JetSeq DNA Library Preparation Kit. Product Manual
|
|
|
- Adelia Dennis
- 5 years ago
- Views:
Transcription
1 JetSeq DNA Library Preparation Kit Product Manual
2 2
3 JetSeq DNA Library Preparation Kit JetSeq DNA Library Preparation Kit TABLE OF CONTENTS 1 Kit contents 04 2 Description 05 3 Storage 06 4 Safety information 06 5 Product specifications 06 6 Equipment and reagents to be supplied by user 08 7 Important notes Recommended DNA preparation methods Recommendations for DNA fragmentation Recommendations for bead-based clean-up and size selection Recommendations for quality control throughout the library preparation 09 8 Protocol Protocol A (target enrichment) Protocol B (no target enrichment) 18 Appendix A: Adapter indexes 25 Appendix B: Low multiplexing guidelines 25 GENERAL INFORMATION A Technical support and troubleshooting 26 B Associated products 26 C Product warranty and disclaimer 26 D Trademark and licensing information 26 Research use only 3
4 1. KIT CONTENTS Cap color JetSeq DNA Library Preparation Reagents Volume Step 1: ER buffer 160 μl Step 1: ER enzyme mix 96 μl Step 2: Ligation buffer 48 μl Step 2: Adapter A 80 μl Step 2: Adapter B 80 μl Step 2: Ligase 32 μl Step 3: PCR buffer 80 μl Step 3: Primer mix 80 μl Step 3: DNA polymerase 32 μl Step 4: PCR buffer 80 μl Step 4: Primer 16 μl Step 4: DNA polymerase 32 μl DEPC-treated water 1.8 ml Box 1 Cap color JetSeq DNA Library Preparation Index Set Volume Index 1 Index 2 Index 3 Index 4 Index 5 Index 6 Index 7 Index 8 Index 9 Index 10 Index 11 Index 12 Index 13 Index 14 Index 15 Index 16 Box 2 4
5 JetSeq DNA Library Preparation Kit 2. DESCRIPTION The success of next-generation sequencing is dependent upon the precise and accurate processing of the input DNA. This requires high-quality library preparation of sheared DNA using a coordinated series of standard molecular biology reactions whilst maintaining high-yields during the intermediate purification steps. The JetSeq DNA Library Preparation Kit is designed to generate high-quality next generation sequencing (NGS) libraries suitable for sequencing on Illumina MiniSeq, MiSeq, NextSeq or HiSeq instruments. The kit contains all of the enzymes and buffers necessary for end-repair, A-tailing, ligation and amplification in convenient master mix formulations as well as 16 barcoded adapters that can be used for single or multiplex reads. The JetSeq DNA Library Preparation Kit has been developed and optimized for a variety of DNA sequencing applications, including whole genome sequencing, exome sequencing and targeted sequencing. Low input: 1 ng - 3 μg fragmented DNA Increased speed: sequencing ready library in under 3 hours Improved confidence: simpler protocol improves reproducibility Improved quality: maximum coverage from all sample types Maximum convenience: all-in-one kit Flexible: compatible with target enrichment By combining end-repair and A-tailing in one unique step, the JetSeq DNA Library Preparation Kit is able to reduce total NGS library preparation time and minimize the variability caused by additional handling, as well as the risk of contamination and material loss. In the present manual two different protocols are described: A step-by-step protocol, recommended for all applications, in particular for library preparation protocols that involve enrichment or capture step (Protocol A, section 8.1): quantity and quality of the library is assessed at any step, to ensure optimal results with any enrichment or capture method. A streamlined protocol, recommended for whole genome sequencing and all the sequencing methods which do not need enrichment step (Protocol B, section 8.2). Please read this manual carefully to familiarize yourself with the JetSeq DNA Library Preparation protocol before starting (also available on Research use only 5
6 3. STORAGE When stored under the recommended conditions and handled correctly, full activity of reagents is retained until the expiry date indicated on the outer box label. The kit components should be stored at -20 C. It is recommended that the user avoid repeated freeze-thaw cycles. 4. SAFETY INFORMATION When working with chemicals, always wear suitable personal protective equipment, including lab coat, gloves and safety glasses. For detailed information, please consult the material safety data sheets available on our website at 5. PRODUCT SPECIFICATIONS The JetSeq DNA Library Preparation Kit is designed for Illumina library construction workflows for a wide range of NGS applications, including: targeted sequencing (capture), whole genome sequencing, de novo sequencing, whole exome sequencing and ChIP sequencing. 6
7 JetSeq DNA Library Preparation Kit FRAGMENTED DNA Cap color 5 Adapter A P A A P Adapter B SINGLE TUBE END-REPAIR/A-TAILING T P A A P Cleanup T Cleanup (optional) LIGATION ADAPTER EXTENSION INDEXING Sequence of interest Index Flow cell binding sequence (P5) Flow cell binding sequence (P7) Sequencing primer 1 binding site Indexing primer read 3/ Sequencing primer 2 binding site Fig. 1 Principles of JetSeq DNA Library Preparation Kit Research use only 7
8 SAMPLE I 6. EQUIPMENT AND REAGENTS TO BE SUPPLIED BY THE USER The following additional items are required: Thermal cycler or heat block Equipment for the determination of DNA concentration such as Nanodrop, Qubit, Tapestation, Bioanalyzer or equivalent Equipment for the determination of DNA size distribution such as Tapestation, Bioanalyzer or equivalent Equipment for the purification and size selection of DNA fragments such as JetSeq Clean, AMPure, Dynabeads, SPRI beads or other equivalent column-based (with magnetic device) 10 mm Tris-HCl, ph mm Tris-HCl, ph 8.0, 100 μm EDTA, 50 mm NaCl Vortex Mixer DNase-free plastic ware (0.2 ml tubes, 96-well plates, pipette tips) Molecular grade water Freshly prepared 70% ethanol 7. IMPORTANT NOTES 7.1 Recommended DNA preparation methods The most important prerequisite for any NGS library preparation is high-quality DNA. Sample handling and DNA isolation procedures are therefore critical to the success of the experiment. Residual traces of proteins, salts or other contaminants will degrade the DNA or decrease the efficiency of the enzymatic activities necessary for optimal library preparation. Depending on the sample, we recommend one of the following extraction kits: ISOLATE II Genomic DNA Kit (BIO-52066) for the preparation of genomic DNA from fresh tissues and cells. ISOLATE II Plant DNA Kit (BIO-52069) for isolation of genomic DNA from plants. ISOLATE II PCR and Gel Kit (BIO 52059) for the purification of PCR products and for the isolation of DNA fragments from agarose gels or PCR reactions For more DNA extraction kits, please refer to our ISOLATE II selection tool ( 7.2 Recommendations for DNA fragmentation 8
9 JetSeq DNA Library Preparation Kit DNA can be fragmented using one of the following methods: Mechanical fragmentation (acoustics, sonication, nebulization). Enzymatic fragmentation. To ensure the quality of DNA fragmentation needed for library preparation, only use the recommended parameters given in the manufacturer s instructions. Check the fragmented DNA to ensure a correct size distribution is obtained. 7.3 Recommendations for bead-based clean-up and size selection DNA fragments and libraries can be cleaned up and/or size selected using paramagnetic beads. For these applications, JetSeq Clean beads are recommended ( Alternatively, AMPure XP beads or similar can be used. Conditions and beads volumes listed in the protocol below are valid for both JetSeq Clean and AMPure XP. Beads from any other source should be used following the parameters given by the manufacturer. 7.4 Recommendations for quality control throughout the library preparation Quality of input DNA and DNA libraries can be assessed using Tapestation, Bioanalyzer or equivalent. 8. PROTOCOLS JetSeq DNA Library Preparation Kit can be used following two different protocols, depending on the user requirements. Please refer to Fig.2 for a graphical representation of the two protocols. Protocol A (target enrichment, section 8.1): it is recommended for the vast majority of DNA library preparations and it is particularly suitable for applications that will include a target enrichment step. The quantification analysis recommended at different stages of the protocol will ensure that the right amount of library is used, independently from the efficiency of each step, n particular of the target enrichment. The suggested starting material is fragmented DNA from 1 ng to 3 µg of input. Protocol B (section 8.2): this protocol is a streamlined version of the previous one and is not compatible with target enrichment. It is has been optimized to deliver high-quality, high-yield libraries ready for sequencing, in an efficient and reliable way. The suggested starting material is fragmented DNA from 1 ng to 100 ng of input. Research use only 9
10 For both protocols, all the required enzymes, buffers, adapters, indexes and primers are provided. Fragmented DNA 10 ng - 3 μg Fragmented DNA 10 ng ng End Repair / A-Tailing End Repair / A-Tailing Adapter Ligation Adapter Ligation Cleanup Cleanup Adapter Extension (PCR1) Adapter Extension (PCR1) Cleanup Cleanup Target Enrichment/Capture Indexing (PCR2) Indexing (PCR2) Cleanup Cleanup Sequencing WITH ENRICHMENT Sequencing NO ENRICHMENT Fig. 2 Protocols A and B workflows. Protocol A includes a target enrichment step, protocol B is a standard, direct protocol. 10
11 JetSeq DNA Library Preparation Kit 8.1 Protocol A (target enrichment) End-repair Remove the Step 1 reagents (green cap) and the nuclease free water (blue cap) from storage (-20 C) and allow them to thaw on ice. 1. Prepare reaction mix on ice using the volumes shown below and mix by pipetting up and down. Table 1. End-repair reaction mix Cap color Reagent Quantity Fragmented DNA µg Step 1: ER buffer 10 µl Step 1: ER enzyme mix 6 µl DEPC-treated water up to 50 µl 2. Incubate for 30 min at 20 C then 30 min at 72 C. 3. Transfer the reaction tube on ice (4 C) Adapter Ligation Remove the Step 2 reagents (yellow cap) from storage (-20 C) and allow them to thaw on ice. Preparation of adapter solution Prepare Adapter A and Adapter B Solutions by diluting adapter stocks in a 1 mm Tris-HCl (ph 8.0), 100 μm EDTA, 50 mm NaCl buffer according to Table 2 for both each of the adapters. Research use only 11
