JAMP: Joint Genetic Association of Multiple Phenotypes
|
|
|
- Patience Underwood
- 5 years ago
- Views:
Transcription
1 JAMP: Joint Genetic Association of Multiple Phenotypes Manual, version /06/2012 D Posthuma AE van Bochoven Ctglab.nl 1
2 JAMP is a free, open source tool to run multivariate GWAS. It combines information from multiple phenotypes to obtain one combined P- value per SNP. As JAMP uses permutation to determine P- values, raw genotype data is required as input. If you use JAMP, please refer to Posthuma D, van Bochoven AE et al., JAMP: A tool to assess joint association of multiple phenotypes in GWAS datasets. Forthcoming Download JAMP is written in Python and runs on Mac OSX and Linux. It can be downloaded from After download, copy all the JAMP files to your working directory. JAMP is a command line tool, you need to open a terminal window and type commands at the prompt to perform analyses with JAMP. JAMP can be invoked by typing./jamp, from the directory where JAMP is installed. If you want to invoke JAMP from any directory you need to add the path to JAMP to PATH. How to do this, depends on the environment you are working in. Alternatively you can add (a link to) the executable to a folder that is already in your PATH. Typing PATH shows the current PATH settings. If you are using a bash shell in a MAC/UNIX/Linux environment, you need to modify the file.profile (for MacOSX) or.bashrc (for some other linux versions) (in your home directory). Simply add the following line to this script: alias jamp= [insert path to the jamp executable] This allows invoking jamp from any directory by typing jamp at the prompt. The following files are available for download: jamp [the program] JAMP_manual.pdf [the manual] example.bed [example binary pedigree file] example.bim [example binary map file] example.fam [example fam file] Pheno_b.txt [an alternate phenotype file containing 4 binary traits] Pheno_q.txt [an alternate phenotype file containing 4 quantitative traits] Pheno_bq.txt [an alternate phenotype file containing 4 binary and 4 quantitative traits] The example input files are based on the CEU hapmap genotypes (10 SNPs) with randomly generated phenotypes. Besides JAMP you need to have PLINK from Shaun Purcell installed, which can be downloaded freely from here: In addition, python v2.6 or higher is required and if not already installed can be downloaded from: 2
3 Input Files JAMP uses the same input files as PLINK, which can be in plain text rectangular (.ped and.map) format, in transposed or long format, or in the more efficient and widely used binary format (.bed,.bim. and.fam files). Please consult the PLINK documentation ( if you are unfamiliar with these file formats. JAMP expects that multivariate phenotypes are available in a separate file, in the alternate phenotype format as specified by PLINK, with the exception that the phenotype file is not allowed to have a header row for use in JAMP. The first two columns in the phenotype file must contain Family ID and Individual ID, the consecutive columns contain the multiple phenotypes. Also, the delimiter must be a space, not a tab. IMPORTANT: The alternate phenotype file should not contain a header and should have space as a delimiter. Quickstart - running JAMP To obtain a combined P- value based on 100 permutations of all phenotypes for each SNP, provide PLINK input files and an alternative phenotype file without a header and with spaces as delimiter, then type:./jamp - - bfile example - - assoc - - pheno pheno_b.txt - - all- pheno - - jperm 100 This will invoke JAMP which first calls PLINK to run genetic association tests on all SNPs available in the input files and for all phenotypes in pheno_b.txt, then permutes and calls PLINK again to run association on the permuted phenotypes, to obtain multivariate P- values. The - - all- pheno option must always be provided to ensure PLINK runs association on all phenotypes. If it is not included, JAMP will add it. JAMP assumes all phenotypes in the alternate phenotype file need to be analyzed. If you only wish to analyze a subset of your phenotypes it is required to generate a new alternate phenotype file that includes only those phenotypes that you wish to analyze. While permuting JAMP keeps all phenotypic scores from one individual together in order to retain the phenotypic structure in the original data. JAMP thus corrects for the correlational structure between phenotypes and does not make any assumptions on the multivariate nature of the phenotypic data. The phenotypes can be binary, quantitative or a combination of these. Crude permutation controlling family wise error rate The command - - jperm invokes crude permutation and runs the same number of permutations for each SNP. the command - - jperm 1000 runs 1000 permutations for all SNPs and provides an empirical P value (EMP_P) based on 1000 permutations. The empirical P- value is calculated as follows: JAMP first calculates and stores the Σ- log 10(P) across all phenotypes for each SNP based on the original dataset. Then for each permutation JAMP also calculates the Σ- log 10(P). When finished permuting, JAMP obtains the empirical P- value (P M) for each SNP by dividing the number of times the Σ- log 10(P) from the permuted analyses exceeds or equals the Σ- log 10(P) from the original analysis (hits, H) by the number of permutations run (M): 3
