 Collaborative Filtering Fall 2017 Kostas Stefanidis kostas.stefanidis@uta.fi https://coursepages.uta.fi/tiets43/ selection Amazon generates 35% of their sales through recommendations
GEOG432: Remote sensing Lab 3 Unsupervised classification
 GEOG432: Remote sensing Lab 3 Unsupervised classification Goal: This lab involves identifying land cover types by using agorithms to identify pixels with similar Digital Numbers (DN) and spectral signatures
GEOG432: Remote sensing Lab 3 Unsupervised classification Goal: This lab involves identifying land cover types by using agorithms to identify pixels with similar Digital Numbers (DN) and spectral signatures
Homework Assignment (20 points): MORPHOMETRICS (Bivariate and Multivariate Analyses)
 Fossils and Evolution Due: Tuesday, Jan. 31 Spring 2012 Homework Assignment (20 points): MORPHOMETRICS (Bivariate and Multivariate Analyses) Introduction Morphometrics is the use of measurements to assess
Fossils and Evolution Due: Tuesday, Jan. 31 Spring 2012 Homework Assignment (20 points): MORPHOMETRICS (Bivariate and Multivariate Analyses) Introduction Morphometrics is the use of measurements to assess
Emitter Location in the Presence of Information Injection
 in the Presence of Information Injection Lauren M. Huie Mark L. Fowler lauren.huie@rl.af.mil mfowler@binghamton.edu Air Force Research Laboratory, Rome, N.Y. State University of New York at Binghamton,
in the Presence of Information Injection Lauren M. Huie Mark L. Fowler lauren.huie@rl.af.mil mfowler@binghamton.edu Air Force Research Laboratory, Rome, N.Y. State University of New York at Binghamton,
Projecting Fantasy Football Points
 Projecting Fantasy Football Points Brian Becker Gary Ramirez Carlos Zambrano MATH 503 A/B October 12, 2015 1 1 Abstract Fantasy Football has been increasing in popularity throughout the years and becoming
Projecting Fantasy Football Points Brian Becker Gary Ramirez Carlos Zambrano MATH 503 A/B October 12, 2015 1 1 Abstract Fantasy Football has been increasing in popularity throughout the years and becoming
BASIC PATTERN RECOGNITION AND DIGITAL IMAGE PROCESSING USING
 BASIC PATTERN RECOGNITION AND DIGITAL IMAGE PROCESSING USING SAS/AF FRAME Abhishek Lall Department of Mathematics and Statistics, Sam Houston State University, Huntsville, Texas Abstract The principal
BASIC PATTERN RECOGNITION AND DIGITAL IMAGE PROCESSING USING SAS/AF FRAME Abhishek Lall Department of Mathematics and Statistics, Sam Houston State University, Huntsville, Texas Abstract The principal
Image Enhancement using Image Fusion
 Image Enhancement using Image Fusion Ajinkya A. Jadhav Student,ME(Electronics &Telecommunication) Mr. S. R. Khot Associate Professor, Department of Electronics, Mrs. P. S. Pise Associate Professor, Department
Image Enhancement using Image Fusion Ajinkya A. Jadhav Student,ME(Electronics &Telecommunication) Mr. S. R. Khot Associate Professor, Department of Electronics, Mrs. P. S. Pise Associate Professor, Department
Community Detection and Labeling Nodes
 and Labeling Nodes Hao Chen Department of Statistics, Stanford Jan. 25, 2011 (Department of Statistics, Stanford) Community Detection and Labeling Nodes Jan. 25, 2011 1 / 9 Community Detection - Network:
and Labeling Nodes Hao Chen Department of Statistics, Stanford Jan. 25, 2011 (Department of Statistics, Stanford) Community Detection and Labeling Nodes Jan. 25, 2011 1 / 9 Community Detection - Network:
DURING the past several years, independent component
