Procedure & Checklist - Iso-Seq Template Preparation for Sequel Systems
|
|
|
- Jessica Bryan
- 5 years ago
- Views:
Transcription
1 Procedure & Checklist - Iso-Seq Template Preparation for Sequel Systems Before You Begin The Sequel System generates long reads that are well-suited for characterizing full-length transcripts produced from high-quality RNA samples. This document describes two methods to construct Iso-Seq SMRTbell libraries with and without size selection allowing detection of full-length transcripts Section 1 of this document describes the construction of a non-size selected Iso-Seq library. This procedure allows detection of full-length transcripts up to 4 kb (without doing size-selection). Using the Clontech SMARTer PCR cdna Synthesis Kit, RNA is synthesized to cdna and subsequently amplified to generate double-stranded cdna. Without size selection, the cdna is constructed to a SMRTbell library for sequencing. Section 2 describes a combined procedure to construct and pool a non-size selected Iso-Seq library with a size selected Iso-Seq library for sequencing on the Sequel System. By incorporating a size selection step using BluePippin or SageELF, this procedure enables one to increase the sequencing yield of >4 kb transcripts. Using the Clontech SMARTer PCR cdna Synthesis Kit, RNA is synthesized to cdna and subsequently amplified to generate double-stranded cdna. One portion of the amplified cdna product is used directly to construct a non-size selected SMRTbell library. In parallel, a second portion of the amplified cdna product is first size selected using either BluePippin or SageELF, and then the sizefractionated cdna is used to construct a size-selected SMRTbell library. Finally, the non-size selected SMRTbell library and the size-selected SMRTbell library are separately annealed and bound, and then pooled together for sequencing on the Sequel System. Users should proceed with following either Section 1 or Section 2 of this document to construct the desired type of Iso-Seq library that is most appropriate for their experimental design. PacBio recommends performing a no-size selection experiment first to better understand diversity of transcripts in your samples. Sections Procedure Target Size 1 Size Selection No Size-Selection Iso-Seq Template Preparation for Sequel Systems < 4kb No 2 Iso-Seq Template Preparation for Sequel Systems with Size selection For enriching >4kb transcripts Yes Page 1
2 Materials and Kits Needed Item Vendor SMARTer PCR cdna Synthesis Kit Clontech ( or ) PrimeSTAR GXL DNA Polymerase Clontech (R050A or R050B) Additional 5 PCR Primer IIA Any Oligo Synthesis Vendor 5 AAG CAG TGG TAT CAA CGC AGA GTA C 3 1.2% FlashGel system or 0.80% Agarose Gels Lonza (57023) or (57029) or Any MLS FlashGel DNA Marker (100bp 3 kb or 100 bp - 4 kb) Lonza Qubit dsdna BR Assay Kit or HS Assay Kit DNA Kit or HS DNA Kit SMRTbell Template Prep Kit Sequel Binding Kit Sequel Sequencing Kit AMPure PB Beads Optional for Size Selection: Blue Pippin System and Consumables: o BluePippin system with Software v5.90 or later o PacBio SMRTbell cassette definition set 0.75% DF 2 6kb Marker S1 o 0.75% Dye-Free Agarose Gel Cassettes Loading Solution o S1 Marker o Electrophoresis Buffer SageELF System and Consumables: o SageELF system with Software v0.57 or later o PacBio SMRTbell cassette definition set 0.75% Agarose, 1 kb-18 kb o 0.75% Dye-Free Agarose Gel Cassettes o Loading Solution o DNA Marker Invitrogen Agilent Pacific Biosciences Sage Science Page 2
3 Section 1: No Size-Selection Iso-Seq Template Preparation for Sequel Systems Page 3
4 Preparing cdna from RNA Samples RNA Input Requirements First strand cdna synthesis employs the Clontech SMARTer PCR cdna Synthesis Kit. The CDS Primer IIA is first annealed to the polya+ tail of transcripts, followed by first-strand synthesis with SMARTScribe Reverse Transcriptase. The first-strand product is diluted with Elution Buffer (EB) to an appropriate volume and subsequently used for large-scale PCR. If starting with Total RNA, prepare two or more first-strand cdna synthesis reactions. Consult your local Field Applications Scientist (FAS) for recommendations if starting with poly A+ RNA. First-Strand Synthesis This procedure has been optimized using 800 to1000 ng of total RNA as input for each cdna synthesis reaction. Note that as low as 2 ng of total RNA may be used in each cdna synthesis reaction however, the resulting amount of cdna product will not be sufficient to proceed with this procedure. In this case, PacBio recommends constructing an Iso-Seq library using a 1X AMPure PB bead purification method. Please contact your local Field Applications Scientist for recommendations on how to proceed with low input total RNA samples. For each sample, combine the reagents below. When using >1 µg of Total RNA as input, do not scale up the reaction volumes. Instead, perform multiple reactions separately (with each reaction in a 4.5 µl Total Volume). Reagent Volume Notes Total RNA ( ng) μl 3 SMART CDS Primer IIA (12 μm) 1 μl Nuclease-Free Water X Total Volume 4.5 μl 1. Mix contents and spin the tubes briefly in a microcentrifuge. 2. Incubate the tubes at 72 C in a hot -lid thermal cycler for 3 min; slowly ramp to 42 C at 0.1 C/sec, then let sit for 2 minutes. During this step, prepare a Master Mix for all reaction tubes, at room temperature, by combining the following reagents in the order shown. Important: go immediately into step 4 after step 3. However, add the reverse transcriptase to the master mix just prior to use. Mix well by pipetting and spin the tube briefly in a microcentrifuge. Reagent Volume Notes 5X First-Strand Buffer DTT (100 mm) dntp (10 mm) SMARTer IIA Oligonucleotide (12 μm) RNase Inhibitor SMARTScribe Reverse Transcriptase (100 U) - add immediately before use Total Volume added per reaction 2 μl 0.25 μl 1 μl 1 μl 0.25 μl 1 μl 5.5 μl Page 4
5 3. Place the master mix at 42 C for 1 min to bring it up to temperature and proceed immedi ately to step Aliquot 5.5 μl of the Master Mix into each reaction tube. Mix the contents of the tubes by gently pipetting, and spin the tubes briefly to collect the contents at the bottom. 5. Incubate the tubes at 42 C for 90 minutes. 6. Terminate the reaction by heating the tubes at 70 C for 10 min. 7. Dilute the first-strand reaction product by adding 90 μl of PacBio Elution Buffer (EB): Input Sample Total RNA (2 ng - 1 μg) Volume of EB Added 90 μl 8. Pool the diluted first-strand reactions for large-scale amplification. 9. Proceed to PCR cycle optimization PCR Cycle Optimization RNA samples from different sources behave differently during amplification. PacBio highly recommends performing cycle number optimizations to minimize PCR bias (prevent under or over-amplification). For 800 ng 1000 ng input total RNA into 1 st strand synthesis, the optimal number of cycles is typically If input total RNA is less than 800 ng, it is highly recommended to perform cycle optimizations to determine the best condition for large scale PCR. 1. Add the following reagents to an appropriately sized PCR tube: 5X PrimeSTAR GXL buffer Reagent Volume Notes Diluted first-strand cdna from step 7 above dntp Mix (2.5 mm each) 10 μl 10 μl 4 μl 5 PCR Primer IIA (12 μm) 1 μl Nuclease-free water PrimeSTAR GXL DNA Polymerase (1.25 U/μL) Total Volume 24 μl 1 μl 50 μl 2. Cycle the reaction with the following conditions (using the default heated lid setting): Initial denaturation: 98 C for 30s 10 cycles at the following temperatures and times: 98 C for 10 seconds 65 C for 15 seconds 68 C for 10 minutes Final extension: 68 C for 5 minutes 3. After the initial 10 cycles, remove 5 μl of the reaction and transfer it to a tube labeled Return the remaining 45 μl PCR reaction to the thermocycler and run two cycles of the above amplification conditions. 2 cycles at the following temperatures and times: 98 C for 10 seconds 65 C for 15 seconds 68 C for 10 minutes Final extension: 68 C for 5 minutes Page 5
6 5. Remove 5 μl again and transfer to a tube labeled Repeat steps 4-5 for 14 cycles. Note that the number of cycles is dependent on the sample input. Typically, ng input of total RNA requires 10 to 12 cycles of PCR amplification. Input total RNA of <800 ng may require more cycles. 7. Load the 3 aliquots on a 1% Agarose gel or Lonza flash Gel. Figure 1. This sample is Human Brain Total RNA from Biochain. One (1) µg Total RNA was used for 1 st strand synthesis. With 1 ug input, the optimal number of PCR cycles is cycles. To demonstrate cdna distribution, PCR aliquots (8, 10, 12, 14, 16, 18) collected during PCR optimization were run on agarose electrophoresis (1.2% Lonza FlashGel, Lonza FlashGel Ladder (100 bp 3 kb). The numbers above lanes indicate cycle number. In this example, 10 cycles were determined to be optimal for large-scale amplification. Smear distribution from 8 and 10 cycles look similar, however 10 cycles shows more products than 8 cycles. Cycles above 12 show signs of over-amplification which will result in biased sequencing representation. Page 6
7 Large-Scale PCR Use the cycle number (as determined in the PCR Cycle Optimization step) to generate a sufficient amount of double-stranded cdna product for SMRTbell library construction. 1. Set up 16 X 50 μl PCR reactions using the diluted first-strand cdna as input. 2. Make a master mix by adding the following reagents: Reagent Volume (1 rxn) Volume (16 rxns) Notes 5X PrimeSTAR GXL Buffer 10 μl 160 μl Diluted first-strand cdna 10 μl 160 μl dntp Mix (2.5 mm each) 4 μl 64 μl 5 PCR Primer IIA (12 μm) 1 μl 16 μl Nuclease-free water 24 μl 384 μl PrimeSTAR GXL DNA Polymerase (1.25 U/μL) 1 μl 16 μl Total Volume 50 μl 800 μl 3. Transfer 50 μl aliquots into 16 PCR tubes and perform PCR using the optimal cycle number determined from the optimization step. Cycle the reaction with the following conditions (using a heated lid): Initial denaturation: 98 C for 30 seconds N cycles (optimal cycle determined in the optimization step) at the following temperatures and times: 98 C for 1 0 seconds 65 C for 15 seconds 68 C for 10 minutes Final extension: 68 C for 5 minutes Page 7
8 AMPure PB Bead Purification of Large-Scale PCR Products This procedure requires the use of AMPure purification methods, 1X and 0.40X. Fraction 1 (pool of 6 tubes x 50 µl) is purified twice using 1X AMPure PB beads. Fraction 2 (pool of 10 tubes x 50 µl) is purified once using 0.40X AMPure PB beads. STEP 1X and 0.40X AMPure PB Bead Purification Notes 1 Pool 6 x 50 µl PCR reactions and add 1X volume of AMPure PB magnetic beads. This is Fraction 1. 2 Pool 10 x 50 µl PCR reactions and add 0.40X volume of AMPure PB magnetic beads in a separate 1.5mL LoBind tube. This is Fraction 2. 3 Process both tubes in parallel by mixing the bead/dna solution thoroughly. 4 Quickly spin down the tubes (for 1 second) to collect the beads. 5 Allow the DNA to bind to beads by shaking in a VWR vortex mixer at 2000 rpm for 10 minutes at room temperature. 6 Spin down both tubes (for 1 second) to collect beads. 7 Place the tubes in a magnetic bead rack until the beads collect to the side of the tubes and the solution appears clear. The actual time required to collect the beads to the side depends on the volume of beads added. 8 With the tubes still on the magnetic bead rack, slowly pipette off cleared supernatant and save in other tubes. Avoid disturbing the bead pellet. If the DNA is not recovered at the end of this procedure, you can add equal volumes of AMPure PB beads to the saved supernatant and repeat the AMPure PB bead purification steps to recover the DNA. 9 Wash beads with freshly prepared 70% ethanol. Note that 70% ethanol is hygroscopic and should be prepared FRESH to achieve optimal results. Also, 70% ethanol should be stored in a tightly capped polypropylene tube for no more than 3 days. 10 Repeat step 9. Do not remove the tubes from the magnetic rack. Use a sufficient volume of 70% ethanol to fill the tubes (1.5 ml for 1.5 ml tubes or 2 ml for 2 ml tubes). Slowly dispense the 70% ethanol against the side of the tubes opposite the beads. Do not disturb the bead pellet. After 30 seconds, pipette and discard the 70% ethanol. 11 Remove residual 70% ethanol. Remove tubes from magnetic bead rack and spin to pellet beads. Both the beads and any residual 70% ethanol will be at the bottom of the tubes. Place the tubes back on magnetic bead rack. Pipette off any remaining 70% ethanol. 12 Check for any remaining droplets in the tubes. If droplets are present, repeat step Remove the tubes from the magnetic bead rack and allow beads to air-dry (with the tube caps open) for seconds. Page 8
9 14 Add the Elution Buffer volume (see table below) to your beads. Tap the tubes with finger to mix until beads are uniformly re-suspended. Do not pipet to mix. Elute the DNA by letting the mix stand at room temperature for 2 minutes Spin the tube down to pellet beads, then place the tube back on the magnetic bead rack. Let beads separate fully. Then without disturbing the bead pellet, transfer supernatant to a new 1.5 ml Lo-Bind tube. Discard the beads. Fractions Elution Buffer Volume Fraction 1 (1X AMPure) 100 µl Fraction 2 (0.40X AMPure) 22 µl 15 Fraction 1 requires a second round of AMPure PB bead purification. Proceed directly to the next section ( Second Purification ). Fraction 2 does not require a second AMPure PB bead purification. Set this tube aside on ice and measure the DNA concentration along with Fraction 1 after the second 1X AMPure PB bead purification for Fraction 1 is completed. Page 9