4 P M = H M As the same number of permutations for every SNP is run, JAMP calculates an additional empirical P- value (EMP_P_COR) that controls for the family wise error due to testing multiple SNPs. This is achieved by comparing every observed Σ- log 10(P) with the maximum Σ- log 10(P) obtained across all SNPs for each permutation. The empirical P valued based on the Σ- log 10(P) test statistic, tests the hypothesis that the multivariate pattern of P- values of all phenotypes is significantly different than what is expected under the null hypothesis of no association. A significant P value is thus suggestive of multivariate association with a SNP. In addition to this test, JAMP produces a second empirical P- value (EMP_Pmin) to test the hypothesis that at least one of the phenotypes is significantly associated with a SNP, given the multivariate nature of the phenotypes. For each SNP, the smallest P value from the original range of P- values from all phenotypes is evaluated against the smallest P- value from all univariate P- values obtained in each permutation, thus correcting for the multivariate nature of the data. In addition, the original smallest P- value is evaluated against the smallest of the smallest P- values across all SNPs, providing the EMP_Pmin_CORR wihich is corrected for testing multiple SNPs. The output of - - jperm thus produces two P- values per tested hypothesis: one empirical P- value which is corrected for testing multiuple phenotypes, but uncorrected for testing multiple SNPs (and which needs to be evaluated against a generally accepted genome- wide significance level based on Bonferroni correction for multiple testing) and one empirical P- value which is corrected testing both multiple phenotypes and multiple SNPs which can be evaluated against a nominal significance level of 0.01 or This latter P- value corrects for multiple testing conditional on the genomic data and tends to be less conservative compared to Bonferroni. As it is usually sufficient to show that the corrected P- value is <.05 or.01, only permutations are needed with the crude permutation scheme. When finished, a file called jamp.empp is created, containing the following columns: CHR The name of the chromosome SNP The SNP- id NPHENO The number of phenotypes for which the analysis was run P_P1 The P- value of the 1 st phenotype, as produced by PLINK P_P2 The P- value of the 2 nd phenotype, as produced by PLINK.... P_Pn The P- value of the n th phenotype, as produced by PLINK SUMLOGP The Σ- log 10(P) across all phenotypes for one SNP NPERMS The number of permutations run for each SNP EMP_P The empirical P- value of the multivariate SNP association EMP_P_COR EMP_P corrected for the family- wise error rate of testing multiple SNPs EMP_Pmin The empirical P- value of the test that at least one phenotype is significantly associated given the multivariate nature of the phenotypes 4
5 EMP_Pmin_COR EMP_Pmin corrected for the family- wise error rate of testing multiple SNPs Note that the - - jperm [number] option not only specifies how many permutations need to be carried out, but it also specifies the seed numbers, as JAMP takes the number of the permutation as seed number. This can come in handy if you wish to reproduce exactly the same results. However if you wish to split up the permutations in two batches of 500 each, you need to ensure that you do not obtain two batches of exactly the same permutations. The command - - jperm specifies that 500 permutations will be run with permutation (seed) numbers whereas the command - - jperm specifies a different set of 500 permutations with seed numbers It is generally practical to use this in combination with the option - - out which adds a prefix to the JAMP output files. For example:./jamp - - bfile example - - assoc - - pheno pheno.txt - - all- pheno - - jperm out run1 and./jamp - - bfile example - - assoc - - pheno pheno.txt - - all- pheno - - jperm out run2 will generate run1.jamp.empp and