 912 IEEE TRANSACTIONS ON NEURAL NETWORKS, VOL. 10, NO. 4, JULY 1999 Principal Independent Component Analysis Jie Luo, Bo Hu, Xie-Ting Ling, Ruey-Wen Liu Abstract Conventional blind signal separation algorithms
912 IEEE TRANSACTIONS ON NEURAL NETWORKS, VOL. 10, NO. 4, JULY 1999 Principal Independent Component Analysis Jie Luo, Bo Hu, Xie-Ting Ling, Ruey-Wen Liu Abstract Conventional blind signal separation algorithms
FEASIBILITY STUDY OF PHOTOPLETHYSMOGRAPHIC SIGNALS FOR BIOMETRIC IDENTIFICATION. Petros Spachos, Jiexin Gao and Dimitrios Hatzinakos
 FEASIBILITY STUDY OF PHOTOPLETHYSMOGRAPHIC SIGNALS FOR BIOMETRIC IDENTIFICATION Petros Spachos, Jiexin Gao and Dimitrios Hatzinakos The Edward S. Rogers Sr. Department of Electrical and Computer Engineering,
FEASIBILITY STUDY OF PHOTOPLETHYSMOGRAPHIC SIGNALS FOR BIOMETRIC IDENTIFICATION Petros Spachos, Jiexin Gao and Dimitrios Hatzinakos The Edward S. Rogers Sr. Department of Electrical and Computer Engineering,
Automobile Independent Fault Detection based on Acoustic Emission Using FFT
 SINCE2011 Singapore International NDT Conference & Exhibition, 3-4 November 2011 Automobile Independent Fault Detection based on Acoustic Emission Using FFT Hamid GHADERI 1, Peyman KABIRI 2 1 Intelligent
SINCE2011 Singapore International NDT Conference & Exhibition, 3-4 November 2011 Automobile Independent Fault Detection based on Acoustic Emission Using FFT Hamid GHADERI 1, Peyman KABIRI 2 1 Intelligent
Privacy preserving data mining multiplicative perturbation techniques
 Privacy preserving data mining multiplicative perturbation techniques Li Xiong CS573 Data Privacy and Anonymity Outline Review and critique of randomization approaches (additive noise) Multiplicative data
Privacy preserving data mining multiplicative perturbation techniques Li Xiong CS573 Data Privacy and Anonymity Outline Review and critique of randomization approaches (additive noise) Multiplicative data
Scatter Plots, Correlation, and Lines of Best Fit
 Lesson 7.3 Objectives Interpret a scatter plot. Identify the correlation of data from a scatter plot. Find the line of best fit for a set of data. Scatter Plots, Correlation, and Lines of Best Fit A video
Lesson 7.3 Objectives Interpret a scatter plot. Identify the correlation of data from a scatter plot. Find the line of best fit for a set of data. Scatter Plots, Correlation, and Lines of Best Fit A video
ELEC E7210: Communication Theory. Lecture 11: MIMO Systems and Space-time Communications
 ELEC E7210: Communication Theory Lecture 11: MIMO Systems and Space-time Communications Overview of the last lecture MIMO systems -parallel decomposition; - beamforming; - MIMO channel capacity MIMO Key
ELEC E7210: Communication Theory Lecture 11: MIMO Systems and Space-time Communications Overview of the last lecture MIMO systems -parallel decomposition; - beamforming; - MIMO channel capacity MIMO Key
From Morphological Box to Multidimensional Datascapes
 From Morphological Box to Multidimensional Datascapes S. George Center for Data-Driven Discovery and Dept. of Astronomy, Caltech AstroInformatics 2016, Sorrento, Italy, October 2016 Big Data is like teenage
From Morphological Box to Multidimensional Datascapes S. George Center for Data-Driven Discovery and Dept. of Astronomy, Caltech AstroInformatics 2016, Sorrento, Italy, October 2016 Big Data is like teenage
Augment the Spatial Resolution of Multispectral Image Using PCA Fusion Method and Classified It s Region Using Different Techniques.