10 STEP Second Purification Notes 1 Perform a second round of AMPure PB bead purification for Fraction 1 (now in 100 µl of EB) using 1X volume of AMPure PB magnetic beads 2 Quickly spin down the tube (for 1 second) to collect the beads. 3 Allow the DNA to bind to beads by shaking in a VWR vortex mixer at 2000 rpm for 10 minutes at room temperature. 4 Spin down both tube (for 1 second) to collect beads. 5 Place the tube in a magnetic bead rack until the beads collect to the side of the tube and the solution appears clear. The actual time required to collect the beads to the side depends on the volume of beads added. 6 With the tube still on the magnetic bead rack, slowly pipette off cleared supernatant and save in another tube. Avoid disturbing the bead pellet. If the DNA is not recovered at the end of this Procedure, you can add equal volumes of AMPure PB beads to the saved supernatant and repeat the AMPure PB bead purification steps to recover the DNA. 7 Wash beads with freshly prepared 70% ethanol. Note that 70% ethanol is hygroscopic and should be prepared FRESH to achieve optimal results. Also, 70% ethanol should be stored in a tightly capped polypropylene tube for no more than 3 days. 8 Repeat step 7. Do not remove the tube from the magnetic rack. Use a sufficient volume of 70% ethanol to fill the tube (1.5 ml for 1.5 ml tube or 2 ml for 2 ml tube). Slowly dispense the 70% ethanol against the side of the tube opposite the beads. Do not disturb the bead pellet. After 30 seconds, pipette and discard the 70% ethanol. 9 Remove residual 70% ethanol. Remove tube from magnetic bead rack and spin to pellet beads. Both the beads and any residual 70% ethanol will be at the bottom of the tube. Place the tube back on magnetic bead rack. Pipette off any remaining 70% ethanol. 10 Check for any remaining droplets in the tube. If droplets are present, repeat step Remove the tube from the magnetic bead rack and allow beads to air-dry (with the tube caps open) for seconds. 12 Add 22 μl of Elution Buffer volume to your beads. Tap the tube with finger to mix until beads are uniformly re-suspended. Do not pipet to mix. Elute the DNA by letting the mix stand at room temperature for 2 minutes Spin the tube down to pellet beads, then place the tube back on the magnetic bead rack. Let beads separate fully. Then without disturbing the bead pellet, transfer supernatant to a new 1.5 ml Lo-Bind tube. Discard the beads. Page 10
11 13 Verify the DNA amount and concentration of Fraction 1 and Fraction 2 using a Qubit quantitation platform. Measure the DNA concentration using a Qubit fluorometer. Using 1 μl of the eluted sample, make a 1:10 dilution in EB. Use 1 µl of this 1:10 dilution to measure the DNA concentration using a Qubit dsdna BR Assay kit and the dsdna HS Assay kit according to the manufacturer s recommendations. 14 Perform qualitative and quantitative analysis using a Bioanalyzer instrument with the DNA Kit. To determine the average library size, select the region of interest by defining the start and end points of the smear. Pooling Fraction 1 (1X) and Fraction 2 (0.40X) for SMRTbell Template Preparation The two purified fractions are pooled for library construction. STEP Pooling Notes 1 Based on sample information from the Qubit and BioAnalyzer, determine the molarity of the two fractions, which can be calculated by the following equation: concentration in ng/ul X 10 6 = concentration in nm (660 g/mol x average library size in bp*) *To determine the average library size, select the region of interest by defining the start and end points of the smear. 2 Pool equal molar quantities of the two fractions in a clean LoBind microcentrifuge tube. The total combined mass must be at least 1 µg. 3 Proceed to SMRTbell Template Preparation below. You need at least 1 µg of pooled cdna for library construction. Page 11
12 SMRTbell Template Preparation Repair DNA Damage 1. In a LoBind microcentrifuge tube, add the following reagents: Reagent Tube Cap Color Stock Conc. Volume Final Conc. Pooled cdna - μl for 1 to 5 μg - DNA Damage Repair Buffer 10 X 5.0 μl 1 X NAD+ 100 X 0.5 μl 1 X Notes ATP high 10 mm 5.0 μl 1 mm dntp 10 mm 0.5 μl 0.1 mm DNA Damage Repair Mix 2.0 μl H 2 O - μl to adjust to 50.0 μl - Total Volume 50.0 μl - 2. Mix the reaction well by pipetting or flicking the tube. 3. Spin down contents of tube with a quick spin in a microfuge. 4. Incubate at 37ºC for 20 minutes, then return the reaction to 4ºC for 1 minute. Repair Ends Use the following table to prepare your reaction then purify the DNA. Reagent Pooled cdna (Damage Repaired) Tube Cap Color Stock Conc. Volume Final Conc μl - Notes End Repair Mix 20 X 2.5 μl 1X Total Volume 52.5 μl - 1. Mix the reaction well by pipetting or flicking the tube. 2. Spin down contents of tube with a quick spin in a microfuge. 3. Incubate at 25ºC for 5 minutes, return the reaction to 4ºC. Page 12
13 STEP Purify DNA Notes 1 Add 1X volume of AMPure PB beads. 2 Mix the bead/dna solution thoroughly. 3 Quickly spin down the tube (for 1 second) to collect the beads. Do not pellet beads. 4 Allow the DNA to bind to beads by shaking in a VWR vortex mixer at 2000 rpm for 10 minutes at room temperature. 5 Spin down the tube (for 1 second) to collect beads. 6 Place the tube in a magnetic bead rack until the beads collect to the side of the tubes and the solution appears clear. The actual time required to collect the beads to the side depends on the volume of beads added. 7 With the tube still on the magnetic bead rack, slowly pipette off cleared supernatant and save in other tubes. Avoid disturbing the bead pellet. If the DNA is not recovered at the end of this procedure, you can add equal volumes of AMPure PB beads to the saved supernatant and repeat the AMPure PB bead purification steps to recover the DNA. 8 Wash beads with freshly prepared 70% ethanol. Note that 70% ethanol is hygroscopic and should be prepared FRESH to achieve optimal results. Also, 70% ethanol should be stored in a tightly capped polypropylene tube for no more than 3 days. Do not remove the tube from the magnetic rack. Use a sufficient volume of 70% ethanol to fill the tubes (1.5 ml for 1.5 ml tubes or 2 ml for 2 ml tubes). Slowly dispense the 70% ethanol against the side of the tubes opposite the beads. Do not disturb the bead pellet. After 30 seconds, pipette and discard the 70% ethanol. 9 Repeat step 8 above. 10 Remove residual 70% ethanol. Remove tube from magnetic bead rack and spin to pellet beads. Both the beads and any residual 70% ethanol will be at the bottom of the tubes. Place the tubes back on magnetic bead rack. Pipette off any remaining 70% ethanol. 11 Check for any remaining droplets in the tube. If droplets are present, repeat step Remove the tube from the magnetic bead rack and allow beads to air-dry (with tube caps open) for 30 to 60 seconds. 13 Add 32 μl of Elution Buffer volume to your beads. Tap the tube with finger to mix until beads are uniformly re-suspended. Do not pipet to mix. Elute the DNA by letting the mix stand at room temperature for 2 minutes Spin the tube down to pellet beads, then place the tube back on the magnetic bead rack. Let beads separate fully. Then without disturbing the bead pellet, transfer supernatant to a new 1.5 ml Lo-Bind tube. Discard the beads. 14 Optional: Verify your DNA amount and concentration using a Nanodrop or Qubit quantitation platform, as appropriate. 15 Optional: Perform qualitative and quantitative analysis using a Bioanalyzer instrument with the DNA Kit. 16 The End-Repaired DNA can be stored overnight at 4ºC or (or -20ºC for longer). 17 Enter your actual recovery per μl and total available sample material: Page 13
14 Prepare Blunt Ligation Reaction Use the following table to prepare your blunt ligation reaction: 1. In a LoBind microcentrifuge tube (on ice), add the following reagents in the order shown. If preparing a Master Mix, ensure that the adapter is NOT mixed with the ligase prior to introduction of the inserts. Add the adapter to the well with the DNA. All other components, including the ligase, should be added to the Master Mix. Reagent Pooled cdna (End Repaired) Tube Cap Color Stock Conc μl Volume Final Conc. Notes Blunt Adapter (20 μm) 20 μm 2.0 μl 1 μm Mix before proceeding Template Prep Buffer 10 X 4.0 μl 1X ATP low 1 mm 2.0 μl 0.05 mm Mix before proceeding Ligase 30 U/μL 1.0 μl 0.75 U/μL H 2 O - - μl to adjust to 40.0 μl - Total Volume μl - 2. Mix the reaction well by pipetting or flicking the tube. 3. Spin down contents of tube with a quick spin in a microfuge. 4. Incubate at 25ºC for 15 minutes. At this point, the ligation can be extended up to 24 hours or cooled to 4ºC (for storage up to 24 hours). 5. Incubate at 65ºC for 10 minutes to inactivate the ligase, then return the reaction to 4ºC. You must proceed with adding exonuclease after this step. Add Exonuclease to Remove Failed Ligation Products Reagent Tube Cap Color Stock Conc. Volume Ligated cdna Mix reaction well by pipetting 40 μl Exo III U/μL 1.0 μl Exo VII 10.0 U/μL 1.0 μl Total Volume 1. Mix the reaction well by pipetting or flicking the tube. 2. Spin down contents of tube with a quick spin in a microfuge. 3. Incubate at 37ºC for 1 hour, then return the reaction to 4ºC. You must proceed with purification after this step. 42 μl Page 14
15 Purify SMRTbell Templates STEP Purify SMRTbell Templates - First Purification Notes 1 Add 1X volume of AMPure PB beads. 2 Mix the bead/dna solution thoroughly. 3 Quickly spin down the tube (for 1 second) to collect the beads. Do not pellet beads. 4 Allow the DNA to bind to beads by shaking in a VWR vortex mixer at 2000 rpm for 10 minutes at room temperature. 5 Spin down the tube (for 1 second) to collect beads. 6 Place the tube in a magnetic bead rack until the beads collect to the side of the tubes and the solution appears clear. The actual time required to collect the beads to the side depends on the volume of beads added. 7 With the tube still on the magnetic bead rack, slowly pipette off cleared supernatant and save in other tubes. Avoid disturbing the bead pellet. If the DNA is not recovered at the end of this procedure, you can add equal volumes of AMPure PB beads to the saved supernatant and repeat the AMPure PB bead purification steps to recover the DNA. 8 Wash beads with freshly prepared 70% ethanol. Note that 70% ethanol is hygroscopic and should be prepared FRESH to achieve optimal results. Also, 70% ethanol should be stored in a tightly capped polypropylene tube for no more than 3 days. Do not remove the tube from the magnetic rack. Use a sufficient volume of 70% ethanol to fill the tubes (1.5 ml for 1.5 ml tubes or 2 ml for 2 ml tubes). Slowly dispense the 70% ethanol against the side of the tubes opposite the beads. Do not disturb the bead pellet. After 30 seconds, pipette and discard the 70% ethanol. 9 Repeat step 8 above. 10 Remove residual 70% ethanol. Remove tube from magnetic bead rack and spin to pellet beads. Both the beads and any residual 70% ethanol will be at the bottom of the tubes. Place the tubes back on magnetic bead rack. Pipette off any remaining 70% ethanol. 11 Check for any remaining droplets in the tube. If droplets are present, repeat step Remove the tube from the magnetic bead rack and allow beads to air-dry (with tube caps open) for 30 to 60 seconds. 13 Add 50 μl of Elution Buffer volume to your beads. Tap the tube with finger to mix until beads are uniformly re-suspended. Do not pipet to mix. Elute the DNA by letting the mix stand at room temperature for 2 minutes Spin the tube down to pellet beads, then place the tube back on the magnetic bead rack. Let beads separate fully. Then without disturbing the bead pellet, transfer supernatant to a new 1.5 ml Lo-Bind tube. Discard the beads. Page 15
16 STEP Purify SMRTbell Templates - Second Purification Notes 1 Add 1X volume of AMPure PB beads. 2 Mix the bead/dna solution thoroughly. 3 Quickly spin down the tube (for 1 second) to collect the beads. Do not pellet beads. 4 Allow the DNA to bind to beads by shaking in a VWR vortex mixer at 2000 rpm for 10 minutes at room temperature. 5 Spin down the tube (for 1 second) to collect beads. 6 Place the tube in a magnetic bead rack until the beads collect to the side of the tubes and the solution appears clear. The actual time required to collect the beads to the side depends on the volume of beads added. 7 With the tube still on the magnetic bead rack, slowly pipette off cleared supernatant and save in other tubes. Avoid disturbing the bead pellet. If the DNA is not recovered at the end of this procedure, you can add equal volumes of AMPure PB beads to the saved supernatant and repeat the AMPure PB bead purification steps to recover the DNA. 8 Wash beads with freshly prepared 70% ethanol. Note that 70% ethanol is hygroscopic and should be prepared FRESH to achieve optimal results. Also, 70% ethanol should be stored in a tightly capped polypropylene tube for no more than 3 days. Do not remove the tube from the magnetic rack. Use a sufficient volume of 70% ethanol to fill the tubes (1.5 ml for 1.5 ml tubes or 2 ml for 2 ml tubes). Slowly dispense the 70% ethanol against the side of the tubes opposite the beads. Do not disturb the bead pellet. After 30 seconds, pipette and discard the 70% ethanol. 9 Repeat step 8 above. 10 Remove residual 70% ethanol. Remove tube from magnetic bead rack and spin to pellet beads. Both the beads and any residual 70% ethanol will be at the bottom of the tubes. Place the tubes back on magnetic bead rack. Pipette off any remaining 70% ethanol. 11 Check for any remaining droplets in the tube. If droplets are present, repeat step Remove the tube from the magnetic bead rack and allow beads to air-dry (with tube caps open) for 30 to 60 seconds. 13 Add 10 μl of Elution Buffer volume to your beads. Tap the tube with finger to mix until beads are uniformly re-suspended. Do not pipet to mix. Elute the DNA by letting the mix stand at room temperature for 2 minutes Spin the tube down to pellet beads, then place the tube back on the magnetic bead rack. Let beads separate fully. Then without disturbing the bead pellet, transfer supernatant to a new 1.5 ml Lo-Bind tube. Discard the beads. Page 16