12 Table 2. Recommended* adapter dilution factors for varying starting amounts of input DNA Input DNA Amount Dilution Factor 3 1 µg no dilution required µg 1: ng 1: ng 1: ng 1: ng 1:40 * Adapter concentrations calculations are based on DNA fragments of 180 bp. Users are advised to use this table as a guideline to optimize the dilution of Adapters to be used for different fragment sizes. Adapter ligation set-up 1. Using the end-repair reaction from section assemble the following reagents on ice. Mix by pipetting up and down. Table 3. Adapter ligation reaction mix Cap color Reagent Volumes End-repair reaction from section µl Step 2: Ligation buffer 3 µl Step 2: Adapter A solution 5 µl Step 2: Adapter B solution 5 µl Step 2: Ligase 2 µl Total 65 µl 2. Incubate for 15 min at 20 C. 3. Clean up and size select the adapter-ligated library. It is important at this stage to remove unligated adapters and unwanted adapter-dimers. Note: Equipment and reagents are not provided, see section 6 12
13 JetSeq DNA Library Preparation Kit Post-ligation Clean-up This protocol has been optimized using JetSeq Clean beads and AMPure XP beads. Users are advised to optimize the clean-up conditions when working with different beads. All volumes refer to reactions carried out in 0.2 ml tubes or 96-well plates. a) b) c) d) e) f) Allow JetSeq Clean beads to equilibrate at room temperature for at least 30 min. Vortex thoroughly to ensure beads are homogenously suspended. Perform a 1.8x bead-based clean-up by adding 117 µl of homogenous JetSeq Clean beads to each adapter ligated DNA sample. Mix well by pipetting up and down at least 10 times. Incubate at room temperature for 5 min. Put the tube(s)/plate in the magnetic stand to capture the beads until the solution is clear (approximately 3-5 min). Keeping the tube(s)/plate in the magnetic stand, carefully remove and discard the supernatant without disturbing the beads. Continue to keep the tube(s)/plate in the magnetic stand whilst adding 200 µl of 70% freshly prepared ethanol to each tube. IMPORTANT: Always use freshly prepared 70% ethanol. Incubate the tube(s)/plate on the magnetic stand at room temperature for 1 min and remove the ethanol. g) Repeat wash (step e and f). h) i) j) After the second wash, remove all residual ethanol without disturbing the beads. TIP: Use µl tips to aspirate small volumes of residual ethanol. Leave the lids open and dry the beads at room temperature for 3-5 min or until the residual ethanol has completely evaporated. IMPORTANT: Do not over-dry the beads as this will decrease yield. The bead pellet is dry when the appearance of the surface changes from shiny to matt. Remove tube(s)/plate from the magnetic stand. Add 32 µl of 10 mm Tris-HCl ph 8.0 to the bead pellet, mix well by pipetting up and down at least 10 times. Incubate for 3 min at room temperature. Place tube(s)/plate back on magnetic stand for 2-3 min or until the solution is clear. k) Remove 30 µl of the supernatant and transfer to a fresh tube(s)/plate. Discard the beads. l) Assess quality and concentration of the cleaned up adapter-ligated DNA: Confirm the DNA library size distribution and the absence of adapter-dimers on a Bioanalyzer, Tapestation or equivalent. An increase of 58 bp should be measured following the ligation of the adapters. Determine concentration of the purified adapter-ligated DNA using Nanodrop, Qubit or equivalent. m) The purified DNA can be stored at -20 C up to one week. Research use only 13
14 8.1.4 Adapter extension (PCR 1) Remove the Step 3 reagents (orange cap) from storage (-20 C) and allow them to thaw on ice. 1. Assemble the following reaction on ice using the quantities shown below. Mix by pipetting up and down. Table 4. PCR 1 reaction mix Cap color Reagent Volumes Purified adapter-ligated library from ng Step 3: PCR buffer 5 µl Step 3: Primer mix 5 µl Step 3: DNA polymerase 2 µl DEPC-treated water up to 50 µl 2. Run the PCR using the following conditions. Table 5. PCR 1 cycling conditions Temperature ( C) Time Cycles 98 C 3 min 1 98 C 30 s 65 C 30 s See table 6 72 C 1 min 72 C 10 min 1 4 C Hold Table 6. Number of cycles recommended according to the amount of purified adapter-ligated DNA used DNA quantity in PCR1 Number of PCR cycles >20 ng ng ng ng ng 9 <1 ng 10 or more Note: It is not recommended to perform >10 cycles as this will increase the percentage of duplicates. 14
15 JetSeq DNA Library Preparation Kit 3. Optional: Check the quality of the library on a Bioanalyzer, Tapestation or similar equipment. This is to ensure the absence of adapter-dimers. If adapter-dimers are observed it is recommended that a clean-up of the adapter extension (PCR 1) is performed in order to remove these unwanted products Post-PCR 1 clean-up This protocol has been optimized using JetSeq Clean beads and AMPure XP beads. Users are advised to optimize the clean-up conditions when working with different beads. All volumes refer to reactions carried out in 0.2 ml tubes or 96-well plates. a) b) c) d) e) f) Allow JetSeq Clean beads to equilibrate at room temperature for 30 min. Vortex beads thoroughly to ensure homogenous resuspension. Add 90 µl of homogenous JetSeq Clean beads to each library amplification. Mix well by pipetting up and down. Incubate at room temperature for 5 min. Put the tube(s)/plate in the magnetic stand to capture the beads until the solution is clear (approximately 3-5 min). Keeping the tube(s)/plate in the magnetic stand, carefully remove and discard the supernatant without disturbing the beads. Continue to keep the tube(s)/plate in the magnetic stand and add 200 µl of 70% ethanol to each tube. IMPORTANT: Always use freshly prepared 70% ethanol. Incubate the tube(s)/plate on the magnetic stand at room temperature for 1 min and remove the ethanol. g) Repeat wash (step e and f). h) i) j) After the second wash, remove all residual ethanol without disturbing the beads. TIP: Use 10 µl or 20 µl tips to aspirate small volumes of residual ethanol. Leave the lids open and dry the beads at room temperature for 3-5 min or until the residual ethanol has completely evaporated. IMPORTANT: Do not over dry the beads as this will decrease yield. Remove tube(s)/plate from the magnetic stand. Add 32 µl of 10mM Tris-HCl ph 8.0 to the bead pellet, mix well by pipetting up and down. Incubate for 3 min at room temperature. Place on a magnetic stand for 2 3 min or until the solution is clear. k) Remove 30 µl of the supernatant and transfer to a fresh tube(s)/plate. Discard the beads. l) Determine the PCR product concentration using a Nanodrop, Qubit or equivalent. m) The purified, amplified libraries can be stored at 4 C for up to two weeks, or at -20 C for longer periods of time. If target selection or DNA capture is used, please proceed from here, following the protocol provided by the manufacturer. The enriched, purified libraries will be used as DNA input for the adapter completion and indexing step (PCR 2, section 8.1.6). Research use only 15
16 8.1.6 Adapter completion and indexing (PCR 2) Remove the Step 4 reagents (purple cap) from storage (-20 C) and allow them to thaw on ice. 1. Prepare the following reaction mix on ice using the quantities shown below. Mix by pipetting up and down. Table 7. Adapter completion and indexing (PCR 2) reaction mix Cap color Reagent Quantity PCR product from ng ** Index (1-16) 5 µl Step 4: PCR buffer 5 µl Step 4: Primer 1 µl Step 4: DNA polymerase 2 µl DEPC-treated water Up to 50 µl ** if target selection or DNA capture is used the DNA may be at a too low concentration to be measured. In this case we would suggest to use 14 µl of the enriched fraction. 2. Run the PCR with the following cycling conditions: Table 8. Adapter completion and indexing (PCR 2) cycling conditions Temperature ( C) Time Cycles 98 C 3 min 1 98 C 30 s 65 C 30 s 9 ** 72 C 1 min 72 C 10 min 1 4 C Hold ** if target selection or DNA capture is used the DNA may be at a too low concentration to be measured. In this case we would suggest to use 20 cycles. 3. Optional: Check the quality of the library on a Bioanalyzer, Tapestation or similar equipment. This is to ensure the absence of unreacted primers and adapter-dimers. If these are observed it is recommended that a clean-up of the adapter completion and indexing (PCR 2) is performed in order to remove these unwanted products. 16
17 JetSeq DNA Library Preparation Kit Post-PCR 2 clean-up This protocol has been optimized using JetSeq Clean beads and AMPure XP beads. Users are advised to optimize the clean-up conditions when working with different beads. All volumes refer to reactions carried out in 0.2 ml tubes or 96-well plates. a) b) c) d) e) f) Allow JetSeq Clean beads to equilibrate at room temperature for 30 min. Vortex beads thoroughly to ensure homogenous resuspension. Add 90 µl of homogenous JetSeq Clean beads to each library amplification. Mix well by pipetting up and down. Incubate at room temperature for 5 min. Put the tube(s)/plate in the magnetic stand to capture the beads until the solution is clear (approximately 3-5 min). Keeping the tube(s)/plate in the magnetic stand, carefully remove and discard the supernatant without disturbing the beads. Continue to keep the tube(s)/plate in the magnetic stand and add 200 µl of 70% ethanol to each tube. IMPORTANT: Always use freshly prepared 70% ethanol. Incubate the tube(s)/plate on the magnetic stand at room temperature for 1 min and remove the ethanol. g) Repeat wash (step e and f). h) i) j) After the second wash, remove all residual ethanol without disturbing the beads. TIP: Use 10 µl or 20 µl tips to aspirate small volumes of residual ethanol. Leave the lids open and dry the beads at room temperature for 3 5 min or until the residual ethanol has completely evaporated. IMPORTANT: Do not over dry the beads as this will decrease yield. Remove tube(s)/plate from the magnetic stand. Add 32 µl of 10mM Tris-HCl ph 8.0 to the bead pellet, mix well by pipetting up and down. Incubate for 3 min at room temperature. Place on a magnetic stand for 2 3 min or until the solution is clear. k) Remove 30 µl of the supernatant and transfer to a fresh tube(s)/plate. Discard the beads. l) m) Assess quality and concentration of the DNA libraries: Confirm the DNA library size distribution and the absence of unbound primers or adapterdimers on a Bioanalyzer, Tapestation or equivalent. When comparing the products of PCR 1 (adapter extension) and PCR 2 (adapter completion and indexing reaction), an increase of approximatively 70 bp of the DNA size should be observed. Determine the PCR product concentration using Nanodrop, Qubit or equivalent. For accurate measurement we recommend the JetSeq Library Quantification Kit. The purified, amplified libraries can be stored at 4 C for up to two weeks, or at -20 C for longer periods of time. The DNA library is ready for sequencing on MiniSeq, MiSeq, NextSeq and HiSeq platforms and can be pooled if necessary. When loading the library in the sequencing machine we recommend following the manufacturer s instructions. Research use only 17