run2.jamp.empp The empirical P- value based on all 1000 permutations can easily be obtained afterwards with the command jmerge:./jamp - - jmerge run1.jamp.empp run2.jamp.empp - - out all Or, if you have a long list of runs you can use a wildcard:./jamp - - jmerge run*.jamp.empp - - out all.empp This will generate an output file called all.empp.jamp.merged that includes an empirical P value based on the total number of permutations. If JAMP is used to run multiple permutations simultaneously (for example using a cluster), JAMP will start each set of permutations with running PLINK on the actual data. In some cases (i.e. when the original analysis takes a long time) it may be convenient to provide JAMP with output files from the actual run to avoid running the same analyses multiple times and to save computing time. The command - - jstart invokes this behavior. For example./jamp - - bfile example - - assoc - - pheno pheno.txt - - all- pheno - - jperm out run1 - - jstart run0 Will cause jamp to search for the following files run0.jamp.chr_snp run0.npheno_pheno_x run0.jamp.sumlogp 5
6 JAMP will then skip running PLINK on the actual data and will start permutation right away. These files from the original run can be obtained by running jamp with zero permutations:./jamp - - bfile example - - assoc - - pheno pheno.txt - - all- pheno - - jperm out run0 IMPORTANT: When using - - jstart it is important that the provided files are based on exactly the same dataset as specified with the - - bfile option Supported PLINK options JAMP currently supports the following options for association in PLINK: - - assoc - - linear - - logistic - - trend - - model - - dosage JAMP currently does not support the - - mh, - - adjust or any of the family based association options from PLINK. In theory all other options that are used in PLINK can be added on the command line when calling JAMP. However, since JAMP uses permutation, it is often a good idea to pre- run options that require some time, especially when running 100 or 1000 permutations. In particular options intended to clean the datafiles are advised to be used with - - make- bed in PLINK prior to running JAMP (e.g. - - extract - - remove - - hwe - - maf ). Some PLINK options require special attention when running JAMP: - - out [prefix] - - sex - - covar [file.txt] - - adjust The PLINK option - - out changes all prefixes in PLINK, and also in JAMP If you wish to correct for sex, you have to put the sex codes in a covariate file and use - - covar, do not use the - - sex option with JAMP When you use this option, JAMP will permute all covariates with the phenotypes, i.e. the relation between the covariates and the phenotypes is retained and the analyses on the permuted datasets are carried out using the same corrected phenotypes as in the original analyses. The file containing covariates should be in the PLINK covariate file format, except that it should not contain a header (whereas PLINK accepts both with and without a header), and have spaces and not tabs. Note that the JAMP output will also contain p- values for the covariates JAMP currently does not support taking the GC corrected P- values, if you use - - adjust and - - assoc, JAMP will work with the output from - - assoc and - - adjust will only slow down the permutation procedure. 6
Genome-Wide Association Exercise - Data Quality Control
 Genome-Wide Association Exercise - Data Quality Control The Rockefeller University, New York, June 25, 2016 Copyright 2016 Merry-Lynn McDonald & Suzanne M. Leal Introduction In this exercise, you will
Genome-Wide Association Exercise - Data Quality Control The Rockefeller University, New York, June 25, 2016 Copyright 2016 Merry-Lynn McDonald & Suzanne M. Leal Introduction In this exercise, you will
Linkage Analysis in Merlin. Meike Bartels Kate Morley Danielle Posthuma
 Linkage Analysis in Merlin Meike Bartels Kate Morley Danielle Posthuma Software for linkage analyses Genehunter Mendel Vitesse Allegro Simwalk Loki Merlin. Mx R Lisrel MERLIN software Programs: MERLIN
Linkage Analysis in Merlin Meike Bartels Kate Morley Danielle Posthuma Software for linkage analyses Genehunter Mendel Vitesse Allegro Simwalk Loki Merlin. Mx R Lisrel MERLIN software Programs: MERLIN
Illumina GenomeStudio Analysis
 Illumina GenomeStudio Analysis Paris Veltsos University of St Andrews February 23, 2012 1 Introduction GenomeStudio is software by Illumina used to score SNPs based on the Illumina BeadExpress platform.
Illumina GenomeStudio Analysis Paris Veltsos University of St Andrews February 23, 2012 1 Introduction GenomeStudio is software by Illumina used to score SNPs based on the Illumina BeadExpress platform.