17 14 Verify the DNA amount and concentration using a Qubit quantitation platform. Measure the DNA concentration using a Qubit fluorometer. Using 1 μl of the eluted sample, make a 1:10 dilution in EB. Use 1 µl of this 1:10 dilution to measure the DNA concentration using a Qubit dsdna BR Assay kit and the dsdna HS Assay kit according to the manufacturer s recommendations. 15 Perform qualitative and quantitative analysis using a Bioanalyzer instrument with the DNA Kit. To determine the average library size, select the region of interest by defining the start and end points of the smear. Anneal and Bind Non-Size Selected SMRTbell Templates Follow the SMRT Link Sample Setup instructions for primer annealing and polymerase binding conditions to prepare your non-size selected Iso-Seq library for sequencing. Sequence MagBead loading is recommended for Iso-Seq libraries prepared using this procedure. PacBio recommends performing loading titrations to determine an appropriate loading concentration. Sequencing recommendations: Loading: >40 pm on-plate concentration, Target P1 >50% (Performing loading titrations to determine the appropriate loading concentration is recommended.) Movie Collection time: minutes Pre-extension time: 120 minutes Sample Cleanup: Not required Page 17
18 Section 2: Iso-Seq Template Preparation for Sequel Systems with Size Selection Sample Considerations for Performing Size Selection To sequence a broad range of transcripts (5 kb -10 kb) using the Sequel System, consider including a size selection step as part of the Iso-Seq library preparation workflow. Amplification and sequencing of full-length transcripts > 5 kb can be achieved depending on the sample type and quality. Sample 1 below is an example of a sample that would benefit from size-selection since the Bioanalyzer sizing QC plot indicates the presence of transcripts between 5 kb to 8 kb. However, Sample 2 will likely not benefit from size selection because there are no transcripts >5 kb visible in the QC plot. Sample 1 Sample 2 This section of the procedure describes how to construct Iso-Seq libraries using a parallel workflow that incorporates both no-size selection steps and size selection steps (see flow chart below). Page 18
19 Page 19
20 First-Strand Synthesis Requirements The table below summarizes the total number of first-strand cdna synthesis reactions required for carrying out parallel non-size selection and size-selection Iso-Seq library construction. RNA Input Type Total RNA Recommended Number of First-Strand Synthesis Reactions 3 or more Refer to Preparing cdna from RNA Samples in Section 1 for detailed instructions on how to prepare Firststrand cdna synthesis reactions. For each cdna synthesis reaction, dilute the first-strand reaction product by adding the appropriate volume of PacBio Elution Buffer (EB) as follows: Input Sample Total RNA (2 ng - 1 μg) Volume of EB Added 90 μl Large-Scale PCR 1. Set up 24 x 50 μl PCR reactions if 3 first-strand cdna synthesis reaction with Total RNA were prepared in the previous step above. Scale up the large-scale PCR reaction volumes if additional firststrand cdna synthesis reactions are needed for your experiment. 2. Make a master mix by adding the following reagents: Reagent Volume (1 rxn) Volume (24 rxns) Notes 5X PrimeSTAR GXL Buffer 10 μl 240 μl Diluted first-strand cdna 10 μl 240 μl dntp Mix (2.5mM each) 4 μl 96 μl 5 PCR Primer IIA (12 μm) 1 μl 24 μl Nuclease-free water 24 μl 576 μl PrimeSTAR GXL DNA Polymerase (1.25 U/μL) 1 μl 24 μl Total Volume 50 μl 1200 μl 3. Transfer 50 μl aliquots into 24 PCR tubes and perform PCR using the optimal cycle number determined from the optimization step. Cycle the reaction with the following conditions (using a heated lid): Initial denaturation: 98 C for 30 seconds N cycles (optimal cycle determined in the optimization step) at the following temperatures and times: 98 C for 1 0 seconds 65 C for 15 seconds 68 C for 10 minutes Final extension: 68 C for 5 minutes Page 20
21 AMPure PB Bead Purification of Large-Scale PCR Products Fraction 1 (pool of 6 tubes x 50 µl) is purified using 1X AMPure PB beads. Fraction 2 (pool of 10 tubes x 50 µl) is purified using 0.40X AMPure PB beads. Fraction 3 (pool of 8 tubes x 50 µl) is purified using 1X AMPure PB beads and will be used for size selection. STEP 1X and 0.40X AMPure PB Bead Purification Notes 1 Pool 6 x 50 µl PCR reactions and add 1X volume of AMPure PB magnetic beads. This is Fraction 1. 2 Pool 10 x 50 µl PCR reactions and add 0.40X volume of AMPure PB magnetic beads in a separate 1.5mL LoBind tube. This is Fraction 2. 3 Pool 8 x 50 µl PCR reactions and add 1X volume of AMPure PB magnetic beads in a separate 1.5mL LoBind tube. This is Fraction 3. 4 Process all tubes in parallel by mixing the bead/dna solution thoroughly. 5 Quickly spin down the tubes (for 1 second) to collect the beads. 6 Allow the DNA to bind to beads by shaking in a VWR vortex mixer at 2000 rpm for 10 minutes at room temperature. 7 Spin down tubes (for 1 second) to collect beads. 8 Place the tubes in a magnetic bead rack until the beads collect to the side of the tubes and the solution appears clear. The actual time required to collect the beads to the side depends on the volume of beads added. 9 With the tubes still on the magnetic bead rack, slowly pipette off cleared supernatant and save in other tubes. Avoid disturbing the bead pellet. If the DNA is not recovered at the end of this procedure, you can add equal volumes of AMPure PB beads to the saved supernatant and repeat the AMPure PB bead purification steps to recover the DNA. 10 Wash beads with freshly prepared 70% ethanol. Note that 70% ethanol is hygroscopic and should be prepared FRESH to achieve optimal results. Also, 70% ethanol should be stored in a tightly capped polypropylene tube for no more than 3 days. 11 Repeat step 10. Do not remove the tubes from the magnetic rack. Use a sufficient volume of 70% ethanol to fill the tubes (1.5 ml for 1.5 ml tubes or 2 ml for 2 ml tubes). Slowly dispense the 70% ethanol against the side of the tubes opposite the beads. Do not disturb the bead pellet. After 30 seconds, pipette and discard the 70% ethanol. 12 Remove residual 70% ethanol. Remove tubes from magnetic bead rack and spin to pellet beads. Both the beads and any residual 70% ethanol will be at the bottom of the tubes. Place the tubes back on magnetic bead rack. Pipette off any remaining 70% ethanol. 13 Check for any remaining droplets in the tubes. If droplets are present, repeat step 12. Page 21
22 14 Remove the tubes from the magnetic bead rack and allow beads to air-dry (with the tube caps open) for seconds. 15 Add the Elution Buffer volume (see table below) to your beads. Tap the tubes with finger to mix until beads are uniformly re-suspended. Do not pipet to mix. Elute the DNA by letting the mix stand at room temperature for 2 minutes Spin the tube down to pellet beads, then place the tube back on the magnetic bead rack. Let beads separate fully. Then without disturbing the bead pellet, transfer supernatant to a new 1.5 ml Lo-Bind tube. Discard the beads. Fractions Elution Buffer Volume Fraction 1 (1X AMPure) 100 µl Fraction 2 (0.40X AMPure) 22 µl Fraction 3 (1X AMPure) 100 ul 16 Fraction 1 and Fraction 3 require a second purification. Proceed directly to the next section ( Second Purification ). Fraction 2 does not require a second purification. Set aside in ice and measure concentration along with Fraction 1 and Fraction 3 after the second AMPure PB bead purification. Page 22
23 STEP Second Purification Notes 1 Perform a second 1X AMPure PB bead purification for Fraction 1 and Fraction 3. 2 Quickly spin down the tubes (for 1 second) to collect the beads. 3 Allow the DNA to bind to beads by shaking in a VWR vortex mixer at 2000 rpm for 10 minutes at room temperature. 4 Spin down both tubes (for 1 second) to collect beads. 5 Place the tubes in a magnetic bead rack until the beads collect to the side of the tube and the solution appears clear. The actual time required to collect the beads to the side depends on the volume of beads added. 6 With the tubes still on the magnetic bead rack, slowly pipette off cleared supernatant and save in another tube. Avoid disturbing the bead pellet. If the DNA is not recovered at the end of this Procedure, you can add equal volumes of AMPure PB beads to the saved supernatant and repeat the AMPure PB bead purification steps to recover the DNA. 7 Wash beads with freshly prepared 70% ethanol. Note that 70% ethanol is hygroscopic and should be prepared FRESH to achieve optimal results. Also, 70% ethanol should be stored in a tightly capped polypropylene tube for no more than 3 days. 8 Repeat step 7. Do not remove the tubes from the magnetic rack. Use a sufficient volume of 70% ethanol to fill the tube (1.5 ml for 1.5 ml tube or 2 ml for 2 ml tube). Slowly dispense the 70% ethanol against the side of the tube opposite the beads. Do not disturb the bead pellet. After 30 seconds, pipette and discard the 70% ethanol. 9 Remove residual 70% ethanol. Remove tubes from magnetic bead rack and spin to pellet beads. Both the beads and any residual 70% ethanol will be at the bottom of the tube. Place the tube back on magnetic bead rack. Pipette off any remaining 70% ethanol. 10 Check for any remaining droplets in the tube. If droplets are present, repeat step Remove the tubes from the magnetic bead rack and allow beads to air-dry (with the tube caps open) for seconds. 12 Add 22 μl of Elution Buffer volume to your beads. Tap the tube with finger to mix until beads are uniformly re-suspended. Do not pipet to mix. Elute the DNA by letting the mixes stand at room temperature for 2 minutes Spin the tubes down to pellet beads, then place the tube back on the magnetic bead rack. Let beads separate fully. Then without disturbing the bead pellet, transfer supernatant to a new 1.5 ml Lo-Bind tube. Discard the beads. Page 23
24 13 Verify the DNA amount and concentration of Fractions 1, 2 and 3 using a Qubit quantitation platform. Measure the DNA concentration using a Qubit fluorometer. Using 1 μl of the eluted sample, make a 1:10 dilution in EB. Use 1 µl of this 1:10 dilution to measure the DNA concentration using a Qubit dsdna BR Assay kit and the dsdna HS Assay kit according to the manufacturer s recommendations. 14 Perform qualitative and quantitative analysis using a Bioanalyzer instrument with the DNA Kit. To determine the average library size, select the region of interest by defining the start and end points of the smear. 15 Use Fraction 3 for size selection. Use Fraction 1 and Fraction 2 for Pooling in the next section. Pooling Fraction 1 (1X) and Fraction 2 (0.40X) for Non-Size Selected SMRTbell Template Preparation: STEP Pooling Notes 1 Based on sample information from the Qubit and BioAnalyzer, determine the molarity of the two fractions, which can be calculated by the following equation: concentration in ng/ul X 10 6 = concentration in nm (660 g/mol x average library size* in bp) *To determine the average library size, select the region of interest by defining the start and end points of the smear. 2 Pool equal molar of the two fractions. The total mass must be at least 1 µg. 3 The pooled fractions can now be taken directly into SMRTbell Library construction starting with Repair DNA Damage. You need at least 1 µg cdna for library construction. Page 24
25 Size Selection Procedure for Fraction 3 (1X AMPure Purified): The Fraction 3 (1X) AMPure PB bead-purified sample is size-selected using a BluePippin System or a SageELF System. Option 1 - Size Selection of Fraction 3 (1X) using the BluePippin System For BluePippin, 500 ng 5000 ng is required. STEP Running the BluePippin System Notes 1 Follow the BluePippin Manual and instructions to calibrate your instrument. A new calibration is recommended before each BluePippin run. 2 Inspect the gel cassette (using Sage Sciences BluePippin manual). Ensure that the buffer wells are full. Ensure that there is no separation of the gel from the cassette. 3 Prepare the gel cassette: Remove all bubbles from the elution buffer chamber by tilting the cassette and tapping it until all air bubbles move into the buffer chamber. Place the gel cassette in the BluePippin System and carefully remove the plastic seals on the cassette. Remove the buffer from the elution well and fill with 40 µl of fresh Electrophoresis Buffer. Keep the pipette down the center of the well and avoid creating a vacuum in the well. The bottom of the well is okay to touch. If the well bubbles over when adding the buffer to the well, remove buffer and try again. If the well continues to bubble over, then use this well for the S1 marker. The elution well may be damaged and should not be used for sample collection. Cover the elution wells with a clear adhesive tape Remove the buffer from the sample well and fill with 70 µl of fresh Electrophoresis Buffer Close the lid and perform a Continuity Test. 4 Prepare samples for loading >500 ng (up to 5 µg) is required. Use Elution Buffer to dilute sample to 30 µl. Add 10 µl of the Loading Solution and vortex to mix well. 5 Load samples: Remove 40 µl of buffer from each well. Load all 40 µl of the sample prepared in step 4 into each lane. Load 40 µl of S1 Marker in one of the lanes. Page 25