18 8.2 Protocol B (no target enrichment) Protocol B has been optimized to provide a faster, streamlined version of the previous protocol to be used when the target enrichment step is not required. The intermediate DNA fragments obtained after each step are purified using paramagnetic beads and directly used in the following step without the need of quantifying End-repair Remove the Step 1 reagents (green cap) and the DEPC-treated water (blue cap) from storage (-20 C) and allow them to thaw on ice. 1. Prepare reaction on ice using the volumes shown below and mix by pipetting up and down. Table 9. End-repair reaction Cap color Reagent Volumes Fragmented DNA ng Step 1: ER buffer 10 µl Step 1: ER enzyme mix 6 µl DEPC-treated water Up to 50 µl 2. Incubate for 30 min at 20 C, then 5 min at 72 C. If a thermocycler is used, we recommend setting the heated lid at 85 C 3. Cool down at 4 C or transfer the reaction tubes on ice Adapter ligation Remove the Step 2 reagents (yellow cap) from storage (-20 C) and allow them to thaw on ice. Preparation of adapter solution Prepare Adapter A and Adapter B Solutions by diluting adapter stocks in a 1 mm Tris-HCl (ph 8.0), 100 µm EDTA, 50 mm NaCl buffer according to Table 2 for each of the adapters. 18
19 JetSeq DNA Library Preparation Kit Table 10. Recommended* adapter dilution factors for varying starting amounts of input DNA. Input DNA Amount Dilution Factor ng 1: ng 1: ng 1:40 * Adapter concentrations calculations are based on DNA fragments of 180 bp. Users are advised to use this table as a guideline to optimize the dilution of Adapters to be used for different fragment sizes. Adapter ligation set-up 1. Using the end-repaired reaction from section 8.2.1, prepare an adapter-ligation mix by assembling the following reagents on ice. Mix by pipetting up and down. Table 11. Adapter ligation reaction mix Cap color Reagent Volumes End-repair reaction from section µl Step 2: Ligation buffer 3 µl Step 2: Adapter A solution 5 µl Step 2: Adapter B solution 5 µl Step 2: Ligase 2 µl Total 65 µl 2. Incubate for 15 min at 20 C. 3. Clean up the adapter-ligated library. It is important at this stage to remove unwanted adapter-dimers. Research use only 19
20 8.2.3 Post-ligation clean-up This protocol has been optimized using JetSeq Clean beads and AMPure XP beads. Users are advised to optimize the clean-up conditions when working with different beads. Note: Equipment and reagents are not provided, see section 6. a) b) c) d) e) f) Allow JetSeq Clean beads to equilibrate at room temperature for at least 30 min. Vortex thoroughly to ensure beads are homogenously suspended. For starting DNA input 10 ng: Perform a 1.8x bead-based clean-up by adding 117 µl of homogenous JetSeq Clean beads to each adapter ligated DNA sample. For starting DNA input <10 ng: Perform a 0.8x bead-based clean-up by adding 52 µl of homogenous JetSeq Clean beads to each adapter ligated DNA sample. Mix well by pipetting up and down at least 10 times. Incubate at room temperature for 5 min. Put the tube(s)/plate in the magnetic stand to capture the beads until the solution is clear (approximately 3-5 min). Keeping the tube(s)/plate in the magnetic stand, carefully remove and discard the supernatant without disturbing the beads. Continue to keep the tube(s)/plate in the magnetic stand whilst adding 200 µl of 70% freshly prepared ethanol to each tube. IMPORTANT: Always use freshly prepared 70% ethanol. Incubate the tube(s)/plate on the magnetic stand at room temperature for 1 min and remove the ethanol. g) Repeat wash (step e to f). h) i) j) After the second wash, remove all residual ethanol without disturbing the beads. TIP: Use µl tips to aspirate small volumes of residual ethanol. Leave the lids open and dry the beads at room temperature for 3-5 min or until the residual ethanol has completely evaporated. IMPORTANT: Do not over-dry the beads as this will decrease yield. The bead pellet is dry when the appearance of the surface changes from shiny to matt. Remove tube(s)/plate from the magnetic stand. Add 32 µl of 10 mm Tris-HCl ph 8.0 to the bead pellet, mix well by pipetting up and down at least 10 times. Incubate for 3 min at room temperature. Place tube(s)/plate back on magnetic stand for 2-3 min or until the solution is clear. k) Remove 30 µl of the supernatant and transfer to a fresh tube(s)/plate. Discard the beads. l) The purified DNA can be stored at -20 C up to 1 week. 20
21 JetSeq DNA Library Preparation Kit Adapter extension (PCR 1) 1. Remove the Step 3 reagents (orange cap) from storage (-20 C) and allow them to thaw on ice. 2. Assemble the following reaction on ice using the quantities show below. Mix by pipetting up and down. Table 12. Adapter extension (PCR 1) Cap color Reagent Volumes Purified adapter-ligated library from µl PCR Buffer 5 µl Primer Mix 5 µl DNA Polymerase 2 µl DEPC-treated water 8 µl Total 50 µl 1. Run the PCR using the following conditions Table 13. PCR 1 cycling Temperature Time Cycles 98 C 3 min 1 98 C 30 s 65 C 30 s See table 6 72 C 1 min 72 C 10 min 1 4 C Hold Table 14. Number of cycles recommended according to the amount of DNA used DNA input quantity Recommended number of PCR cycles ng 3 10 ng 4 1 ng 6 2. It is recommended that a clean-up of the adapter extension (PCR 1) is performed in order to remove these unwanted products. This step is crucial to remove unwanted adapters and adapter-dimers from the library. Research use only 21
22 8.2.5 Post adapter extension clean-up This protocol has been optimized using JetSeq Clean beads and AMPure XP beads. Users are advised to optimize the clean-up conditions when working with different beads. Note: Equipment and reagents are not provided, see section 6. a) b) c) d) e) f) Allow JetSeq Clean beads to equilibrate at room temperature for at least 30 min. Vortex thoroughly to ensure beads are homogenously suspended. For starting DNA input 10 ng: Perform a 1.8x bead-based clean-up by adding 90 µl of homogenous JetSeq Clean beads to each adapter ligated DNA sample. For starting DNA input <10 ng: Perform a 1x bead-based clean-up by adding 50 µl of homogenous JetSeq Clean beads to each adapter ligated DNA sample. Mix well by pipetting up and down at least 10 times. Incubate at room temperature for 5 min. Put the tube(s)/plate in the magnetic stand to capture the beads until the solution is clear (approximately 3-5 min). Keeping the tube(s)/plate in the magnetic stand, carefully remove and discard the supernatant without disturbing the beads. Continue to keep the tube(s)/plate in the magnetic stand whilst adding 200 µl of 70% freshly prepared ethanol to each tube. IMPORTANT: Always use freshly prepared 70% ethanol. Incubate the tube(s)/plate on the magnetic stand at room temperature for 1 min and remove the ethanol. g) Repeat wash (step e and f). h) i) j) After the second wash, remove all residual ethanol without disturbing the beads. TIP: Use µl tips to aspirate small volumes of residual ethanol. Leave the lids open and dry the beads at room temperature for 3-5 min or until the residual ethanol has completely evaporated. IMPORTANT: Do not over-dry the beads as this will decrease yield. The bead pellet is dry when the appearance of the surface changes from shiny to matt. Remove tube(s)/plate from the magnetic stand. Add 32 µl of 10 mm Tris-HCl ph 8.0 to the bead pellet, mix well by pipetting up and down at least 10 times. Incubate for 3 min at room temperature. Place tube(s)/plate back on magnetic stand for 2-3 min or until the solution is clear. k) Remove 30 µl of the supernatant and transfer to a fresh tube(s)/plate. Discard the beads. l) The purified, amplified libraries can be stored at 4 C for up to two weeks, or at 20 C for longer periods of time. 22
23 8.2.6 Adapter completion and indexing (PCR 2) 1. Remove the Step 4 reagents (purple cap) from storage (-20 C) and allow them to thaw on ice. 2. Prepare the following reaction mix on ice using the quantities shown below. Mix by pipetting up and down. Table 15. Adapter completion and indexing (PCR 2) Cap Color Reagent Volumes PCR product from µl Index (1-16) 5 µl PCR Buffer 5 µl Step 4: Primer 1 µl DNA Polymerase 2 µl DEPC-treated water 7 µl Run the PCR with the following cycling conditions: Table 16. Adapter completion and indexing (PCR 2) cycling conditions Temperature Time Cycles 98 C 3 min 1 98 C 30 s 65 C 30 s See table 9 72 C 1 min 72 C 10 min 1 4 C Hold Table 17. Number of cycles recommended according to the amount of DNA used DNA input quantity Recommended number of PCR cycles ng ng ng It is crucial to perform post adapter completion and indexing (PCR 2) clean-up in order to remove unbound primers or unwanted adapter-dimers from the library. 23