 Augment the Spatial Resolution of Multispectral Image Using PCA Fusion Method and Classified It s Region Using Different Techniques. Israa Jameel Muhsin 1, Khalid Hassan Salih 2, Ebtesam Fadhel 3 1,2 Department
Augment the Spatial Resolution of Multispectral Image Using PCA Fusion Method and Classified It s Region Using Different Techniques. Israa Jameel Muhsin 1, Khalid Hassan Salih 2, Ebtesam Fadhel 3 1,2 Department
CSC 320 H1S CSC320 Exam Study Guide (Last updated: April 2, 2015) Winter 2015
 Question 1. Suppose you have an image I that contains an image of a left eye (the image is detailed enough that it makes a difference that it s the left eye). Write pseudocode to find other left eyes in
Question 1. Suppose you have an image I that contains an image of a left eye (the image is detailed enough that it makes a difference that it s the left eye). Write pseudocode to find other left eyes in
Auto-tagging The Facebook
 Auto-tagging The Facebook Jonathan Michelson and Jorge Ortiz Stanford University 2006 E-mail: JonMich@Stanford.edu, jorge.ortiz@stanford.com Introduction For those not familiar, The Facebook is an extremely
Auto-tagging The Facebook Jonathan Michelson and Jorge Ortiz Stanford University 2006 E-mail: JonMich@Stanford.edu, jorge.ortiz@stanford.com Introduction For those not familiar, The Facebook is an extremely
How to Use the Method of Multivariate Statistical Analysis Into the Equipment State Monitoring. Chunhua Yang
 4th International Conference on Mechatronics, Materials, Chemistry and Computer Engineering (ICMMCCE 205) How to Use the Method of Multivariate Statistical Analysis Into the Equipment State Monitoring
4th International Conference on Mechatronics, Materials, Chemistry and Computer Engineering (ICMMCCE 205) How to Use the Method of Multivariate Statistical Analysis Into the Equipment State Monitoring
SIGNAL PROCESSING OF POWER QUALITY DISTURBANCES
 SIGNAL PROCESSING OF POWER QUALITY DISTURBANCES MATH H. J. BOLLEN IRENE YU-HUA GU IEEE PRESS SERIES I 0N POWER ENGINEERING IEEE PRESS SERIES ON POWER ENGINEERING MOHAMED E. EL-HAWARY, SERIES EDITOR IEEE
SIGNAL PROCESSING OF POWER QUALITY DISTURBANCES MATH H. J. BOLLEN IRENE YU-HUA GU IEEE PRESS SERIES I 0N POWER ENGINEERING IEEE PRESS SERIES ON POWER ENGINEERING MOHAMED E. EL-HAWARY, SERIES EDITOR IEEE
F2 - Fire 2 module: Remote Sensing Data Classification
 F2 - Fire 2 module: Remote Sensing Data Classification F2.1 Task_1: Supervised and Unsupervised classification examples of a Landsat 5 TM image from the Center of Portugal, year 2005 F2.1 Task_2: Burnt
F2 - Fire 2 module: Remote Sensing Data Classification F2.1 Task_1: Supervised and Unsupervised classification examples of a Landsat 5 TM image from the Center of Portugal, year 2005 F2.1 Task_2: Burnt
Instruction Manual. Mark Deimund, Zuyi (Jacky) Huang, Juergen Hahn
 Instruction Manual Mark Deimund, Zuyi (Jacky) Huang, Juergen Hahn This manual is for the program that implements the image analysis method presented in our paper: Z. Huang, F. Senocak, A. Jayaraman, and
Instruction Manual Mark Deimund, Zuyi (Jacky) Huang, Juergen Hahn This manual is for the program that implements the image analysis method presented in our paper: Z. Huang, F. Senocak, A. Jayaraman, and
6. Multivariate EDA. ACE 492 SA - Spatial Analysis Fall 2003
 1 Objectives 6. Multivariate EDA ACE 492 SA - Spatial Analysis Fall 2003 c 2003 by Luc Anselin, All Rights Reserved This lab covers some basic approaches to carry out EDA with a focus on discovering multivariate
1 Objectives 6. Multivariate EDA ACE 492 SA - Spatial Analysis Fall 2003 c 2003 by Luc Anselin, All Rights Reserved This lab covers some basic approaches to carry out EDA with a focus on discovering multivariate