26 STEP Running the BluePippin System Notes 6 Set up the run Protocol: 7 Start the run. Click on the New button to create a new protocol. Select the 0.75% DF 2 6kb Marker S1 cassette definition file. Click on the box below End Run when Elution is Complete. Set the lane where the S1 Marker is loaded as the reference Ref lane. Click on the Range button and enter the following: Library Size BP Start BP End 5 kb - 10 kb After the run, collect approximately 40 µl the respective fractions from each lane. 9 Wash the elution well with 40uL of EB, pipet up and down 10 times, let the buffer sit for 10 minutes before removing from well. Combine the 40uL wash with the 40uL of eluate The samples can be stored in - 20 C or used directly in the next step ( Large-Scale PCR Post-Size Selection ). Note that DNA quantification using a Qubit system is not necessary at this point. Page 26
27 Option 2 Size Selection of Fraction 3 (1X) using the SageELF System For Sage ELF, 1000 ng ng is required. STEP Running the SageELF System Notes 1 Follow the SageELF Manual and instructions to calibrate your instrument. A new calibration is recommended before each run. 2 Inspect the gel cassette (using Sage Science s SageELF manual). Ensure that the buffer wells are full. Ensure that there is no separation of the gel from the cassette. 3 Prepare the gel cassette: While the cassette is sealed, remove all bubbles from the elution buffer chamber by tilting the cassette and tapping it until all air bubbles move into the buffer chamber. Hold the cassette firmly on the bench top and carefully remove the plastic seals on the cassette. Remove the buffer from the elution well and fill with 30 μl of fresh Electrophoresis Buffer. Keep the pipette down the center of the well and avoid creating a vacuum in the well. The bottom of the well is okay to touch. If the well bubbles over when adding the buffer to the well, remove buffer and try again. Cover the elution wells with a clear adhesive tape and verify that it is tightly sealed. Remove the buffer from the sample well and fill with 70 μl of fresh Electrophoresis Buffer. Do not touch the sides and bottom of the sample well. Carefully place the gel cassette in the SageELF System. Verify that the moat on both sides of the cassette, that connect the electrode reservoirs, are full. Add additional electrophoresis buffer to fill up the moat, if necessary. Close the lid and perform a Current Test. 4 Prepare samples for loading 5 Load samples: Prepare 30 μl tube with 1-5 μg of amplified cdna. It is highly recommended to start with >3 μg of cdna. Add 10 μl of Sage Science s Marker 75. Mix well and do a quick spin down. Remove 40 μl of buffer from the sample well. Load all 40 μl of the sample prepared in step 4 into the sample well. If necessary, top off well with additional Electrophoresis Buffer. Do not overflow the well. Page 27
28 6 Set up the run Protocol: 7 Start the run. In the Protocol Editor tab, click on the New Protocol button. Select the 0.75% Dye Free 1-18kb in the cassette definition menu. Select size-based for separation mode. Fill in the Target Value field and use the slider to select well #10. Enter 1500 bp in the target value. Save as new protocol. On the Main screen, clear previous run data, select cassette description, cassette definition and protocol, enter sample ID(s). Select in the Nest Selector the cartridge that will be run. 8 Once the run is complete, (approximately 3 hours), collect 30 μl of the respective fractions from the elution wells. Rinse each well by adding 30 μl of fresh Elution Buffer into the empty elution well. Rinse by pipetting up and down several times, and collect the rinse into the same tube. 9 Measure concentration by Qubit. 10 Check the sizes of all 12 fractions by loading on a Bioanalyzer system using a DNA kit. To determine the average library size, select the region of interest by defining the start and end points of the smears. 11 This is a safe stopping point. The 12 fractions can be stored at - 20 C for future use. 12 Pool fractions 1-4. This pool will contain >5 kb cdna. When pooling, mix fractions in equimolar quantities. (Note that this is the reason accurate quantitation using the Qubit system is essential). The samples can be stored in - 20 C or used directly in the next step ( Large-Scale PCR Post-Size Selection ). Note that DNA quantification using a Qubit system is not necessary at this point. Page 28
29 Large-Scale PCR Post-Size Selection The amount of recovered size-selected Fraction 3 (1X) sample is not sufficient for SMRTbell library construction. It is necessary to perform additional amplification to enrich for >5 kb cdna. 1. Set up 6 X 50 µl PCR reactions. 2. Add the following reagents to an appropriately sized PCR tube: Reagent Volume (1 rxn) Volume (6 rxns) Notes 5X PrimeSTAR GXL Buffer 10 µl 60 µl Eluted DNA from size selection 10 µl 60 µl dntp Mix (2.5 mm) 4 µl 24 µl 5 PCR Primer IIA (12 µm) 1 µl 6 µl Nuclease-free water 24 µl 144 µl PrimeSTAR GXL DNA Polymerase (1.25U/µL) 1 µl 6 µl Total Volume 50 µl 300 µl 3. Aliquot 50 µl into 6 PCR tubes and perform PCR using the cycle number and extension parameters below. Size Desired Extension Time Number of Cycles 5 kb - 10 kb 10 min 6-10 * *The number of cycles depends on your recovery after size selection. If concentration is > 4ng/µL, we recommend 6 cycles. If <4 ng/µl, use 10 cycles. 4. Cycle the reaction with the following conditions (using a heated lid): Initial denaturation: 98 C for 30s N (see above recommendation) cycles at the following temperatures and times: 98 C for 10 seconds 65 C for 15 seconds 68 C for 10 minutes (for this step, see extension times in table above) Final extension: 68 C for 5 minutes Page 29
30 Purifying the Large-Scale PCR Products After PCR, pool the PCR reactions into a 1.5 ml tube and purify with 0.5X AMPure PB beads. STEP AMPure PB Bead Purification Notes 1 Add 0.5X volume of AMPure PB magnetic beads to the amplified cdna sample. 2 Mix the bead/dna solution thoroughly. 3 Quickly spin down the tube (for 1 second) to collect the beads. 4 Allow the DNA to bind to beads by shaking in a VWR vortex mixer at 2000 rpm for 10 minutes at room temperature. 5 Spin down the tube (for 1 second) to collect beads. 6 Place the tube in a magnetic bead rack until the beads collect to the side of the tubes and the solution appears clear. The actual time required to collect the beads to the side depends on the volume of beads added. 7 With the tube still on the magnetic bead rack, slowly pipette off cleared supernatant and save in other tubes. Avoid disturbing the bead pellet. If the DNA is not recovered at the end of this procedure, you can add equal volumes of AMPure PB beads to the saved supernatant and repeat the AMPure PB bead purification steps to recover the DNA. 8 Wash beads with freshly prepared 70% ethanol. 9 Repeat step Remove residual 70% ethanol. Remove tube from magnetic bead rack and spin to pellet beads. Both the beads and any residual 70% ethanol will be at the bottom of the tubes. Place the tube back on magnetic bead rack. Pipette off any remaining 70% ethanol. 11 Check for any remaining droplets in the tube. If droplets are present, repeat step Remove the tube from the magnetic bead rack and allow beads to air-dry (with the tube caps open) for seconds. 13 Add the ul Elution Buffer volume to your beads. Tap the tubes with finger to mix until beads are uniformly re-suspended. Do not pipet to mix. Elute the DNA by letting the mix stand at room temperature for 2 minutes Spin the tube down to pellet beads, then place the tube back on the magnetic bead rack. Let beads separate fully. Then without disturbing the bead pellet, transfer supernatant to a new 1.5 ml Lo-Bind tube. Discard the beads. 14 Verify the DNA amount and concentration using a Qubit quantitation platform. Measure the DNA concentration using a Qubit fluorometer. 15 Perform qualitative and quantitative analysis using a Bioanalyzer instrument with the DNA Kit. To determine the average library size, select the region of interest by defining the start and end points of the smear. 16 The enriched size selected sample can now be taken into SMRTbell Library construction starting with Repair DNA Damage. You need at least 1 µg cdna for library construction. Page 30
31 SMRTbell Template Preparation Prepare a non-size selected SMRTbell library using the (non-size selected) equimolar-pooled Fraction 1 (1X) and Fraction 2 (0.40X) cdna samples. In parallel, prepare a second SMRTbell library using the size-selected Fraction 3 (1X) cdna sample. For detailed instructions on how to prepare SMRTbell libraries, see the SMRTbell Template Preparation procedure in Section 1. After completing the SMRTbell Library construction steps, proceed to Anneal and Bind Non-Size Selected and Size-Selected SMRTbell Templates below. Anneal and Bind Non-Size Selected and Size-Selected SMRTbell Templates Prepare your non-size selected Iso-Seq library and size-selected Iso-Seq library for sequencing by following the SMRT Link Sample Setup instructions for primer annealing and polymerase binding conditions: 1. Perform primer annealing and polymerase binding of the pooled non-size-selected Iso-Seq library. 2. In parallel, perform primer annealing and polymerase binding of the size-selected Iso-Seq library 3. The non-size-selected binding complex and size-selected binding complex can now be pooled together in a 5:1 molar ratio for sequencing on the same Sequel SMRT Cell. 4. [Optional: The non-size selected complex can also be sequenced on its own Sequel SMRT Cell (i.e., without pooling with the size-selected complex) if desired] Sequence MagBead loading is recommended for Iso-Seq libraries prepared using this procedure. PacBio recommends performing loading titrations to determine an appropriate loading concentration. Sequencing recommendations: Loading: >40 pm on-plate concentration, Target P1 50% (Performing loading titrations to determine the appropriate loading concentration is recommended.) Movie Collection time: minutes Pre-extension time: 120 minutes Sample Cleanup: Not required Revision History (Description) Version Date Incorporates (adds in) procedure on constructing and pooling a non-size selected Iso-Seq library with a size selected Iso-Seq library. On page 14 (Blunt Ligation Reaction table), sample volume (Pooled cdna) showed 32 µl. It has been corrected to 31 µl. On pages 17 and 31, on-plate concentration changed from pm to >40 pm. Error fixed in flowchart of page 19: AMPure volume after large-scale PCR (for Fraction 3) changed from 1X to 0.5X AMPure. 03 October October November 2017 For Research Use Only. Not for use in diagnostic procedures. Copyright 2017, Pacific Biosciences of California, Inc. All rights reserved. Information in this document is subject to change without notice. Pacific Biosciences assumes no responsibility for any errors or omissions in this document. Certain notices, terms, conditions and/o r use restrictions may pertain to your use of Pacific Biosciences products and/or third p arty products. Please refer to the applicable Pacific Biosciences Terms and Conditions of S ale and to the applicable license terms at nses.html. Pacific Biosciences, the Pacific Biosciences logo, PacBio, S MRT, SMRTbell, Iso-Seq and Sequel are trademarks of Pacific Biosciences. BluePippin and SageELF are trademarks of Sage Science, Inc. NGS-go and NGSengine are trademarks of GenDx. FEMTO Pulse and Fragment Analyzer are trademarks of Advanced Analytical Technologies. All other trademarks are the sole property of their respective owners. Page 31
Procedure & Checklist - Iso-Seq Template Preparation for Sequel Systems
 Procedure & Checklist - Iso-Seq Template Preparation for Sequel Systems Before You Begin The long read lengths of the PacBio System are well-suited for characterizing full-length transcripts produced from
Procedure & Checklist - Iso-Seq Template Preparation for Sequel Systems Before You Begin The long read lengths of the PacBio System are well-suited for characterizing full-length transcripts produced from
Procedure & Checklist - Greater Than 10 kb Template Preparation Using AMPure PB Beads
 Procedure & Checklist - Greater Than 10 kb Template Preparation Using AMPure PB Beads Before You Begin This procedure can be used to prepare greater than 10 kb libraries from 5 μg of sheared and concentrated
Procedure & Checklist - Greater Than 10 kb Template Preparation Using AMPure PB Beads Before You Begin This procedure can be used to prepare greater than 10 kb libraries from 5 μg of sheared and concentrated
Procedure & Checklist >20 kb Template Preparation Using BluePippin Size-Selection System (15-20 kb Cutoff) for Sequel Systems
 Procedure & Checklist >20 kb Template Preparation Using BluePippin Size-Selection System (15-20 kb Cutoff) for Sequel Systems Before You Begin To perform this procedure, you must have the PacBio Template