24 8.2.6 Post adapter completion and indexing clean-up This protocol has been optimized using JetSeq Clean beads and AMPure XP beads. Users are advised to optimize the clean-up conditions when working with different beads. Note: Equipment and reagents are not provided, see section 6. a) b) c) d) e) f) Allow JetSeq Clean beads to equilibrate at room temperature for at least 30 min. Vortex thoroughly to ensure beads are homogenously suspended. For starting DNA input 10 ng: Perform a 1.8x bead-based clean-up by adding 90 µl of homogenous JetSeq Clean beads to each adapter ligated DNA sample. For starting DNA input <10 ng: Perform a 1x bead-based clean-up by adding 50 µl of homogenous JetSeq Clean beads to each adapter ligated DNA sample. Mix well by pipetting up and down at least 10 times. Incubate at room temperature for 5 min. Put the tube(s)/plate in the magnetic stand to capture the beads until the solution is clear (approximately 3-5 min). Keeping the tube(s)/plate in the magnetic stand, carefully remove and discard the supernatant without disturbing the beads. Continue to keep the tube(s)/plate in the magnetic stand whilst adding 200 µl of 70% freshly prepared ethanol to each tube. IMPORTANT: Always use freshly prepared 70% ethanol. Incubate the tube(s)/plate on the magnetic stand at room temperature for 1 min and remove the ethanol. g) Repeat wash (step e to f). h) i) j) After the second wash, remove all residual ethanol without disturbing the beads. TIP: Use µl tips to aspirate small volumes of residual ethanol. Leave the lids open and dry the beads at room temperature for 3-5 min or until the residual ethanol has completely evaporated. IMPORTANT: Do not over-dry the beads as this will decrease yield. The bead pellet is dry when the appearance of the surface changes from shiny to matt. Remove tube(s)/plate from the magnetic stand. Add 32 µl of 10 mm Tris-HCl ph 8.0 to the bead pellet, mix well by pipetting up and down at least 10 times. Incubate for 3 min at room temperature. Place tube(s)/plate back on magnetic stand for 2-3 min or until the solution is clear. k) Remove 30 µl of the supernatant and transfer to a fresh tube(s)/plate. Discard the beads. l) m) Assess quality and concentration of the DNA libraries Confirm the DNA library size distribution and the absence of unbound primers or adapterdimers on a Bioanalyzer, Tapestation or equivalent. When comparing the final libraries with the starting material, an increase of approximatively 120 bp of the DNA size should be observed. Determine the PCR product concentration using Nanodrop, Qubit or equivalent. For accurate measurement, we recommend the JetSeq Library Quantification Kit. The purified, amplified libraries can be stored at 4 C for up to two weeks, or at -20 C for longer periods of time. The DNA library is ready for sequencing on MiniSeq, MiSeq, NextSeq and HiSeq platforms and can be pooled if necessary. When loading the library in the sequencing machine we recommend following the manufacturer s instructions. 24
25 JetSeq DNA Library Preparation Kit Appendix A: Adapter indexes The nucleotide sequences for the 16 indexes provided are detailed in the table below. Index Number Sequence 1 AACGTGAT 2 AAACATCG 3 AGTGGTCA 4 ACCACTGT 5 GATAGACA 6 GTGTTCTA 7 TGGAACAA 8 TGGTGGTA 9 ACATTGGC 10 CAGATCTG 11 CATCAAGT 12 AGTACAAG 13 AGATCGCA 14 GACTAGTA 15 GGTGCGAA 16 TGAAGAGA Appendix B: Low multiplexing guidelines Illumina platform such as MiSeq and HiSeq use a red laser to sequence A/C and a green laser to sequence G/T. To ensure accurate registration of the index read, both a red and green signal must be present at each cycle. It is also important to maintain color balance where possible. If pooling less than eight samples in the final sequencing pool we suggest using the following index combinations Number of Index samples in pool 1 Any index 2 2 & 6 Option A: 4, 6 & 7 3 Option B: 1, 11 & 16 Option A: 2, 6, 10 & 14 4 Option B: 9, 12, 15 & 16 Option A: 1, 2, 4, 6, 7 & 8 6 Option B: 2, 8, 9, 12, 15 & 16 Option A: Option B: 9-16 Research use only 25
26 A B TECHNICAL SUPPORT AND TROUBLESHOOTING For technical assistance or more information on this product, please us at tech@bioline.com ASSOCIATED PRODUCTS Product Size Cat. # ISOLATE II Genomic DNA Kit 50 prep BIO ISOLATE II FFPE RNA/DNA Kit 50 prep BIO ISOLATE II Plant DNA Kit 50 prep BIO JetSeq Library Quantification Hi-ROX Kit 500 Reactions BIO JetSeq Library Quantification Lo-ROX Kit 500 Reactions BIO JetSeq Clean 50 ml BIO C D PRODUCT WARRANTY AND DISCLAIMER Bioline warrants that its products will conform to the standards stated in its product specification sheets in effect at the time of shipment. Bioline will replace any product that does not conform to the specifications free of charge. This warranty limits Bioline s liability to only the replacement of the product. TRADEMARK AND LICENSING INFORMATION JetSeq was developed jointly by OGT and Bioline. JetSeq (Bioline Reagents Ltd), HiSeq, MiSeq, NextSeq (Illumina Inc.); Qubit (ThermoFisher Scientific); Dynabeads (Dynal Inc.); AMPure (Beckman Coulter Inc.) 26
27 Ordering Information Product Size Cat. # JetSeq DNA Library Preparation Kit 16 Reactions BIO PM0118V1.5 USA info.us@bioline.com Order Toll Free: France info.fr@bioline.com Tel: +33 (0) United Kingdom info.uk@bioline.com Tel: +44 (0) Australia info.au@bioline.com Tel: +61 (0) Germany info.de@bioline.com Tel: +49 (0) Singapore info.sg@bioline.com Toll Free: 1800 BIOLINE ( ) bioline.com/jetseq To find a Bioline distributor in your country, visit bioline.com/distributors
JetSeq Clean. Product Manual
 JetSeq Clean Product Manual 2 Product Manual bioline.com/jetseq JetSeq Clean JetSeq Clean TABLE OF CONTENTS 1 Kit contents 04 2 Description 05 3 Equipment and reagents to be supplied by user 06 4 Storage
JetSeq Clean Product Manual 2 Product Manual bioline.com/jetseq JetSeq Clean JetSeq Clean TABLE OF CONTENTS 1 Kit contents 04 2 Description 05 3 Equipment and reagents to be supplied by user 06 4 Storage
Additional reagents and materials that are not supplied
 sparq PureMag Beads Cat. No. 95196-005 Size: 5 ml Store at 2 C to 8 C 95196-060 60 ml 95196-450 450 ml Description sparq PureMag Beads uses reversible nucleic acid-binding properties of magnetic beads
sparq PureMag Beads Cat. No. 95196-005 Size: 5 ml Store at 2 C to 8 C 95196-060 60 ml 95196-450 450 ml Description sparq PureMag Beads uses reversible nucleic acid-binding properties of magnetic beads
NR601. VAHTS TM mrna-seq V2 Library Prep Kit for Illumina
 NR601 VAHTS TM mrna-seq V2 Library Prep Kit for Illumina v Vazyme Biotech Co., Ltd Website: www.vazyme.com Order: global@vazyme.com Support: support@vazyme.com Service: service@vazyme.com SYSTEMS www.vazyme.com
NR601 VAHTS TM mrna-seq V2 Library Prep Kit for Illumina v Vazyme Biotech Co., Ltd Website: www.vazyme.com Order: global@vazyme.com Support: support@vazyme.com Service: service@vazyme.com SYSTEMS www.vazyme.com
USER GUIDE For Illumina Platform
 USER GUIDE For Illumina Platform Copyright Nimagen B.V. P.O. Box 91 6500 AB Nijmegen The Netherlands Tel. +31 (0)24 820 0241 Fax. +31 (0)24 358 0259 info@nimagen.com VAT#: NL850011243B01 Rabobank Nijmegen:
USER GUIDE For Illumina Platform Copyright Nimagen B.V. P.O. Box 91 6500 AB Nijmegen The Netherlands Tel. +31 (0)24 820 0241 Fax. +31 (0)24 358 0259 info@nimagen.com VAT#: NL850011243B01 Rabobank Nijmegen:
VAHTS Stranded mrna-seq Library Prep Kit for Illumina
 Instruction Manual VAHTS Stranded mrna-seq Library Prep Kit for Illumina Vazyme Cat #NR602 Vazyme Biotech Co., Ltd Web: www.vazyme.com Tel: 400-600-9335 Sales: Sales@vazyme.com Support: Support@ vazyme.com
Instruction Manual VAHTS Stranded mrna-seq Library Prep Kit for Illumina Vazyme Cat #NR602 Vazyme Biotech Co., Ltd Web: www.vazyme.com Tel: 400-600-9335 Sales: Sales@vazyme.com Support: Support@ vazyme.com
Illumina TruSeq Stranded mrna (LT) Protocol 1
 Illumina TruSeq Stranded mrna (LT) Protocol 1 Performed using the TruSeq Stranded mrna Sample Preparation Kit (A cat#fc-122-2101, B cat#fc122-2102) Purify and Fragment mrna NOTE: Use 500ng of Total RNA
Illumina TruSeq Stranded mrna (LT) Protocol 1 Performed using the TruSeq Stranded mrna Sample Preparation Kit (A cat#fc-122-2101, B cat#fc122-2102) Purify and Fragment mrna NOTE: Use 500ng of Total RNA
illumina TruSeq RNA Sample Prep. (LT) Protocol 1
 illumina TruSeq RNA Sample Prep. (LT) Protocol 1 Performed using the TruSeq RNA Sample Preparation Kit (A cat#fc-122-1001, B cat#fc122-1002) Purify and Fragment mrna NOTE: Use 3ug of Total RNA to initiate
illumina TruSeq RNA Sample Prep. (LT) Protocol 1 Performed using the TruSeq RNA Sample Preparation Kit (A cat#fc-122-1001, B cat#fc122-1002) Purify and Fragment mrna NOTE: Use 3ug of Total RNA to initiate
NEBNext DNA Library Prep Master Mix Set for Illumina
 LIBRARY PREPARATION NEBNext DNA Library Prep Master Mix Set for Illumina Instruction Manual NEB #E6040S/L 12/60 reactions Version 8.0 9/18 be INSPIRED drive DISCOVERY stay GENUINE This product is intended
LIBRARY PREPARATION NEBNext DNA Library Prep Master Mix Set for Illumina Instruction Manual NEB #E6040S/L 12/60 reactions Version 8.0 9/18 be INSPIRED drive DISCOVERY stay GENUINE This product is intended
Select-a-Size DNA Clean & Concentrator MagBead Kit Catalog No. D4084 & D4085
 INSTRUCTION MANUAL Select-a-Size DNA Clean & Concentrator MagBead Kit Catalog No. D4084 & D4085 Highlights Tunable: Size selection can be tuned from 100 bp to 1000 bp with left, right, or double size selection
INSTRUCTION MANUAL Select-a-Size DNA Clean & Concentrator MagBead Kit Catalog No. D4084 & D4085 Highlights Tunable: Size selection can be tuned from 100 bp to 1000 bp with left, right, or double size selection
NxSeq UltraLow DNA Library Kit, 12 Reactions
 NxSeq UltraLow DNA Library Kit, 12 Reactions Illumina-compatible FOR RESEARCH USE ONLY. NOT FOR HUMAN OR DIAGNOSTIC USE Lucigen Corporation 2905 Parmenter St, Middleton, WI 53562 USA Toll Free: (888) 575-9695
NxSeq UltraLow DNA Library Kit, 12 Reactions Illumina-compatible FOR RESEARCH USE ONLY. NOT FOR HUMAN OR DIAGNOSTIC USE Lucigen Corporation 2905 Parmenter St, Middleton, WI 53562 USA Toll Free: (888) 575-9695
Procedure & Checklist - 1 kb Template Preparation and Sequencing
 Procedure & Checklist - 1 kb Template Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit. Fragment and Concentrate DNA Important: The distribution
Procedure & Checklist - 1 kb Template Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit. Fragment and Concentrate DNA Important: The distribution
Procedure & Checklist - 2 kb Template Preparation and Sequencing
 Procedure & Checklist - 2 kb Template Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio DNA Template Prep Kit (verify you have the correct kit for your insert
Procedure & Checklist - 2 kb Template Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio DNA Template Prep Kit (verify you have the correct kit for your insert
Procedure & Checklist bp Amplicon Library Preparation and Sequencing
 Procedure & Checklist - 250 bp Amplicon Library Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit. This procedure is optimized for SMRTbell
Procedure & Checklist - 250 bp Amplicon Library Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit. This procedure is optimized for SMRTbell
Stranded mrna-seq Lib Prep Kit for Illumina
 Stranded mrna-seq Lib Prep Kit for Illumina RK20301 (10ng-1ug Input Total RNA) (Illumina Compatible) C U G A www.abclonal.com version: N12G13v1.0 Contents 1.Introduction 01 2.Components 02 3.Additional
Stranded mrna-seq Lib Prep Kit for Illumina RK20301 (10ng-1ug Input Total RNA) (Illumina Compatible) C U G A www.abclonal.com version: N12G13v1.0 Contents 1.Introduction 01 2.Components 02 3.Additional
Procedure & Checklist bp Template Preparation and Sequencing
 Procedure & Checklist - 500 bp Template Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit. This procedure is optimized for SMRTbell template
Procedure & Checklist - 500 bp Template Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit. This procedure is optimized for SMRTbell template
Procedure & Checklist - 10 kb Template Preparation and Sequencing
 Procedure & Checklist - 10 kb Template Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit. This procedure can be used to prepare 10 kb libraries
Procedure & Checklist - 10 kb Template Preparation and Sequencing Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit. This procedure can be used to prepare 10 kb libraries
Procedure & Checklist Preparing SMRTbell Libraries using PacBio Barcoded Adapters for Multiplex SMRT Sequencing
 Procedure & Checklist Preparing SMRTbell Libraries using PacBio Barcoded Adapters for Multiplex SMRT Sequencing Before You Begin This document describes a procedure for multiplexing 5 Mb microbial genomes
Procedure & Checklist Preparing SMRTbell Libraries using PacBio Barcoded Adapters for Multiplex SMRT Sequencing Before You Begin This document describes a procedure for multiplexing 5 Mb microbial genomes
Procedure & Checklist - Preparing SMRTbell Libraries using PacBio Barcoded Adapters for Multiplex SMRT Sequencing
 Procedure & Checklist - Preparing SMRTbell Libraries using PacBio Barcoded Adapters for Multiplex SMRT Sequencing Before You Begin This document describes methods for generating SMRTbell libraries using
Procedure & Checklist - Preparing SMRTbell Libraries using PacBio Barcoded Adapters for Multiplex SMRT Sequencing Before You Begin This document describes methods for generating SMRTbell libraries using
MEDIP-SEQUENCING PROTOCOL
 MEDIP-SEQUENCING PROTOCOL MAGMEDIP KIT Cat. No. C02010020 Table 1 The GenDNA module provides you with an excess of buffer for the preparation of DNA. Sufficient buffer is given for the preparation of several
MEDIP-SEQUENCING PROTOCOL MAGMEDIP KIT Cat. No. C02010020 Table 1 The GenDNA module provides you with an excess of buffer for the preparation of DNA. Sufficient buffer is given for the preparation of several
Procedure & Checklist - Iso-Seq Template Preparation for Sequel Systems
 Procedure & Checklist - Iso-Seq Template Preparation for Sequel Systems Before You Begin The long read lengths of the PacBio System are well-suited for characterizing full-length transcripts produced from
Procedure & Checklist - Iso-Seq Template Preparation for Sequel Systems Before You Begin The long read lengths of the PacBio System are well-suited for characterizing full-length transcripts produced from
Procedure & Checklist Multiplex Genomic DNA Target Capture Using IDT xgen Lockdown Probes
 Procedure & Checklist Multiplex Genomic DNA Target Capture Using IDT xgen Lockdown Probes Before You Begin This procedure describes capture and enrichment of regions of interest by using IDT xgen Lockdown
Procedure & Checklist Multiplex Genomic DNA Target Capture Using IDT xgen Lockdown Probes Before You Begin This procedure describes capture and enrichment of regions of interest by using IDT xgen Lockdown
10 kb to 20 kb Template Preparation and Sequencing with Low-Input DNA
 Please note: the shared protocols described herein may not have been validated by Pacific Biosciences and are provided as-is and without any warranty. Use of these protocols is offered to those customers
Please note: the shared protocols described herein may not have been validated by Pacific Biosciences and are provided as-is and without any warranty. Use of these protocols is offered to those customers
Procedure & Checklist - 10 kb Template Preparation and Sequencing (with Low-Input DNA)
 Procedure & Checklist - 10 kb Template Preparation and Sequencing (with Low-Input DNA) Before You Begin To perform this procedure, you must have the PacBio : Template Prep Kit DNA/Polymerase Binding Kit
Procedure & Checklist - 10 kb Template Preparation and Sequencing (with Low-Input DNA) Before You Begin To perform this procedure, you must have the PacBio : Template Prep Kit DNA/Polymerase Binding Kit
Procedure and Checklist - 20 kb Template Preparation Using BluePippin Size-Selection System
 Procedure and Checklist - 20 kb Template Preparation Using BluePippin Size-Selection System Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit and have reviewed the
Procedure and Checklist - 20 kb Template Preparation Using BluePippin Size-Selection System Before You Begin To perform this procedure, you must have the PacBio Template Prep Kit and have reviewed the
xgen hybridization capture of DNA libraries
 xgen hybridization capture of DNA libraries For NGS target enrichment Uses IDT s: xgen Hybridization and Wash Kit xgen Universal Blockers TS Mix, 10 bp TS Mix, or NXT Mix xgen Lockdown Panels and Probes
xgen hybridization capture of DNA libraries For NGS target enrichment Uses IDT s: xgen Hybridization and Wash Kit xgen Universal Blockers TS Mix, 10 bp TS Mix, or NXT Mix xgen Lockdown Panels and Probes
MinION PROTOCOL. Adapted from Janneke Wit by Robyn Tanny May Company Kit/Item Catalog Number
 MinION PROTOCOL Adapted from Janneke Wit by Robyn Tanny May 2016 Company Kit/Item Catalog Number Fisher Eppendorf DNA/RNA LoBind Tubes 13-698-791 Fisher Covaris g-tube NC0380758 NEB NEBNext FFPE Repair
MinION PROTOCOL Adapted from Janneke Wit by Robyn Tanny May 2016 Company Kit/Item Catalog Number Fisher Eppendorf DNA/RNA LoBind Tubes 13-698-791 Fisher Covaris g-tube NC0380758 NEB NEBNext FFPE Repair
We want to thank and acknowledge the authors for sharing this protocol and their contributions to the field.
 We adopted the protocol described in the Extended Experimental Procedures section I.a.1 of the 2014 Cell paper by Rao and Huntley et. al: A 3D Map of the Human Genome at Kilobase Resolution Reveals Principles
We adopted the protocol described in the Extended Experimental Procedures section I.a.1 of the 2014 Cell paper by Rao and Huntley et. al: A 3D Map of the Human Genome at Kilobase Resolution Reveals Principles
Procedure & Checklist - 20 kb Template Preparation Using BluePippin Size-Selection System
 Procedure & Checklist - 20 kb Template Preparation Using BluePippin Size-Selection System Before You Begin To perform this procedure, you must have the PacBio DNA Template Prep Kit 2.0 (3 kb to 10 kb)
Procedure & Checklist - 20 kb Template Preparation Using BluePippin Size-Selection System Before You Begin To perform this procedure, you must have the PacBio DNA Template Prep Kit 2.0 (3 kb to 10 kb)
Procedure & Checklist >20 kb Template Preparation Using BluePippin Size-Selection System (15-20 kb Cutoff) for Sequel Systems
 Procedure & Checklist >20 kb Template Preparation Using BluePippin Size-Selection System (15-20 kb Cutoff) for Sequel Systems Before You Begin To perform this procedure, you must have the PacBio Template
Procedure & Checklist >20 kb Template Preparation Using BluePippin Size-Selection System (15-20 kb Cutoff) for Sequel Systems Before You Begin To perform this procedure, you must have the PacBio Template
Procedure & Checklist - cdna Capture Using SeqCap EZ Libraries
 Procedure & Checklist - cdna Capture Using SeqCap EZ Libraries Before You Begin This document describes the process for capturing cdna prepared with the SMARTer PCR cdna Synthesis Kit (Clontech) and pulled-down
Procedure & Checklist - cdna Capture Using SeqCap EZ Libraries Before You Begin This document describes the process for capturing cdna prepared with the SMARTer PCR cdna Synthesis Kit (Clontech) and pulled-down
Target Sequence Capture Using Roche NimbleGen SeqCap EZ Library
 Please note: the shared protocols described herein may not have been validated by Pacific Biosciences and are provided as-is and without any warranty. Use of these protocols is offered to those customers
Please note: the shared protocols described herein may not have been validated by Pacific Biosciences and are provided as-is and without any warranty. Use of these protocols is offered to those customers
DNA Size Selection Magnetic Beads
 DNA Size Selection Magnetic Beads Catalog #: 801-117 User Manual Last revised July 30 th, 2018 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
DNA Size Selection Magnetic Beads Catalog #: 801-117 User Manual Last revised July 30 th, 2018 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
NGS clean-up and size selection
 NGS clean-up and size selection User manual NucleoMag NGS Clean-up and Size Select May 2014 / Rev. 01 NGS clean-up and size selection Table of contents 1 Components 4 1.1 Kit contents 4 1.2 Equipment and
NGS clean-up and size selection User manual NucleoMag NGS Clean-up and Size Select May 2014 / Rev. 01 NGS clean-up and size selection Table of contents 1 Components 4 1.1 Kit contents 4 1.2 Equipment and
Project: RADseqReady Plate # Library # Name: Date: Section 1: DNA Standardization
 BestRAD Library Preparation Based on protocol of Ali et al. 2015 (10.1534/genetics.115.183665) Adapted by Linda Rutledge many times but this version was done on August 16, 2016 Section 1: DNA Standardization
BestRAD Library Preparation Based on protocol of Ali et al. 2015 (10.1534/genetics.115.183665) Adapted by Linda Rutledge many times but this version was done on August 16, 2016 Section 1: DNA Standardization