Procedure & Checklist >20 kb Template Preparation Using BluePippin Size-Selection System (15-20 kb Cutoff) for Sequel Systems Before You Begin To perform this procedure, you must have the PacBio Template
Procedure and Checklist - 20 kb Template Preparation Using BluePippin Size-Selection System
 Procedure and Checklist - 20 kb Template Preparation Using BluePippin Size-Selection System Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit and have reviewed the
Procedure and Checklist - 20 kb Template Preparation Using BluePippin Size-Selection System Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit and have reviewed the
Procedure & Checklist - Preparing SMRTbell Libraries using PacBio Barcoded Adapters for Multiplex SMRT Sequencing
 Procedure & Checklist - Preparing SMRTbell Libraries using PacBio Barcoded Adapters for Multiplex SMRT Sequencing Before You Begin This document describes methods for generating SMRTbell libraries using
Procedure & Checklist - Preparing SMRTbell Libraries using PacBio Barcoded Adapters for Multiplex SMRT Sequencing Before You Begin This document describes methods for generating SMRTbell libraries using
10 kb to 20 kb Template Preparation and Sequencing with Low-Input DNA
 Please note: the shared protocols described herein may not have been validated by Pacific Biosciences and are provided as-is and without any warranty. Use of these protocols is offered to those customers
Please note: the shared protocols described herein may not have been validated by Pacific Biosciences and are provided as-is and without any warranty. Use of these protocols is offered to those customers
Procedure & Checklist - 1 kb Template Preparation and Sequencing
 Procedure & Checklist - 1 kb Template Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit. Fragment and Concentrate DNA Important: The distribution
Procedure & Checklist - 1 kb Template Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit. Fragment and Concentrate DNA Important: The distribution
Procedure & Checklist - 20 kb Template Preparation Using BluePippin Size-Selection System
 Procedure & Checklist - 20 kb Template Preparation Using BluePippin Size-Selection System Before You Begin To perform this procedure, you must have the PacBio DNA Template Prep Kit 2.0 (3 kb to 10 kb)
Procedure & Checklist - 20 kb Template Preparation Using BluePippin Size-Selection System Before You Begin To perform this procedure, you must have the PacBio DNA Template Prep Kit 2.0 (3 kb to 10 kb)
Procedure & Checklist - cdna Capture Using SeqCap EZ Libraries
 Procedure & Checklist - cdna Capture Using SeqCap EZ Libraries Before You Begin This document describes the process for capturing cdna prepared with the SMARTer PCR cdna Synthesis Kit (Clontech) and pulled-down
Procedure & Checklist - cdna Capture Using SeqCap EZ Libraries Before You Begin This document describes the process for capturing cdna prepared with the SMARTer PCR cdna Synthesis Kit (Clontech) and pulled-down
Required Materials. Page 1 PN Version 06 (February 2018)
 Procedure & Checklist - Preparing >30 kb SMRTbell Libraries Using Megaruptor Shearing and BluePippin Size-Selection for PacBio RS II and Sequel Systems This document provides recommendations for preparing
Procedure & Checklist - Preparing >30 kb SMRTbell Libraries Using Megaruptor Shearing and BluePippin Size-Selection for PacBio RS II and Sequel Systems This document provides recommendations for preparing
Procedure & Checklist bp Template Preparation and Sequencing
 Procedure & Checklist - 500 bp Template Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit. This procedure is optimized for SMRTbell template
Procedure & Checklist - 500 bp Template Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit. This procedure is optimized for SMRTbell template
Procedure & Checklist - 10 kb Template Preparation and Sequencing (with Low-Input DNA)
 Procedure & Checklist - 10 kb Template Preparation and Sequencing (with Low-Input DNA) Before You Begin To perform this procedure, you must have the PacBio : Template Prep Kit DNA/Polymerase Binding Kit
Procedure & Checklist - 10 kb Template Preparation and Sequencing (with Low-Input DNA) Before You Begin To perform this procedure, you must have the PacBio : Template Prep Kit DNA/Polymerase Binding Kit
Procedure & Checklist - 2 kb Template Preparation and Sequencing
 Procedure & Checklist - 2 kb Template Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio DNA Template Prep Kit (verify you have the correct kit for your insert
Procedure & Checklist - 2 kb Template Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio DNA Template Prep Kit (verify you have the correct kit for your insert
Procedure & Checklist - Preparing >15 kb Libraries Using SMRTbell Express Template Preparation Kit
 Procedure & Checklist - Preparing >15 kb Libraries Using SMRTbell Express Template Preparation Kit This document provides recommendations for preparing >15 kb size-selected SMRTbell libraries from 3-5
Procedure & Checklist - Preparing >15 kb Libraries Using SMRTbell Express Template Preparation Kit This document provides recommendations for preparing >15 kb size-selected SMRTbell libraries from 3-5
Procedure & Checklist bp Amplicon Library Preparation and Sequencing
 Procedure & Checklist - 250 bp Amplicon Library Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit. This procedure is optimized for SMRTbell
Procedure & Checklist - 250 bp Amplicon Library Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit. This procedure is optimized for SMRTbell
Procedure & Checklist - 10 kb Template Preparation and Sequencing
 Procedure & Checklist - 10 kb Template Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit. This procedure can be used to prepare 10 kb libraries
Procedure & Checklist - 10 kb Template Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit. This procedure can be used to prepare 10 kb libraries
Procedure & Checklist cdna Capture Using IDT xgen Lockdown Probes
 Procedure & Checklist cdna Capture Using IDT xgen Lockdown Probes Before You Begin This document describes the process for capturing cdna prepared with the SMARTer PCR cdna Synthesis Kit (Clontech) and
Procedure & Checklist cdna Capture Using IDT xgen Lockdown Probes Before You Begin This document describes the process for capturing cdna prepared with the SMARTer PCR cdna Synthesis Kit (Clontech) and
Procedure & Checklist Preparing SMRTbell Libraries using PacBio Barcoded Adapters for Multiplex SMRT Sequencing
 Procedure & Checklist Preparing SMRTbell Libraries using PacBio Barcoded Adapters for Multiplex SMRT Sequencing Before You Begin This document describes a procedure for multiplexing 5 Mb microbial genomes
Procedure & Checklist Preparing SMRTbell Libraries using PacBio Barcoded Adapters for Multiplex SMRT Sequencing Before You Begin This document describes a procedure for multiplexing 5 Mb microbial genomes
Procedure & Checklist Multiplex Genomic DNA Target Capture Using IDT xgen Lockdown Probes
 Procedure & Checklist Multiplex Genomic DNA Target Capture Using IDT xgen Lockdown Probes Before You Begin This procedure describes capture and enrichment of regions of interest by using IDT xgen Lockdown
Procedure & Checklist Multiplex Genomic DNA Target Capture Using IDT xgen Lockdown Probes Before You Begin This procedure describes capture and enrichment of regions of interest by using IDT xgen Lockdown
Procedure & Checklist Preparing Multiplexed Microbial SMRTbell Libraries for the PacBio Sequel System
 Procedure & Checklist Preparing Multiplexed Microbial SMRTbell Libraries for the PacBio Sequel System Before You Begin This procedure is for preparing multiplexed SMRTbell libraries for sequencing on the
Procedure & Checklist Preparing Multiplexed Microbial SMRTbell Libraries for the PacBio Sequel System Before You Begin This procedure is for preparing multiplexed SMRTbell libraries for sequencing on the
Target Sequence Capture Using Roche NimbleGen SeqCap EZ Library
 Please note: the shared protocols described herein may not have been validated by Pacific Biosciences and are provided as-is and without any warranty. Use of these protocols is offered to those customers
Please note: the shared protocols described herein may not have been validated by Pacific Biosciences and are provided as-is and without any warranty. Use of these protocols is offered to those customers
NR601. VAHTS TM mrna-seq V2 Library Prep Kit for Illumina
 NR601 VAHTS TM mrna-seq V2 Library Prep Kit for Illumina v Vazyme Biotech Co., Ltd Website: www.vazyme.com Order: global@vazyme.com Support: support@vazyme.com Service: service@vazyme.com SYSTEMS www.vazyme.com
NR601 VAHTS TM mrna-seq V2 Library Prep Kit for Illumina v Vazyme Biotech Co., Ltd Website: www.vazyme.com Order: global@vazyme.com Support: support@vazyme.com Service: service@vazyme.com SYSTEMS www.vazyme.com
Illumina TruSeq Stranded mrna (LT) Protocol 1
 Illumina TruSeq Stranded mrna (LT) Protocol 1 Performed using the TruSeq Stranded mrna Sample Preparation Kit (A cat#fc-122-2101, B cat#fc122-2102) Purify and Fragment mrna NOTE: Use 500ng of Total RNA
Illumina TruSeq Stranded mrna (LT) Protocol 1 Performed using the TruSeq Stranded mrna Sample Preparation Kit (A cat#fc-122-2101, B cat#fc122-2102) Purify and Fragment mrna NOTE: Use 500ng of Total RNA
illumina TruSeq RNA Sample Prep. (LT) Protocol 1
 illumina TruSeq RNA Sample Prep. (LT) Protocol 1 Performed using the TruSeq RNA Sample Preparation Kit (A cat#fc-122-1001, B cat#fc122-1002) Purify and Fragment mrna NOTE: Use 3ug of Total RNA to initiate
illumina TruSeq RNA Sample Prep. (LT) Protocol 1 Performed using the TruSeq RNA Sample Preparation Kit (A cat#fc-122-1001, B cat#fc122-1002) Purify and Fragment mrna NOTE: Use 3ug of Total RNA to initiate
20-kb Template Preparation Using BluePippin Size-Selection System (15-kb Size Cutoff)
 Please note: the shared protocols described herein m may not have been validated by Pacific Biosciences and are provided as-is and w without any warranty. Use of these protocols is offered to those customers
Please note: the shared protocols described herein m may not have been validated by Pacific Biosciences and are provided as-is and w without any warranty. Use of these protocols is offered to those customers
MinION PROTOCOL. Adapted from Janneke Wit by Robyn Tanny May Company Kit/Item Catalog Number
 MinION PROTOCOL Adapted from Janneke Wit by Robyn Tanny May 2016 Company Kit/Item Catalog Number Fisher Eppendorf DNA/RNA LoBind Tubes 13-698-791 Fisher Covaris g-tube NC0380758 NEB NEBNext FFPE Repair
MinION PROTOCOL Adapted from Janneke Wit by Robyn Tanny May 2016 Company Kit/Item Catalog Number Fisher Eppendorf DNA/RNA LoBind Tubes 13-698-791 Fisher Covaris g-tube NC0380758 NEB NEBNext FFPE Repair
VAHTS Stranded mrna-seq Library Prep Kit for Illumina
 Instruction Manual VAHTS Stranded mrna-seq Library Prep Kit for Illumina Vazyme Cat #NR602 Vazyme Biotech Co., Ltd Web: www.vazyme.com Tel: 400-600-9335 Sales: Sales@vazyme.com Support: Support@ vazyme.com
Instruction Manual VAHTS Stranded mrna-seq Library Prep Kit for Illumina Vazyme Cat #NR602 Vazyme Biotech Co., Ltd Web: www.vazyme.com Tel: 400-600-9335 Sales: Sales@vazyme.com Support: Support@ vazyme.com
NEBNext DNA Library Prep Master Mix Set for Illumina
 LIBRARY PREPARATION NEBNext DNA Library Prep Master Mix Set for Illumina Instruction Manual NEB #E6040S/L 12/60 reactions Version 8.0 9/18 be INSPIRED drive DISCOVERY stay GENUINE This product is intended
LIBRARY PREPARATION NEBNext DNA Library Prep Master Mix Set for Illumina Instruction Manual NEB #E6040S/L 12/60 reactions Version 8.0 9/18 be INSPIRED drive DISCOVERY stay GENUINE This product is intended
USER GUIDE For Illumina Platform
 USER GUIDE For Illumina Platform Copyright Nimagen B.V. P.O. Box 91 6500 AB Nijmegen The Netherlands Tel. +31 (0)24 820 0241 Fax. +31 (0)24 358 0259 info@nimagen.com VAT#: NL850011243B01 Rabobank Nijmegen:
USER GUIDE For Illumina Platform Copyright Nimagen B.V. P.O. Box 91 6500 AB Nijmegen The Netherlands Tel. +31 (0)24 820 0241 Fax. +31 (0)24 358 0259 info@nimagen.com VAT#: NL850011243B01 Rabobank Nijmegen:
MEDIP-SEQUENCING PROTOCOL
 MEDIP-SEQUENCING PROTOCOL MAGMEDIP KIT Cat. No. C02010020 Table 1 The GenDNA module provides you with an excess of buffer for the preparation of DNA. Sufficient buffer is given for the preparation of several
MEDIP-SEQUENCING PROTOCOL MAGMEDIP KIT Cat. No. C02010020 Table 1 The GenDNA module provides you with an excess of buffer for the preparation of DNA. Sufficient buffer is given for the preparation of several
Stranded mrna-seq Lib Prep Kit for Illumina
 Stranded mrna-seq Lib Prep Kit for Illumina RK20301 (10ng-1ug Input Total RNA) (Illumina Compatible) C U G A www.abclonal.com version: N12G13v1.0 Contents 1.Introduction 01 2.Components 02 3.Additional
Stranded mrna-seq Lib Prep Kit for Illumina RK20301 (10ng-1ug Input Total RNA) (Illumina Compatible) C U G A www.abclonal.com version: N12G13v1.0 Contents 1.Introduction 01 2.Components 02 3.Additional
xgen hybridization capture of DNA libraries
 xgen hybridization capture of DNA libraries For NGS target enrichment Uses IDT s: xgen Hybridization and Wash Kit xgen Universal Blockers TS Mix, 10 bp TS Mix, or NXT Mix xgen Lockdown Panels and Probes
xgen hybridization capture of DNA libraries For NGS target enrichment Uses IDT s: xgen Hybridization and Wash Kit xgen Universal Blockers TS Mix, 10 bp TS Mix, or NXT Mix xgen Lockdown Panels and Probes
We want to thank and acknowledge the authors for sharing this protocol and their contributions to the field.