Procedure & Checklist cdna Capture Using IDT xgen Lockdown Probes
 Procedure & Checklist cdna Capture Using IDT xgen Lockdown Probes Before You Begin This document describes the process for capturing cdna prepared with the SMARTer PCR cdna Synthesis Kit (Clontech) and
Procedure & Checklist cdna Capture Using IDT xgen Lockdown Probes Before You Begin This document describes the process for capturing cdna prepared with the SMARTer PCR cdna Synthesis Kit (Clontech) and
CleanPlex UMI NGS Panel
 CleanPlex UMI NGS Panel User Guide This user guide is for the following products: CleanPlex UMI Lung Cancer Panel CleanPlex UMI Custom NGS Panel Get the latest user guide at: www.paragongenomics.com/product_documents/
CleanPlex UMI NGS Panel User Guide This user guide is for the following products: CleanPlex UMI Lung Cancer Panel CleanPlex UMI Custom NGS Panel Get the latest user guide at: www.paragongenomics.com/product_documents/
Procedure & Checklist Preparing Multiplexed Microbial SMRTbell Libraries for the PacBio Sequel System
 Procedure & Checklist Preparing Multiplexed Microbial SMRTbell Libraries for the PacBio Sequel System Before You Begin This procedure is for preparing multiplexed SMRTbell libraries for sequencing on the
Procedure & Checklist Preparing Multiplexed Microbial SMRTbell Libraries for the PacBio Sequel System Before You Begin This procedure is for preparing multiplexed SMRTbell libraries for sequencing on the
Procedure & Checklist - Preparing >15 kb Libraries Using SMRTbell Express Template Preparation Kit
 Procedure & Checklist - Preparing >15 kb Libraries Using SMRTbell Express Template Preparation Kit This document provides recommendations for preparing >15 kb size-selected SMRTbell libraries from 3-5
Procedure & Checklist - Preparing >15 kb Libraries Using SMRTbell Express Template Preparation Kit This document provides recommendations for preparing >15 kb size-selected SMRTbell libraries from 3-5
Swift Hybridization Capture Kits
 Protocol Swift Hybridization Capture Kits For NGS Target Enrichment Uses: Swift Exome Hyb Panel, Cat. No. 83216 Swift Pan-Cancer Hyb Panel, Cat. No. 83316 Swift Inherited Diseases Hyb Panel, Cat. No. 83416
Protocol Swift Hybridization Capture Kits For NGS Target Enrichment Uses: Swift Exome Hyb Panel, Cat. No. 83216 Swift Pan-Cancer Hyb Panel, Cat. No. 83316 Swift Inherited Diseases Hyb Panel, Cat. No. 83416
Procedure & Checklist - Greater Than 10 kb Template Preparation Using AMPure PB Beads
 Procedure & Checklist - Greater Than 10 kb Template Preparation Using AMPure PB Beads Before You Begin This procedure can be used to prepare greater than 10 kb libraries from 5 μg of sheared and concentrated
Procedure & Checklist - Greater Than 10 kb Template Preparation Using AMPure PB Beads Before You Begin This procedure can be used to prepare greater than 10 kb libraries from 5 μg of sheared and concentrated
DNA extraction Protocol for Agencourt Genfind v2 Blood and Serum Genomic DNA Isolation Kit
 DNA extraction Protocol for Agencourt Genfind v2 Blood and Serum Genomic DNA Isolation Kit Introduction The Agencourt Genfind v2 Blood & Serum DNA Isolation Kit utilizes Agencourt s patented SPRI paramagnetic
DNA extraction Protocol for Agencourt Genfind v2 Blood and Serum Genomic DNA Isolation Kit Introduction The Agencourt Genfind v2 Blood & Serum DNA Isolation Kit utilizes Agencourt s patented SPRI paramagnetic
Procedure & Checklist - Iso-Seq Template Preparation for Sequel Systems
 Procedure & Checklist - Iso-Seq Template Preparation for Sequel Systems Before You Begin The Sequel System generates long reads that are well-suited for characterizing full-length transcripts produced
Procedure & Checklist - Iso-Seq Template Preparation for Sequel Systems Before You Begin The Sequel System generates long reads that are well-suited for characterizing full-length transcripts produced
ChIP Protocol for fresh or frozen cross linked cells
 Prior to starting your ChIPs and Shearing Turn on sonifiers and cooling system allow system to reach -2 C before shearing Cool bench top centrifuge to 4 C Prepare all of your buffers with protease inhibitors
Prior to starting your ChIPs and Shearing Turn on sonifiers and cooling system allow system to reach -2 C before shearing Cool bench top centrifuge to 4 C Prepare all of your buffers with protease inhibitors
Alon s SCN ChIP Protocol
 Prior to starting your ChIPs and Shearing 1. Turn on sonifiers and cooling system allow system to reach -1 C before shearing 2. Cool bench top centrifuge to 4 C 3. Prepare all of your buffers with protease
Prior to starting your ChIPs and Shearing 1. Turn on sonifiers and cooling system allow system to reach -1 C before shearing 2. Cool bench top centrifuge to 4 C 3. Prepare all of your buffers with protease
AGENCOURT GENFIND Blood & Serum Genomic DNA Isolation Kit
 Blood & Serum Genomic DNA Isolation Kit Page 1 of 9 Please refer to http://www.agencourt.com/technical for updated protocols and refer to MSDS instructions when handling or shipping any chemical hazards.
Blood & Serum Genomic DNA Isolation Kit Page 1 of 9 Please refer to http://www.agencourt.com/technical for updated protocols and refer to MSDS instructions when handling or shipping any chemical hazards.
mi-mag mrna Isolation Kit
 mi-mag mrna Isolation Kit Cat. No [50 Reactions] This kit is for research purposes only. Not for use in diagnostic procedures. For in vitro use only. Introduction This kit contains enough materials for
mi-mag mrna Isolation Kit Cat. No [50 Reactions] This kit is for research purposes only. Not for use in diagnostic procedures. For in vitro use only. Introduction This kit contains enough materials for
Evaluation of Omega Mag-Bind TotalPure NGS Beads for DNA Size Selection
 Evaluation of Omega Mag-Bind TotalPure NGS Beads for Size Selection By Maggie Weitzman, M.Sc. (University of Oregon / GC3F) Disclaimer: Neither Maggie Weitzman, the University of Oregon, nor the Genomics
Evaluation of Omega Mag-Bind TotalPure NGS Beads for Size Selection By Maggie Weitzman, M.Sc. (University of Oregon / GC3F) Disclaimer: Neither Maggie Weitzman, the University of Oregon, nor the Genomics
Required Materials. Page 1 PN Version 06 (February 2018)
 Procedure & Checklist - Preparing >30 kb SMRTbell Libraries Using Megaruptor Shearing and BluePippin Size-Selection for PacBio RS II and Sequel Systems This document provides recommendations for preparing
Procedure & Checklist - Preparing >30 kb SMRTbell Libraries Using Megaruptor Shearing and BluePippin Size-Selection for PacBio RS II and Sequel Systems This document provides recommendations for preparing
USER GUIDE. Ovation PART NO. 0344, 0344NB. Ultralow System V2
 USER GUIDE Ovation PART NO. 0344, 0344NB Ultralow System V2 Patents, Licensing and Trademarks 2014 2017 NuGEN Technologies, Inc. All rights reserved. The Encore, Ovation and Applause families of products
USER GUIDE Ovation PART NO. 0344, 0344NB Ultralow System V2 Patents, Licensing and Trademarks 2014 2017 NuGEN Technologies, Inc. All rights reserved. The Encore, Ovation and Applause families of products
EPIGENTEK. EpiQuik Circulating Cell-Free DNA Isolation Kit. Base Catalog # P-1064 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE
 EpiQuik Circulating Cell-Free DNA Isolation Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiQuik Circulating Cell-Free DNA Isolation Kit utilizes magnetic beads based sizefractionation
EpiQuik Circulating Cell-Free DNA Isolation Kit Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiQuik Circulating Cell-Free DNA Isolation Kit utilizes magnetic beads based sizefractionation
Mag-Bind cfdna Kit. M preps M preps M Preps
 Mag-Bind cfdna Kit M3298-00 5 preps M3298-01 50 preps M3298-02 200 Preps March 2018 Mag-Bind cfdna Kit Table of Contents Introduction and Overview...2 Kit Contents/Storage and Stability...3 Preparing Reagents...4
Mag-Bind cfdna Kit M3298-00 5 preps M3298-01 50 preps M3298-02 200 Preps March 2018 Mag-Bind cfdna Kit Table of Contents Introduction and Overview...2 Kit Contents/Storage and Stability...3 Preparing Reagents...4
RayBio mrna Magnetic Beads Kit
 RayBio mrna Magnetic Beads Kit Catalog #: 801-116 User Manual Last revised March 9 th, 2017 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
RayBio mrna Magnetic Beads Kit Catalog #: 801-116 User Manual Last revised March 9 th, 2017 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross,
Formaldehyde Cross-linking of Chromatin from Drosophila
 2 Formaldehyde Cross-linking of Chromatin from Drosophila Protocol from modencode IGSB University of Chicago originally written by Alex Crofts and Sasha Ostapenko and updated by Matt Kirkey. 1. Set centrifuge
2 Formaldehyde Cross-linking of Chromatin from Drosophila Protocol from modencode IGSB University of Chicago originally written by Alex Crofts and Sasha Ostapenko and updated by Matt Kirkey. 1. Set centrifuge
PARP1-Trap_A for Immunoprecipitation of PARP1- Fusion Proteins from cell extract
 PARP1-Trap_A for Immunoprecipitation of PARP1- Fusion Proteins from cell extract Only for research applications, not for diagnostic or therapeutic use. Introduction Specificity Poly(ADP-ribose) polymerase
PARP1-Trap_A for Immunoprecipitation of PARP1- Fusion Proteins from cell extract Only for research applications, not for diagnostic or therapeutic use. Introduction Specificity Poly(ADP-ribose) polymerase
Ampli1 LowPass Kit. USER MANUAL Version 2.0. Low-pass WGS library prep kit for IonTorrent platforms. Content version: January 2017
 For research use only. Not for use in diagnostic procedures. For in vitro use only. Ampli1 LowPass Kit Low-pass WGS library prep kit for IonTorrent platforms USER MANUAL Version 2.0 Content version: January
For research use only. Not for use in diagnostic procedures. For in vitro use only. Ampli1 LowPass Kit Low-pass WGS library prep kit for IonTorrent platforms USER MANUAL Version 2.0 Content version: January
Ampli1 LowPass Kit. USER MANUAL Version 3.0. Low-pass WGS library prep kit for IonTorrent platforms. Content version: July 2017
 For research use only. Not for use in diagnostic procedures. For in vitro use only. Ampli1 LowPass Kit Low-pass WGS library prep kit for IonTorrent platforms USER MANUAL Version 3.0 Content version: July
For research use only. Not for use in diagnostic procedures. For in vitro use only. Ampli1 LowPass Kit Low-pass WGS library prep kit for IonTorrent platforms USER MANUAL Version 3.0 Content version: July
M-Beads Magnetic silica beads DNA 3.0 (COOH) Order #: PR-MAG00078 & PR-MAG00079
 M-Beads Magnetic silica beads DNA 3.0 (COOH) Order #: PR-MAG00078 & PR-MAG00079 MoBiTec GmbH 2015 Page 2 Contents Intended Use... 3 Principle... 3 Silica & Carboxylated M-Beads Magnetic silica beads DNA
M-Beads Magnetic silica beads DNA 3.0 (COOH) Order #: PR-MAG00078 & PR-MAG00079 MoBiTec GmbH 2015 Page 2 Contents Intended Use... 3 Principle... 3 Silica & Carboxylated M-Beads Magnetic silica beads DNA
empcr Amplification Method Manual Lib L LV
 empcr Amplification Method Manual Lib L LV GS FLX+ Series XL+ May 2011 For life science research only. Not for use in diagnostic procedures. Instrument / Kit GS Junior / Junior GS FLX+ / XL+ GS FLX+ /
empcr Amplification Method Manual Lib L LV GS FLX+ Series XL+ May 2011 For life science research only. Not for use in diagnostic procedures. Instrument / Kit GS Junior / Junior GS FLX+ / XL+ GS FLX+ /
ChargeSwitch gdna Rendered Meat Purification Kit
 USER GUIDE ChargeSwitch gdna Rendered Meat Purification Kit Purification of genomic DNA (gdna) from cattle feed, meal, and heparin products Catalog Number CS400-100 Publication Number MAN0000574 Revision
USER GUIDE ChargeSwitch gdna Rendered Meat Purification Kit Purification of genomic DNA (gdna) from cattle feed, meal, and heparin products Catalog Number CS400-100 Publication Number MAN0000574 Revision
Solutions for purifying nucleic acids by solidphase reversible immobilization (SPRI)
 Solutions for purifying nucleic acids by solidphase reversible immobilization (SPRI) Philippe Jolivet and Joseph W. Foley Ludmer Centre for Neuroinformatics and Mental Health October 21, 2015 Based on
Solutions for purifying nucleic acids by solidphase reversible immobilization (SPRI) Philippe Jolivet and Joseph W. Foley Ludmer Centre for Neuroinformatics and Mental Health October 21, 2015 Based on
Ribo-Zero Magnetic Kits*
 * Ribo-Zero Kit Catalog Number 6-Reactions Catalog Number 24-Reactions Ribo-Zero Magnetic Gold Kit (Human/Mouse/Rat) MRZG126 MRZG12324 Ribo-Zero Magnetic Kit (Human/Mouse/Rat) MRZH116 MRZH11124 Ribo-Zero
* Ribo-Zero Kit Catalog Number 6-Reactions Catalog Number 24-Reactions Ribo-Zero Magnetic Gold Kit (Human/Mouse/Rat) MRZG126 MRZG12324 Ribo-Zero Magnetic Kit (Human/Mouse/Rat) MRZH116 MRZH11124 Ribo-Zero
chemagic mrna Direct Kit
 chemagic mrna Direct Kit for general purposes Kit for the direct isolation of mrna from animal and plant tissue and cells. Kit Components M-PVA OdT Magnetic Beads Suspension Buffer 1 Lysis Buffer 2 Wash
chemagic mrna Direct Kit for general purposes Kit for the direct isolation of mrna from animal and plant tissue and cells. Kit Components M-PVA OdT Magnetic Beads Suspension Buffer 1 Lysis Buffer 2 Wash
TrueBlot Protein G Magnetic Beads PG ml. TrueBlot Protein G Magnetic Beads PG ml. Bead Mean Diameter 0.5 µm. Bead Concentration
 Rockland s TrueBlot Protein G Magnetic Beads are uniform, non-aggregating, super-paramagnetic beads coupled with a biomolecule, such as Protein G. These beads are specifically designed, tested and quality
Rockland s TrueBlot Protein G Magnetic Beads are uniform, non-aggregating, super-paramagnetic beads coupled with a biomolecule, such as Protein G. These beads are specifically designed, tested and quality
AffiAmino UltraRapid Agarose
 Product no 1003 AffiAmino UltraRapid Agarose Product Information Lab on a Bead AB Edition 20151030 All rights reserved Copyright 2015 Lab on a Bead AB Table of Contents 1. General information... 3 2. Principle
Product no 1003 AffiAmino UltraRapid Agarose Product Information Lab on a Bead AB Edition 20151030 All rights reserved Copyright 2015 Lab on a Bead AB Table of Contents 1. General information... 3 2. Principle
AGENCOURT ORAPURE Buccal Cell DNA Isolation Kit
 Buccal Cell DNA Isolation Kit Page 1 of 12 Please refer to http://www.agencourt.com/technical for updated protocols and refer to MSDS instructions when handling or shipping any chemical hazards. AGENCOURT
Buccal Cell DNA Isolation Kit Page 1 of 12 Please refer to http://www.agencourt.com/technical for updated protocols and refer to MSDS instructions when handling or shipping any chemical hazards. AGENCOURT
AdnaTest EMT-1/StemCellSelect
 AdnaTest EMT-1/StemCellSelect Enrichment of tumor cells from blood for gene expression analysis For research use only Manual T-1-533 Contents Order Information... 3 Purpose... 3 Abbreviations and Symbols...
AdnaTest EMT-1/StemCellSelect Enrichment of tumor cells from blood for gene expression analysis For research use only Manual T-1-533 Contents Order Information... 3 Purpose... 3 Abbreviations and Symbols...
mag maxi kit Intended use of the mag maxi kits
 mag maxi kit For in vitro diagnostic use 40403 40430 10 288 May 2014 LGC Genomics GmbH Ostendstr. 25 TGS Haus 8 12459 Berlin Germany Tel: +49 (0)30 5304 2200 Fax: +49 (0)30 5304 2201 Intended use of the
mag maxi kit For in vitro diagnostic use 40403 40430 10 288 May 2014 LGC Genomics GmbH Ostendstr. 25 TGS Haus 8 12459 Berlin Germany Tel: +49 (0)30 5304 2200 Fax: +49 (0)30 5304 2201 Intended use of the
User Guide. rrna Depletion Kit V1.2
 rrna Depletion Kit V1.2 User Guide Catalog Number: 037 (RiboCop rrna Depletion Kit V1.2 (Human/Mouse/Rat)) 042 (SENSE Total RNA-Seq Library Prep Kit for Illumina with RiboCop) 037UG073V0201 FOR RESEARCH
rrna Depletion Kit V1.2 User Guide Catalog Number: 037 (RiboCop rrna Depletion Kit V1.2 (Human/Mouse/Rat)) 042 (SENSE Total RNA-Seq Library Prep Kit for Illumina with RiboCop) 037UG073V0201 FOR RESEARCH
User Guide. rrna Depletion Kit
 rrna Depletion Kit User Guide Catalog Number: 037.24 (RiboCop rrna Depletion Kit (Human/Mouse/Rat), 24 preps) 037.96 (RiboCop rrna Depletion Kit (Human/Mouse/Rat), 96 preps) 042.08 (SENSE Total RNA-Seq
rrna Depletion Kit User Guide Catalog Number: 037.24 (RiboCop rrna Depletion Kit (Human/Mouse/Rat), 24 preps) 037.96 (RiboCop rrna Depletion Kit (Human/Mouse/Rat), 96 preps) 042.08 (SENSE Total RNA-Seq
AdnaTest OvarianCancer-2 Select
 AdnaTest OvarianCancer-2 Select Enrichment of tumor cells from blood of ovarian cancer patients for gene expression analysis For research use only Manual T-1-538 Contents Order Information... 3 Purpose...
AdnaTest OvarianCancer-2 Select Enrichment of tumor cells from blood of ovarian cancer patients for gene expression analysis For research use only Manual T-1-538 Contents Order Information... 3 Purpose...
Direct Polysome IP from Brain Tissue Myriam Heiman:
 Direct Polysome IP from Brain Tissue Myriam Heiman: bonillm@rockefeller.edu Protocol below is for 1 IP, scale accordingly General Notes: -7 mouse striata pooled per IP -IP with 50 µg 19C8 and 50 µg 19F7
Direct Polysome IP from Brain Tissue Myriam Heiman: bonillm@rockefeller.edu Protocol below is for 1 IP, scale accordingly General Notes: -7 mouse striata pooled per IP -IP with 50 µg 19C8 and 50 µg 19F7
In nucleus Hi- C protocol for C. elegans embryos
 In nucleus Hi- C protocol for C. elegans embryos Compiled by Erika Anderson, July 2016. Crosslinking, isolating nuclei, and digestion 1. Bleach gravid hermaphrodites to obtain at least 0.5g of embryos.
In nucleus Hi- C protocol for C. elegans embryos Compiled by Erika Anderson, July 2016. Crosslinking, isolating nuclei, and digestion 1. Bleach gravid hermaphrodites to obtain at least 0.5g of embryos.
FastTrack MAG mrna Isolation Kits
 USER GUIDE FastTrack MAG mrna Isolation Kits For isolating high-quality mrna from total RNA, cells, and tissue Catalog Numbers K1580-01 and K1580-02 Document Part Number 25-0754 Publication Number MAN0000475
USER GUIDE FastTrack MAG mrna Isolation Kits For isolating high-quality mrna from total RNA, cells, and tissue Catalog Numbers K1580-01 and K1580-02 Document Part Number 25-0754 Publication Number MAN0000475
ISOFECAL for Beads Beating Manual (First edition)
 Fecal DNA Extraction Kit ISOFECAL for Beads Beating Manual (First edition) Code No. 315-06281 NIPPON GENE CO., LTD. Table of contents I Product description 1 II Contents of kit 1 III Storage 2 IV Precautions
Fecal DNA Extraction Kit ISOFECAL for Beads Beating Manual (First edition) Code No. 315-06281 NIPPON GENE CO., LTD. Table of contents I Product description 1 II Contents of kit 1 III Storage 2 IV Precautions
Poly(A) RNA Selection Kit User Guide
 Poly(A) RNA Selection Kit User Guide 039 (Poly(A) RNA Selection Kit) 009 (SENSE Total RNA-Seq Library Prep Kit for Illumina, including Barcodes) 020 (PCR Add-on Kit for Illumina) 022 (Purification Module
Poly(A) RNA Selection Kit User Guide 039 (Poly(A) RNA Selection Kit) 009 (SENSE Total RNA-Seq Library Prep Kit for Illumina, including Barcodes) 020 (PCR Add-on Kit for Illumina) 022 (Purification Module
Optional Agencourt SPRIPlate Super Magnet Plate (Beckman Coulter A32782)
 V2.2 Serapure B. Faircloth & T. Glenn November 19, 2011 Ecol. and Evol. Biology Univ. of California Los Angeles The goal here is to create a substitute for AMPure XP that is of equal effectiveness in comparison
V2.2 Serapure B. Faircloth & T. Glenn November 19, 2011 Ecol. and Evol. Biology Univ. of California Los Angeles The goal here is to create a substitute for AMPure XP that is of equal effectiveness in comparison
ChargeSwitch NoSpin Plasmid Kits
 USER GUIDE ChargeSwitch NoSpin Plasmid Kits For purification of plasmid DNA from bacterial cells using the MagnaClear Technology Catalog nos. CS10200, CS10201, CS10201-10 Version A 5 January 2005 25-0813
USER GUIDE ChargeSwitch NoSpin Plasmid Kits For purification of plasmid DNA from bacterial cells using the MagnaClear Technology Catalog nos. CS10200, CS10201, CS10201-10 Version A 5 January 2005 25-0813
Strep-tag Purification using MagStrep type3 XT Beads
 Strep-tag Purification using MagStrep type3 XT Beads Instruction manual Last date of revision November 2016 Version PR83-0004 For research only Important licensing information Products featuring Strep-Tactin
Strep-tag Purification using MagStrep type3 XT Beads Instruction manual Last date of revision November 2016 Version PR83-0004 For research only Important licensing information Products featuring Strep-Tactin