 We adopted the protocol described in the Extended Experimental Procedures section I.a.1 of the 2014 Cell paper by Rao and Huntley et. al: A 3D Map of the Human Genome at Kilobase Resolution Reveals Principles
We adopted the protocol described in the Extended Experimental Procedures section I.a.1 of the 2014 Cell paper by Rao and Huntley et. al: A 3D Map of the Human Genome at Kilobase Resolution Reveals Principles
Select-a-Size DNA Clean & Concentrator MagBead Kit Catalog No. D4084 & D4085
 INSTRUCTION MANUAL Select-a-Size DNA Clean & Concentrator MagBead Kit Catalog No. D4084 & D4085 Highlights Tunable: Size selection can be tuned from 100 bp to 1000 bp with left, right, or double size selection
INSTRUCTION MANUAL Select-a-Size DNA Clean & Concentrator MagBead Kit Catalog No. D4084 & D4085 Highlights Tunable: Size selection can be tuned from 100 bp to 1000 bp with left, right, or double size selection
Additional reagents and materials that are not supplied
 sparq PureMag Beads Cat. No. 95196-005 Size: 5 ml Store at 2 C to 8 C 95196-060 60 ml 95196-450 450 ml Description sparq PureMag Beads uses reversible nucleic acid-binding properties of magnetic beads
sparq PureMag Beads Cat. No. 95196-005 Size: 5 ml Store at 2 C to 8 C 95196-060 60 ml 95196-450 450 ml Description sparq PureMag Beads uses reversible nucleic acid-binding properties of magnetic beads
Alon s SCN ChIP Protocol
 Prior to starting your ChIPs and Shearing 1. Turn on sonifiers and cooling system allow system to reach -1 C before shearing 2. Cool bench top centrifuge to 4 C 3. Prepare all of your buffers with protease
Prior to starting your ChIPs and Shearing 1. Turn on sonifiers and cooling system allow system to reach -1 C before shearing 2. Cool bench top centrifuge to 4 C 3. Prepare all of your buffers with protease
NxSeq UltraLow DNA Library Kit, 12 Reactions
 NxSeq UltraLow DNA Library Kit, 12 Reactions Illumina-compatible FOR RESEARCH USE ONLY. NOT FOR HUMAN OR DIAGNOSTIC USE Lucigen Corporation 2905 Parmenter St, Middleton, WI 53562 USA Toll Free: (888) 575-9695
NxSeq UltraLow DNA Library Kit, 12 Reactions Illumina-compatible FOR RESEARCH USE ONLY. NOT FOR HUMAN OR DIAGNOSTIC USE Lucigen Corporation 2905 Parmenter St, Middleton, WI 53562 USA Toll Free: (888) 575-9695
JetSeq DNA Library Preparation Kit. Product Manual
 JetSeq DNA Library Preparation Kit Product Manual 2 JetSeq DNA Library Preparation Kit JetSeq DNA Library Preparation Kit TABLE OF CONTENTS 1 Kit contents 04 2 Description 05 3 Storage 06 4 Safety information
JetSeq DNA Library Preparation Kit Product Manual 2 JetSeq DNA Library Preparation Kit JetSeq DNA Library Preparation Kit TABLE OF CONTENTS 1 Kit contents 04 2 Description 05 3 Storage 06 4 Safety information
Microwell-Seq. High-throughput Single Cell RNA-Seq Kit. Protocol. Index
 Microwell-Seq High-throughput Single Cell RNA-Seq Kit Protocol Index 1 Introduction 2 Kit Reagent 3 Store 4 Application 5 Prepared materials 6 Note 7 Preparation 8 Workflow 1 1 Introduction Microwell-Seq
Microwell-Seq High-throughput Single Cell RNA-Seq Kit Protocol Index 1 Introduction 2 Kit Reagent 3 Store 4 Application 5 Prepared materials 6 Note 7 Preparation 8 Workflow 1 1 Introduction Microwell-Seq
Project: RADseqReady Plate # Library # Name: Date: Section 1: DNA Standardization
 BestRAD Library Preparation Based on protocol of Ali et al. 2015 (10.1534/genetics.115.183665) Adapted by Linda Rutledge many times but this version was done on August 16, 2016 Section 1: DNA Standardization
BestRAD Library Preparation Based on protocol of Ali et al. 2015 (10.1534/genetics.115.183665) Adapted by Linda Rutledge many times but this version was done on August 16, 2016 Section 1: DNA Standardization
DNA Size Selection Magnetic Beads
 DNA Size Selection Magnetic Beads Catalog #: 801-117 User Manual Last revised July 30 th, 2018 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
DNA Size Selection Magnetic Beads Catalog #: 801-117 User Manual Last revised July 30 th, 2018 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
ChIP Protocol for fresh or frozen cross linked cells
 Prior to starting your ChIPs and Shearing Turn on sonifiers and cooling system allow system to reach -2 C before shearing Cool bench top centrifuge to 4 C Prepare all of your buffers with protease inhibitors
Prior to starting your ChIPs and Shearing Turn on sonifiers and cooling system allow system to reach -2 C before shearing Cool bench top centrifuge to 4 C Prepare all of your buffers with protease inhibitors
User Guide. rrna Depletion Kit V1.2
 rrna Depletion Kit V1.2 User Guide Catalog Number: 037 (RiboCop rrna Depletion Kit V1.2 (Human/Mouse/Rat)) 042 (SENSE Total RNA-Seq Library Prep Kit for Illumina with RiboCop) 037UG073V0201 FOR RESEARCH
rrna Depletion Kit V1.2 User Guide Catalog Number: 037 (RiboCop rrna Depletion Kit V1.2 (Human/Mouse/Rat)) 042 (SENSE Total RNA-Seq Library Prep Kit for Illumina with RiboCop) 037UG073V0201 FOR RESEARCH
RayBio mrna Magnetic Beads Kit
 RayBio mrna Magnetic Beads Kit Catalog #: 801-116 User Manual Last revised March 9 th, 2017 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
RayBio mrna Magnetic Beads Kit Catalog #: 801-116 User Manual Last revised March 9 th, 2017 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
JetSeq Clean. Product Manual
 JetSeq Clean Product Manual 2 Product Manual bioline.com/jetseq JetSeq Clean JetSeq Clean TABLE OF CONTENTS 1 Kit contents 04 2 Description 05 3 Equipment and reagents to be supplied by user 06 4 Storage
JetSeq Clean Product Manual 2 Product Manual bioline.com/jetseq JetSeq Clean JetSeq Clean TABLE OF CONTENTS 1 Kit contents 04 2 Description 05 3 Equipment and reagents to be supplied by user 06 4 Storage
Swift Hybridization Capture Kits
 Protocol Swift Hybridization Capture Kits For NGS Target Enrichment Uses: Swift Exome Hyb Panel, Cat. No. 83216 Swift Pan-Cancer Hyb Panel, Cat. No. 83316 Swift Inherited Diseases Hyb Panel, Cat. No. 83416
Protocol Swift Hybridization Capture Kits For NGS Target Enrichment Uses: Swift Exome Hyb Panel, Cat. No. 83216 Swift Pan-Cancer Hyb Panel, Cat. No. 83316 Swift Inherited Diseases Hyb Panel, Cat. No. 83416
empcr Amplification Method Manual Lib L LV
 empcr Amplification Method Manual Lib L LV GS FLX+ Series XL+ May 2011 For life science research only. Not for use in diagnostic procedures. Instrument / Kit GS Junior / Junior GS FLX+ / XL+ GS FLX+ /
empcr Amplification Method Manual Lib L LV GS FLX+ Series XL+ May 2011 For life science research only. Not for use in diagnostic procedures. Instrument / Kit GS Junior / Junior GS FLX+ / XL+ GS FLX+ /
Mayr lab 3'-seq protocol, October 2013
 Mayr lab 3'-seq protocol, October 2013 Time line: Day -1: DNase treat your RNA samples (optional) Day 1: Prepare beads, anneal oligo to beads, 1 st, 2 nd strand synthesis Day 2: Introduce nick (Rnase HII),
Mayr lab 3'-seq protocol, October 2013 Time line: Day -1: DNase treat your RNA samples (optional) Day 1: Prepare beads, anneal oligo to beads, 1 st, 2 nd strand synthesis Day 2: Introduce nick (Rnase HII),
Evaluation of Omega Mag-Bind TotalPure NGS Beads for DNA Size Selection
 Evaluation of Omega Mag-Bind TotalPure NGS Beads for Size Selection By Maggie Weitzman, M.Sc. (University of Oregon / GC3F) Disclaimer: Neither Maggie Weitzman, the University of Oregon, nor the Genomics
Evaluation of Omega Mag-Bind TotalPure NGS Beads for Size Selection By Maggie Weitzman, M.Sc. (University of Oregon / GC3F) Disclaimer: Neither Maggie Weitzman, the University of Oregon, nor the Genomics
DNA extraction Protocol for Agencourt Genfind v2 Blood and Serum Genomic DNA Isolation Kit
 DNA extraction Protocol for Agencourt Genfind v2 Blood and Serum Genomic DNA Isolation Kit Introduction The Agencourt Genfind v2 Blood & Serum DNA Isolation Kit utilizes Agencourt s patented SPRI paramagnetic
DNA extraction Protocol for Agencourt Genfind v2 Blood and Serum Genomic DNA Isolation Kit Introduction The Agencourt Genfind v2 Blood & Serum DNA Isolation Kit utilizes Agencourt s patented SPRI paramagnetic
NGS clean-up and size selection
 NGS clean-up and size selection User manual NucleoMag NGS Clean-up and Size Select May 2014 / Rev. 01 NGS clean-up and size selection Table of contents 1 Components 4 1.1 Kit contents 4 1.2 Equipment and
NGS clean-up and size selection User manual NucleoMag NGS Clean-up and Size Select May 2014 / Rev. 01 NGS clean-up and size selection Table of contents 1 Components 4 1.1 Kit contents 4 1.2 Equipment and
mi-mag mrna Isolation Kit
 mi-mag mrna Isolation Kit Cat. No [50 Reactions] This kit is for research purposes only. Not for use in diagnostic procedures. For in vitro use only. Introduction This kit contains enough materials for
mi-mag mrna Isolation Kit Cat. No [50 Reactions] This kit is for research purposes only. Not for use in diagnostic procedures. For in vitro use only. Introduction This kit contains enough materials for
User Guide. rrna Depletion Kit
 rrna Depletion Kit User Guide Catalog Number: 037.24 (RiboCop rrna Depletion Kit (Human/Mouse/Rat), 24 preps) 037.96 (RiboCop rrna Depletion Kit (Human/Mouse/Rat), 96 preps) 042.08 (SENSE Total RNA-Seq
rrna Depletion Kit User Guide Catalog Number: 037.24 (RiboCop rrna Depletion Kit (Human/Mouse/Rat), 24 preps) 037.96 (RiboCop rrna Depletion Kit (Human/Mouse/Rat), 96 preps) 042.08 (SENSE Total RNA-Seq
CleanPlex UMI NGS Panel
 CleanPlex UMI NGS Panel User Guide This user guide is for the following products: CleanPlex UMI Lung Cancer Panel CleanPlex UMI Custom NGS Panel Get the latest user guide at: www.paragongenomics.com/product_documents/
CleanPlex UMI NGS Panel User Guide This user guide is for the following products: CleanPlex UMI Lung Cancer Panel CleanPlex UMI Custom NGS Panel Get the latest user guide at: www.paragongenomics.com/product_documents/
AGENCOURT GENFIND Blood & Serum Genomic DNA Isolation Kit
 Blood & Serum Genomic DNA Isolation Kit Page 1 of 9 Please refer to http://www.agencourt.com/technical for updated protocols and refer to MSDS instructions when handling or shipping any chemical hazards.