Strep-tag Purification using MagStrep type3 XT Beads
 Strep-tag Purification using MagStrep type3 XT Beads Instruction manual Last date of revision November 2016 Version PR83-0004 For research only Important licensing information Products featuring Strep-Tactin
Strep-tag Purification using MagStrep type3 XT Beads Instruction manual Last date of revision November 2016 Version PR83-0004 For research only Important licensing information Products featuring Strep-Tactin
Total RNA Isolation. User Manual. NucleoMag 96 RNA MACHEREY-NAGEL. January 2010 / Rev. 02
 Total RNA Isolation User Manual NucleoMag 96 RNA January 2010 / Rev. 02 MACHEREY-NAGEL Table of contents 1 Components 4 1.1 Kit contents 4 1.2 Material to be supplied by user 5 2 Product description 6
Total RNA Isolation User Manual NucleoMag 96 RNA January 2010 / Rev. 02 MACHEREY-NAGEL Table of contents 1 Components 4 1.1 Kit contents 4 1.2 Material to be supplied by user 5 2 Product description 6
ChargeSwitch gdna Blood Kits
 Instruction Manual ChargeSwitch gdna Blood Kits For purification of genomic DNA from small volumes of human blood Catalog nos. CS11000, CS11010, and CS11010-10 Version A 6 January 2005 25-0814 ii Table
Instruction Manual ChargeSwitch gdna Blood Kits For purification of genomic DNA from small volumes of human blood Catalog nos. CS11000, CS11010, and CS11010-10 Version A 6 January 2005 25-0814 ii Table
Strep-tag Purification using MagStrep type3 XT Beads
 Strep-tag Purification using MagStrep type3 XT Beads Instruction manual Last date of revision April June 2014 2012 Version PR24-0001 PR83-0004 www.strep-tag.com For research only Important licensing information
Strep-tag Purification using MagStrep type3 XT Beads Instruction manual Last date of revision April June 2014 2012 Version PR24-0001 PR83-0004 www.strep-tag.com For research only Important licensing information
Enriching Beads Oligo (dt) Magnetic Beads for mrna Purification
 Enriching Beads Oligo (dt) Magnetic Beads for mrna Purification Isolate the mrna transcriptome in 15 minutes User Guidance Enriching Biotechnology Rev. 1.0 October 25th. 2018 Why choose Enriching Beads
Enriching Beads Oligo (dt) Magnetic Beads for mrna Purification Isolate the mrna transcriptome in 15 minutes User Guidance Enriching Biotechnology Rev. 1.0 October 25th. 2018 Why choose Enriching Beads
Microwell-Seq. High-throughput Single Cell RNA-Seq Kit. Protocol. Index
 Microwell-Seq High-throughput Single Cell RNA-Seq Kit Protocol Index 1 Introduction 2 Kit Reagent 3 Store 4 Application 5 Prepared materials 6 Note 7 Preparation 8 Workflow 1 1 Introduction Microwell-Seq
Microwell-Seq High-throughput Single Cell RNA-Seq Kit Protocol Index 1 Introduction 2 Kit Reagent 3 Store 4 Application 5 Prepared materials 6 Note 7 Preparation 8 Workflow 1 1 Introduction Microwell-Seq
GS FLX/Junior Titanium Technology
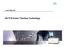 www.454.com GS FLX/Junior Titanium Technology GS FLX and GS Junior Process Steps Overview gdna 1. DNA Library Construction * 4h 2. empcr 3. Sequencing 8 h 10 h Data output DNA Library Preparation Prepare
www.454.com GS FLX/Junior Titanium Technology GS FLX and GS Junior Process Steps Overview gdna 1. DNA Library Construction * 4h 2. empcr 3. Sequencing 8 h 10 h Data output DNA Library Preparation Prepare
Mayr lab 3'-seq protocol, October 2013
 Mayr lab 3'-seq protocol, October 2013 Time line: Day -1: DNase treat your RNA samples (optional) Day 1: Prepare beads, anneal oligo to beads, 1 st, 2 nd strand synthesis Day 2: Introduce nick (Rnase HII),
Mayr lab 3'-seq protocol, October 2013 Time line: Day -1: DNase treat your RNA samples (optional) Day 1: Prepare beads, anneal oligo to beads, 1 st, 2 nd strand synthesis Day 2: Introduce nick (Rnase HII),
M-Beads Magnetic Silica Beads WAX
 M-Beads Magnetic Silica Beads WAX MoBiTec GmbH 2012 Page 2 Contents Technical data... 3 Application... 4 General information... 4 Bead usage... 4 Additional materials needed... 4 Protocols... 5 Order Information,
M-Beads Magnetic Silica Beads WAX MoBiTec GmbH 2012 Page 2 Contents Technical data... 3 Application... 4 General information... 4 Bead usage... 4 Additional materials needed... 4 Protocols... 5 Order Information,
Nature Protocols: doi: /nprot Supplementary Figure 1. Read quality score per sequenced base position.
 Supplementary Figure 1 Read quality score per sequenced base position. (a) Illumina Q-scores for a normal QiSeq run. Typically, more than 80% of reads reach a quality score higher than 30 across all read
Supplementary Figure 1 Read quality score per sequenced base position. (a) Illumina Q-scores for a normal QiSeq run. Typically, more than 80% of reads reach a quality score higher than 30 across all read
pluribead KIT Cell Separation Protocol M-Bead
 pluribead KIT Cell Separation Protocol M-Bead pluriselect@hiss-dx.de 15 14 13 12 11 9 8 7 6 5 4 3 2 1 50 40 30 20 2 Contents Contents pluribead Kit Components & Additional Materials 2 Separation Protocol
pluribead KIT Cell Separation Protocol M-Bead pluriselect@hiss-dx.de 15 14 13 12 11 9 8 7 6 5 4 3 2 1 50 40 30 20 2 Contents Contents pluribead Kit Components & Additional Materials 2 Separation Protocol
LOABeads AffiAmino. Product Manual. Lab on a Bead AB. Revision date Copyright Lab on a Bead AB All rights reserved
 LOABeads AffiAmino Product Manual Lab on a Bead AB Revision date 2016-11-23 Copyright 2015-2016 Lab on a Bead AB All rights reserved Table of Contents 1. General information...3 2. Product data...4 3.
LOABeads AffiAmino Product Manual Lab on a Bead AB Revision date 2016-11-23 Copyright 2015-2016 Lab on a Bead AB All rights reserved Table of Contents 1. General information...3 2. Product data...4 3.
RayBio anti-mouse IgG Magnetic Beads
 RayBio anti-mouse IgG Magnetic Beads Catalog #: 801-103 User Manual Last revised January 4 th, 2017 Caution: Extraordinarily useful information enclosed ISO 1348 Certified 3607 Parkway Lane, Suite 100
RayBio anti-mouse IgG Magnetic Beads Catalog #: 801-103 User Manual Last revised January 4 th, 2017 Caution: Extraordinarily useful information enclosed ISO 1348 Certified 3607 Parkway Lane, Suite 100
AbraMag TM Magnetic Beads
 AbraMag TM Magnetic Beads Abraxis, Inc., founded in 1998, is a biotechnology company that develops, manufactures, markets, and distributes products and services to meet the needs of research and industry.
AbraMag TM Magnetic Beads Abraxis, Inc., founded in 1998, is a biotechnology company that develops, manufactures, markets, and distributes products and services to meet the needs of research and industry.
BoTest Matrix E Botulinum Neurotoxin Detection Kit Protocol
 BoTest Matrix E Botulinum Neurotoxin Detection Kit Protocol 505 S. Rosa Road, Suite 105 Madison, WI 53719 1-608-441-8174 info@biosentinelpharma.com BioSentinel Part No: L1016, Release Date: May 29, 2014
BoTest Matrix E Botulinum Neurotoxin Detection Kit Protocol 505 S. Rosa Road, Suite 105 Madison, WI 53719 1-608-441-8174 info@biosentinelpharma.com BioSentinel Part No: L1016, Release Date: May 29, 2014
CeMM- ChIPmentation protocol v1.1 (2015/10/14)
 CeMM- ChIPmentation protocol v1.1 (2015/10/14) Authors: Christian Schmidl (cschmidl@cemm.oeaw.ac.at) and Christoph Bock (cbock@cemm.oeaw.ac.at). Paper website: http://chipmentation.computational-epigenetics.org
CeMM- ChIPmentation protocol v1.1 (2015/10/14) Authors: Christian Schmidl (cschmidl@cemm.oeaw.ac.at) and Christoph Bock (cbock@cemm.oeaw.ac.at). Paper website: http://chipmentation.computational-epigenetics.org
Amine Magnetic Beads
 588PR-02 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Amine Magnetic Beads (Cat. # 786-906, 786-907) think proteins! think G-Biosciences
588PR-02 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Amine Magnetic Beads (Cat. # 786-906, 786-907) think proteins! think G-Biosciences
Caution: For Laboratory Use. A product for research purposes only. YSi (2 5 μm) Copper His-Tag SPA Beads. Product Numbers: RPNQ0096
 TECHNICAL DATA SHEET SPA Beads Caution: For Laboratory Use. A product for research purposes only. YSi (2 5 μm) Copper His-Tag SPA Beads Product Numbers: RPNQ0096 WARNING For research use only. Not recommended
TECHNICAL DATA SHEET SPA Beads Caution: For Laboratory Use. A product for research purposes only. YSi (2 5 μm) Copper His-Tag SPA Beads Product Numbers: RPNQ0096 WARNING For research use only. Not recommended
Technical Manual No. TM0261 Version
 Donkey Anti-Goat IgG MagBeads Cat. No. L00332 Technical Manual No. TM0261 Version 06272010 Index 1. Product Description 2. Instruction For Use 3. Troubleshooting 4. General Information 1. Product Description
Donkey Anti-Goat IgG MagBeads Cat. No. L00332 Technical Manual No. TM0261 Version 06272010 Index 1. Product Description 2. Instruction For Use 3. Troubleshooting 4. General Information 1. Product Description
Applied Biosystems SOLiD 4 System Templated Bead Preparation Guide
 Applied Biosystems SOLiD 4 System Templated Bead Preparation Guide March 2010 Library Preparation Templated Bead Preparation Instrument Operation For Research Use Only. Not intended for any animal or human
Applied Biosystems SOLiD 4 System Templated Bead Preparation Guide March 2010 Library Preparation Templated Bead Preparation Instrument Operation For Research Use Only. Not intended for any animal or human
LOABeads Protein A. Product no Product Manual. Lab on a Bead AB
 Product no 1001 LOABeads Protein A Product Manual Lab on a Bead AB Revision date 2016-03-08 Copyright 2015-2016 Lab on a Bead AB All rights reserved Table of Contents 1. General information 3 2. Antibody
Product no 1001 LOABeads Protein A Product Manual Lab on a Bead AB Revision date 2016-03-08 Copyright 2015-2016 Lab on a Bead AB All rights reserved Table of Contents 1. General information 3 2. Antibody
CELLSCRIPT RNA for Translation in Cells
 H TM CELLSCRIPT RA for Translation in Cells Cat. o. C-C61025 ITRDUCTI 5'-terminal caps are involved in mra processing, stability and initiation of protein synthesis. 1 Uncapped RA transfected or injected
H TM CELLSCRIPT RA for Translation in Cells Cat. o. C-C61025 ITRDUCTI 5'-terminal caps are involved in mra processing, stability and initiation of protein synthesis. 1 Uncapped RA transfected or injected