Blood & Serum Genomic DNA Isolation Kit Page 1 of 9 Please refer to http://www.agencourt.com/technical for updated protocols and refer to MSDS instructions when handling or shipping any chemical hazards.
Ampli1 LowPass Kit. USER MANUAL Version 3.0. Low-pass WGS library prep kit for IonTorrent platforms. Content version: July 2017
 For research use only. Not for use in diagnostic procedures. For in vitro use only. Ampli1 LowPass Kit Low-pass WGS library prep kit for IonTorrent platforms USER MANUAL Version 3.0 Content version: July
For research use only. Not for use in diagnostic procedures. For in vitro use only. Ampli1 LowPass Kit Low-pass WGS library prep kit for IonTorrent platforms USER MANUAL Version 3.0 Content version: July
Ampli1 LowPass Kit. USER MANUAL Version 2.0. Low-pass WGS library prep kit for IonTorrent platforms. Content version: January 2017
 For research use only. Not for use in diagnostic procedures. For in vitro use only. Ampli1 LowPass Kit Low-pass WGS library prep kit for IonTorrent platforms USER MANUAL Version 2.0 Content version: January
For research use only. Not for use in diagnostic procedures. For in vitro use only. Ampli1 LowPass Kit Low-pass WGS library prep kit for IonTorrent platforms USER MANUAL Version 2.0 Content version: January
EPIGENTEK. EpiQuik Circulating Cell-Free DNA Isolation Kit. Base Catalog # P-1064 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Circulating Cell-Free DNA Isolation Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiQuik Circulating Cell-Free DNA Isolation Kit utilizes magnetic beads based sizefractionation
EpiQuik Circulating Cell-Free DNA Isolation Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiQuik Circulating Cell-Free DNA Isolation Kit utilizes magnetic beads based sizefractionation
Formaldehyde Cross-linking of Chromatin from Drosophila
 2 Formaldehyde Cross-linking of Chromatin from Drosophila Protocol from modencode IGSB University of Chicago originally written by Alex Crofts and Sasha Ostapenko and updated by Matt Kirkey. 1. Set centrifuge
2 Formaldehyde Cross-linking of Chromatin from Drosophila Protocol from modencode IGSB University of Chicago originally written by Alex Crofts and Sasha Ostapenko and updated by Matt Kirkey. 1. Set centrifuge
Applied Biosystems SOLiD 4 System Templated Bead Preparation Guide
 Applied Biosystems SOLiD 4 System Templated Bead Preparation Guide March 2010 Library Preparation Templated Bead Preparation Instrument Operation For Research Use Only. Not intended for any animal or human
Applied Biosystems SOLiD 4 System Templated Bead Preparation Guide March 2010 Library Preparation Templated Bead Preparation Instrument Operation For Research Use Only. Not intended for any animal or human
In nucleus Hi- C protocol for C. elegans embryos
 In nucleus Hi- C protocol for C. elegans embryos Compiled by Erika Anderson, July 2016. Crosslinking, isolating nuclei, and digestion 1. Bleach gravid hermaphrodites to obtain at least 0.5g of embryos.
In nucleus Hi- C protocol for C. elegans embryos Compiled by Erika Anderson, July 2016. Crosslinking, isolating nuclei, and digestion 1. Bleach gravid hermaphrodites to obtain at least 0.5g of embryos.
AGENCOURT ORAPURE Buccal Cell DNA Isolation Kit
 Buccal Cell DNA Isolation Kit Page 1 of 12 Please refer to http://www.agencourt.com/technical for updated protocols and refer to MSDS instructions when handling or shipping any chemical hazards. AGENCOURT
Buccal Cell DNA Isolation Kit Page 1 of 12 Please refer to http://www.agencourt.com/technical for updated protocols and refer to MSDS instructions when handling or shipping any chemical hazards. AGENCOURT
Solutions for purifying nucleic acids by solidphase reversible immobilization (SPRI)
 Solutions for purifying nucleic acids by solidphase reversible immobilization (SPRI) Philippe Jolivet and Joseph W. Foley Ludmer Centre for Neuroinformatics and Mental Health October 21, 2015 Based on
Solutions for purifying nucleic acids by solidphase reversible immobilization (SPRI) Philippe Jolivet and Joseph W. Foley Ludmer Centre for Neuroinformatics and Mental Health October 21, 2015 Based on
FastTrack MAG mrna Isolation Kits
 USER GUIDE FastTrack MAG mrna Isolation Kits For isolating high-quality mrna from total RNA, cells, and tissue Catalog Numbers K1580-01 and K1580-02 Document Part Number 25-0754 Publication Number MAN0000475
USER GUIDE FastTrack MAG mrna Isolation Kits For isolating high-quality mrna from total RNA, cells, and tissue Catalog Numbers K1580-01 and K1580-02 Document Part Number 25-0754 Publication Number MAN0000475
Mag-Bind cfdna Kit. M preps M preps M Preps
 Mag-Bind cfdna Kit M3298-00 5 preps M3298-01 50 preps M3298-02 200 Preps March 2018 Mag-Bind cfdna Kit Table of Contents Introduction and Overview...2 Kit Contents/Storage and Stability...3 Preparing Reagents...4
Mag-Bind cfdna Kit M3298-00 5 preps M3298-01 50 preps M3298-02 200 Preps March 2018 Mag-Bind cfdna Kit Table of Contents Introduction and Overview...2 Kit Contents/Storage and Stability...3 Preparing Reagents...4
Poly(A) RNA Selection Kit User Guide
 Poly(A) RNA Selection Kit User Guide 039 (Poly(A) RNA Selection Kit) 009 (SENSE Total RNA-Seq Library Prep Kit for Illumina, including Barcodes) 020 (PCR Add-on Kit for Illumina) 022 (Purification Module
Poly(A) RNA Selection Kit User Guide 039 (Poly(A) RNA Selection Kit) 009 (SENSE Total RNA-Seq Library Prep Kit for Illumina, including Barcodes) 020 (PCR Add-on Kit for Illumina) 022 (Purification Module
USER GUIDE. Ovation PART NO. 0344, 0344NB. Ultralow System V2
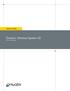 USER GUIDE Ovation PART NO. 0344, 0344NB Ultralow System V2 Patents, Licensing and Trademarks 2014 2017 NuGEN Technologies, Inc. All rights reserved. The Encore, Ovation and Applause families of products
USER GUIDE Ovation PART NO. 0344, 0344NB Ultralow System V2 Patents, Licensing and Trademarks 2014 2017 NuGEN Technologies, Inc. All rights reserved. The Encore, Ovation and Applause families of products
Direct Polysome IP from Brain Tissue Myriam Heiman:
 Direct Polysome IP from Brain Tissue Myriam Heiman: bonillm@rockefeller.edu Protocol below is for 1 IP, scale accordingly General Notes: -7 mouse striata pooled per IP -IP with 50 µg 19C8 and 50 µg 19F7
Direct Polysome IP from Brain Tissue Myriam Heiman: bonillm@rockefeller.edu Protocol below is for 1 IP, scale accordingly General Notes: -7 mouse striata pooled per IP -IP with 50 µg 19C8 and 50 µg 19F7
Ribo-Zero Magnetic Kits*
 * Ribo-Zero Kit Catalog Number 6-Reactions Catalog Number 24-Reactions Ribo-Zero Magnetic Gold Kit (Human/Mouse/Rat) MRZG126 MRZG12324 Ribo-Zero Magnetic Kit (Human/Mouse/Rat) MRZH116 MRZH11124 Ribo-Zero
* Ribo-Zero Kit Catalog Number 6-Reactions Catalog Number 24-Reactions Ribo-Zero Magnetic Gold Kit (Human/Mouse/Rat) MRZG126 MRZG12324 Ribo-Zero Magnetic Kit (Human/Mouse/Rat) MRZH116 MRZH11124 Ribo-Zero
SOLiD EZ Bead Emulsifier
 QUICK REFERENCE SOLiD EZ Bead Emulsifier Pub. Part no. 4441487 Rev. E Rev. Date October 2011 Note: For safety guidelines, refer to the Safety section in the Applied Biosystems SOLiD EZ Bead Emulsifier
QUICK REFERENCE SOLiD EZ Bead Emulsifier Pub. Part no. 4441487 Rev. E Rev. Date October 2011 Note: For safety guidelines, refer to the Safety section in the Applied Biosystems SOLiD EZ Bead Emulsifier
TrueBlot Protein G Magnetic Beads PG ml. TrueBlot Protein G Magnetic Beads PG ml. Bead Mean Diameter 0.5 µm. Bead Concentration
 Rockland s TrueBlot Protein G Magnetic Beads are uniform, non-aggregating, super-paramagnetic beads coupled with a biomolecule, such as Protein G. These beads are specifically designed, tested and quality
Rockland s TrueBlot Protein G Magnetic Beads are uniform, non-aggregating, super-paramagnetic beads coupled with a biomolecule, such as Protein G. These beads are specifically designed, tested and quality
AdnaTest EMT-1/StemCellSelect
 AdnaTest EMT-1/StemCellSelect Enrichment of tumor cells from blood for gene expression analysis For research use only Manual T-1-533 Contents Order Information... 3 Purpose... 3 Abbreviations and Symbols...
AdnaTest EMT-1/StemCellSelect Enrichment of tumor cells from blood for gene expression analysis For research use only Manual T-1-533 Contents Order Information... 3 Purpose... 3 Abbreviations and Symbols...
Drop-seq Protocol, 2015 v. 1.0 (5/21/15)
 page 1 Drop-seq Protocol, 2015 v. 1.0 (5/21/15) Evan Macosko & Melissa Goldman Before you begin: The following is the most up-to-date protocol for Drop-seq, including all equipment and reagents used. Please
page 1 Drop-seq Protocol, 2015 v. 1.0 (5/21/15) Evan Macosko & Melissa Goldman Before you begin: The following is the most up-to-date protocol for Drop-seq, including all equipment and reagents used. Please
ChargeSwitch gdna Rendered Meat Purification Kit
 USER GUIDE ChargeSwitch gdna Rendered Meat Purification Kit Purification of genomic DNA (gdna) from cattle feed, meal, and heparin products Catalog Number CS400-100 Publication Number MAN0000574 Revision
USER GUIDE ChargeSwitch gdna Rendered Meat Purification Kit Purification of genomic DNA (gdna) from cattle feed, meal, and heparin products Catalog Number CS400-100 Publication Number MAN0000574 Revision
AdnaTest OvarianCancer-2 Select
 AdnaTest OvarianCancer-2 Select Enrichment of tumor cells from blood of ovarian cancer patients for gene expression analysis For research use only Manual T-1-538 Contents Order Information... 3 Purpose...
AdnaTest OvarianCancer-2 Select Enrichment of tumor cells from blood of ovarian cancer patients for gene expression analysis For research use only Manual T-1-538 Contents Order Information... 3 Purpose...
PARP1-Trap_A for Immunoprecipitation of PARP1- Fusion Proteins from cell extract
 PARP1-Trap_A for Immunoprecipitation of PARP1- Fusion Proteins from cell extract Only for research applications, not for diagnostic or therapeutic use. Introduction Specificity Poly(ADP-ribose) polymerase
PARP1-Trap_A for Immunoprecipitation of PARP1- Fusion Proteins from cell extract Only for research applications, not for diagnostic or therapeutic use. Introduction Specificity Poly(ADP-ribose) polymerase
ChargeSwitch NoSpin Plasmid Kits
 USER GUIDE ChargeSwitch NoSpin Plasmid Kits For purification of plasmid DNA from bacterial cells using the MagnaClear Technology Catalog nos. CS10200, CS10201, CS10201-10 Version A 5 January 2005 25-0813
USER GUIDE ChargeSwitch NoSpin Plasmid Kits For purification of plasmid DNA from bacterial cells using the MagnaClear Technology Catalog nos. CS10200, CS10201, CS10201-10 Version A 5 January 2005 25-0813
ISOFECAL for Beads Beating Manual (First edition)
 Fecal DNA Extraction Kit ISOFECAL for Beads Beating Manual (First edition) Code No. 315-06281 NIPPON GENE CO., LTD. Table of contents I Product description 1 II Contents of kit 1 III Storage 2 IV Precautions
Fecal DNA Extraction Kit ISOFECAL for Beads Beating Manual (First edition) Code No. 315-06281 NIPPON GENE CO., LTD. Table of contents I Product description 1 II Contents of kit 1 III Storage 2 IV Precautions
BoTest Matrix E Botulinum Neurotoxin Detection Kit Protocol
 BoTest Matrix E Botulinum Neurotoxin Detection Kit Protocol 505 S. Rosa Road, Suite 105 Madison, WI 53719 1-608-441-8174 info@biosentinelpharma.com BioSentinel Part No: L1016, Release Date: May 29, 2014
BoTest Matrix E Botulinum Neurotoxin Detection Kit Protocol 505 S. Rosa Road, Suite 105 Madison, WI 53719 1-608-441-8174 info@biosentinelpharma.com BioSentinel Part No: L1016, Release Date: May 29, 2014
Enriching Beads Oligo (dt) Magnetic Beads for mrna Purification
 Enriching Beads Oligo (dt) Magnetic Beads for mrna Purification Isolate the mrna transcriptome in 15 minutes User Guidance Enriching Biotechnology Rev. 1.0 October 25th. 2018 Why choose Enriching Beads
Enriching Beads Oligo (dt) Magnetic Beads for mrna Purification Isolate the mrna transcriptome in 15 minutes User Guidance Enriching Biotechnology Rev. 1.0 October 25th. 2018 Why choose Enriching Beads
LOABeads AffiAmino. Product Manual. Lab on a Bead AB. Revision date Copyright Lab on a Bead AB All rights reserved
 LOABeads AffiAmino Product Manual Lab on a Bead AB Revision date 2016-11-23 Copyright 2015-2016 Lab on a Bead AB All rights reserved Table of Contents 1. General information...3 2. Product data...4 3.
LOABeads AffiAmino Product Manual Lab on a Bead AB Revision date 2016-11-23 Copyright 2015-2016 Lab on a Bead AB All rights reserved Table of Contents 1. General information...3 2. Product data...4 3.
chemagic mrna Direct Kit
 chemagic mrna Direct Kit for general purposes Kit for the direct isolation of mrna from animal and plant tissue and cells. Kit Components M-PVA OdT Magnetic Beads Suspension Buffer 1 Lysis Buffer 2 Wash
chemagic mrna Direct Kit for general purposes Kit for the direct isolation of mrna from animal and plant tissue and cells. Kit Components M-PVA OdT Magnetic Beads Suspension Buffer 1 Lysis Buffer 2 Wash
Technical Manual No. TM0261 Version
 Donkey Anti-Goat IgG MagBeads Cat. No. L00332 Technical Manual No. TM0261 Version 06272010 Index 1. Product Description 2. Instruction For Use 3. Troubleshooting 4. General Information 1. Product Description
Donkey Anti-Goat IgG MagBeads Cat. No. L00332 Technical Manual No. TM0261 Version 06272010 Index 1. Product Description 2. Instruction For Use 3. Troubleshooting 4. General Information 1. Product Description
BIOL110L-Cell Biology Lab Spring Quarter 2012 Module 3-4 Wednesday May 30, 2012
 BIOL110L-Cell Biology Lab Spring Quarter 2012 Module 3-4 Wednesday May 30, 2012 PART I: Isolation of over-expressed GFP-Karyopherins from yeast extracts by affinity capture on nucleoporins Summary: The
BIOL110L-Cell Biology Lab Spring Quarter 2012 Module 3-4 Wednesday May 30, 2012 PART I: Isolation of over-expressed GFP-Karyopherins from yeast extracts by affinity capture on nucleoporins Summary: The
GS FLX/Junior Titanium Technology
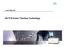 www.454.com GS FLX/Junior Titanium Technology GS FLX and GS Junior Process Steps Overview gdna 1. DNA Library Construction * 4h 2. empcr 3. Sequencing 8 h 10 h Data output DNA Library Preparation Prepare
www.454.com GS FLX/Junior Titanium Technology GS FLX and GS Junior Process Steps Overview gdna 1. DNA Library Construction * 4h 2. empcr 3. Sequencing 8 h 10 h Data output DNA Library Preparation Prepare
Controller Training Kit User Guide
 Library Construction Chromium Controller Training Kit User Guide FOR USE WITH Chromium Controller & Training Reagents, Gel Bead & Chip Kits PN-120245 NOTICES Notices Manual Part Number CG00021 Rev A Legal
Library Construction Chromium Controller Training Kit User Guide FOR USE WITH Chromium Controller & Training Reagents, Gel Bead & Chip Kits PN-120245 NOTICES Notices Manual Part Number CG00021 Rev A Legal
Drop- Seq Laboratory Protocol
 page 1 Drop- Seq Laboratory Protocol version 1.0 (5/21/15) Evan Macosko and Melissa Goldman Steve McCarroll s lab, Harvard Medical School The following is our current protocol for Drop- seq. It should
page 1 Drop- Seq Laboratory Protocol version 1.0 (5/21/15) Evan Macosko and Melissa Goldman Steve McCarroll s lab, Harvard Medical School The following is our current protocol for Drop- seq. It should
RayBio anti-mouse IgG Magnetic Beads
 RayBio anti-mouse IgG Magnetic Beads Catalog #: 801-103 User Manual Last revised January 4 th, 2017 Caution: Extraordinarily useful information enclosed ISO 1348 Certified 3607 Parkway Lane, Suite 100
RayBio anti-mouse IgG Magnetic Beads Catalog #: 801-103 User Manual Last revised January 4 th, 2017 Caution: Extraordinarily useful information enclosed ISO 1348 Certified 3607 Parkway Lane, Suite 100
Drop- Seq Laboratory Protocol
 page 1 Drop- Seq Laboratory Protocol version 3.1 (12/28/15) Evan Macosko and Melissa Goldman Steve McCarroll s lab, Harvard Medical School The following is our most current protocol for Drop- seq. It should
page 1 Drop- Seq Laboratory Protocol version 3.1 (12/28/15) Evan Macosko and Melissa Goldman Steve McCarroll s lab, Harvard Medical School The following is our most current protocol for Drop- seq. It should
Nature Protocols: doi: /nprot Supplementary Figure 1. Read quality score per sequenced base position.
 Supplementary Figure 1 Read quality score per sequenced base position. (a) Illumina Q-scores for a normal QiSeq run. Typically, more than 80% of reads reach a quality score higher than 30 across all read
Supplementary Figure 1 Read quality score per sequenced base position. (a) Illumina Q-scores for a normal QiSeq run. Typically, more than 80% of reads reach a quality score higher than 30 across all read
ChargeSwitch gdna Blood Kits
 Instruction Manual ChargeSwitch gdna Blood Kits For purification of genomic DNA from small volumes of human blood Catalog nos. CS11000, CS11010, and CS11010-10 Version A 6 January 2005 25-0814 ii Table
Instruction Manual ChargeSwitch gdna Blood Kits For purification of genomic DNA from small volumes of human blood Catalog nos. CS11000, CS11010, and CS11010-10 Version A 6 January 2005 25-0814 ii Table
30 Plex Human Luminex (Invitrogen Kit, Single Plate)
 30 Plex Human Luminex (Invitrogen Kit, Single Plate) 1. Defrost samples and bring to room temperature. 2. Bring Kit components to room temperature: Wash solution 20x. Assay Diluent. Incubation buffer.
30 Plex Human Luminex (Invitrogen Kit, Single Plate) 1. Defrost samples and bring to room temperature. 2. Bring Kit components to room temperature: Wash solution 20x. Assay Diluent. Incubation buffer.
CELLSCRIPT RNA for Translation in Cells
 H TM CELLSCRIPT RA for Translation in Cells Cat. o. C-C61025 ITRDUCTI 5'-terminal caps are involved in mra processing, stability and initiation of protein synthesis. 1 Uncapped RA transfected or injected
H TM CELLSCRIPT RA for Translation in Cells Cat. o. C-C61025 ITRDUCTI 5'-terminal caps are involved in mra processing, stability and initiation of protein synthesis. 1 Uncapped RA transfected or injected
LOABeads MagSep 15/50 LOABeads MagSep 500
 LOABeads MagSep 15/50 LOABeads MagSep 500 Product Manual Lab on a Bead AB Edition 2016-05-09 Copyright 2015-2016 Lab on a Bead AB Table of Contents 1. 2. 3. 4. 5. 6. Safety instructions...3 General handling
LOABeads MagSep 15/50 LOABeads MagSep 500 Product Manual Lab on a Bead AB Edition 2016-05-09 Copyright 2015-2016 Lab on a Bead AB Table of Contents 1. 2. 3. 4. 5. 6. Safety instructions...3 General handling
pluribead KIT Cell Separation Protocol M-Bead
 pluribead KIT Cell Separation Protocol M-Bead pluriselect@hiss-dx.de 15 14 13 12 11 9 8 7 6 5 4 3 2 1 50 40 30 20 2 Contents Contents pluribead Kit Components & Additional Materials 2 Separation Protocol
pluribead KIT Cell Separation Protocol M-Bead pluriselect@hiss-dx.de 15 14 13 12 11 9 8 7 6 5 4 3 2 1 50 40 30 20 2 Contents Contents pluribead Kit Components & Additional Materials 2 Separation Protocol
LOABeads Protein A. Product no Product Manual. Lab on a Bead AB
 Product no 1001 LOABeads Protein A Product Manual Lab on a Bead AB Revision date 2016-03-08 Copyright 2015-2016 Lab on a Bead AB All rights reserved Table of Contents 1. General information 3 2. Antibody
Product no 1001 LOABeads Protein A Product Manual Lab on a Bead AB Revision date 2016-03-08 Copyright 2015-2016 Lab on a Bead AB All rights reserved Table of Contents 1. General information 3 2. Antibody
Total RNA Isolation. User Manual. NucleoMag 96 RNA MACHEREY-NAGEL. January 2010 / Rev. 02
 Total RNA Isolation User Manual NucleoMag 96 RNA January 2010 / Rev. 02 MACHEREY-NAGEL Table of contents 1 Components 4 1.1 Kit contents 4 1.2 Material to be supplied by user 5 2 Product description 6
Total RNA Isolation User Manual NucleoMag 96 RNA January 2010 / Rev. 02 MACHEREY-NAGEL Table of contents 1 Components 4 1.1 Kit contents 4 1.2 Material to be supplied by user 5 2 Product description 6
M-Beads Magnetic silica beads DNA 3.0 (COOH) Order #: PR-MAG00078 & PR-MAG00079
 M-Beads Magnetic silica beads DNA 3.0 (COOH) Order #: PR-MAG00078 & PR-MAG00079 MoBiTec GmbH 2015 Page 2 Contents Intended Use... 3 Principle... 3 Silica & Carboxylated M-Beads Magnetic silica beads DNA
M-Beads Magnetic silica beads DNA 3.0 (COOH) Order #: PR-MAG00078 & PR-MAG00079 MoBiTec GmbH 2015 Page 2 Contents Intended Use... 3 Principle... 3 Silica & Carboxylated M-Beads Magnetic silica beads DNA
Buffers. Enzymes. Beads
 Buffers iclip lysis buffer 50 mm Tris-HCl ph 7.4 100 mm NaCl 1% NP-40 (Igepal CA630) 0.1% SDS 0.5% sodium deoxycholate (protect from light) 1:200 Protease Inhibitor Cocktail III (add fresh) High salt wash
Buffers iclip lysis buffer 50 mm Tris-HCl ph 7.4 100 mm NaCl 1% NP-40 (Igepal CA630) 0.1% SDS 0.5% sodium deoxycholate (protect from light) 1:200 Protease Inhibitor Cocktail III (add fresh) High salt wash
Strep-tag Purification using MagStrep type3 XT Beads
 Strep-tag Purification using MagStrep type3 XT Beads Instruction manual Last date of revision November 2016 Version PR83-0004 For research only Important licensing information Products featuring Strep-Tactin
Strep-tag Purification using MagStrep type3 XT Beads Instruction manual Last date of revision November 2016 Version PR83-0004 For research only Important licensing information Products featuring Strep-Tactin