Detecting Heterogeneity in Population Structure Across the Genome in Admixed Populations
 Genetics: Early Online, published on July 20, 2016 as 10.1534/genetics.115.184184 GENETICS INVESTIGATION Detecting Heterogeneity in Population Structure Across the Genome in Admixed Populations Caitlin
Genetics: Early Online, published on July 20, 2016 as 10.1534/genetics.115.184184 GENETICS INVESTIGATION Detecting Heterogeneity in Population Structure Across the Genome in Admixed Populations Caitlin
TDT vignette Use of snpstats in family based studies
 TDT vignette Use of snpstats in family based studies David Clayton April 30, 2018 Pedigree data The snpstats package contains some tools for analysis of family-based studies. These assume that a subject
TDT vignette Use of snpstats in family based studies David Clayton April 30, 2018 Pedigree data The snpstats package contains some tools for analysis of family-based studies. These assume that a subject
Two-point linkage analysis using the LINKAGE/FASTLINK programs
 1 Two-point linkage analysis using the LINKAGE/FASTLINK programs Copyrighted 2018 Maria Chahrour and Suzanne M. Leal These exercises will introduce the LINKAGE file format which is the standard format
1 Two-point linkage analysis using the LINKAGE/FASTLINK programs Copyrighted 2018 Maria Chahrour and Suzanne M. Leal These exercises will introduce the LINKAGE file format which is the standard format
fbat August 21, 2010 Basic data quality checks for markers
 fbat August 21, 2010 checkmarkers Basic data quality checks for markers Basic data quality checks for markers. checkmarkers(genesetobj, founderonly=true, thrsh=0.05, =TRUE) checkmarkers.default(pedobj,
fbat August 21, 2010 checkmarkers Basic data quality checks for markers Basic data quality checks for markers. checkmarkers(genesetobj, founderonly=true, thrsh=0.05, =TRUE) checkmarkers.default(pedobj,
Factors affecting phasing quality in a commercial layer population
 Factors affecting phasing quality in a commercial layer population N. Frioni 1, D. Cavero 2, H. Simianer 1 & M. Erbe 3 1 University of Goettingen, Department of nimal Sciences, Center for Integrated Breeding
Factors affecting phasing quality in a commercial layer population N. Frioni 1, D. Cavero 2, H. Simianer 1 & M. Erbe 3 1 University of Goettingen, Department of nimal Sciences, Center for Integrated Breeding
Detection of Misspecified Relationships in Inbred and Outbred Pedigrees
 Detection of Misspecified Relationships in Inbred and Outbred Pedigrees Lei Sun 1, Mark Abney 1,2, Mary Sara McPeek 1,2 1 Department of Statistics, 2 Department of Human Genetics, University of Chicago,
Detection of Misspecified Relationships in Inbred and Outbred Pedigrees Lei Sun 1, Mark Abney 1,2, Mary Sara McPeek 1,2 1 Department of Statistics, 2 Department of Human Genetics, University of Chicago,
Package RVtests. R topics documented: February 19, 2015
 Type Package Title Rare Variant Tests Version 1.2 Date 2013-05-27 Author, and C. M. Greenwood Package RVtests February 19, 2015 Maintainer Depends R (>= 2.12.1), glmnet,
Type Package Title Rare Variant Tests Version 1.2 Date 2013-05-27 Author, and C. M. Greenwood Package RVtests February 19, 2015 Maintainer Depends R (>= 2.12.1), glmnet,
ville, VA Associate Editor: XXXXXXX Received on XXXXX; revised on XXXXX; accepted on XXXXX
 Robust Relationship Inference in Genome Wide Association Studies Ani Manichaikul 1,2, Josyf Mychaleckyj 1, Stephen S. Rich 1, Kathy Daly 3, Michele Sale 1,4,5 and Wei- Min Chen 1,2,* 1 Center for Public
Robust Relationship Inference in Genome Wide Association Studies Ani Manichaikul 1,2, Josyf Mychaleckyj 1, Stephen S. Rich 1, Kathy Daly 3, Michele Sale 1,4,5 and Wei- Min Chen 1,2,* 1 Center for Public
Package garfield. March 8, 2019
 Package garfield March 8, 2019 Type Package Title GWAS Analysis of Regulatory or Functional Information Enrichment with LD correction Version 1.10.0 Date 2015-12-14 Author Sandro Morganella
Package garfield March 8, 2019 Type Package Title GWAS Analysis of Regulatory or Functional Information Enrichment with LD correction Version 1.10.0 Date 2015-12-14 Author Sandro Morganella
Mapping small-effect and linked quantitative trait loci for complex traits in. backcross or DH populations via a multi-locus GWAS methodology
 Mapping small-effect and linked quantitative trait loci for complex traits in backcross or DH populations via a multi-locus GWAS methodology Shi-Bo Wang 1,2, Yang-Jun Wen 2, Wen-Long Ren 2, Yuan-Li Ni
Mapping small-effect and linked quantitative trait loci for complex traits in backcross or DH populations via a multi-locus GWAS methodology Shi-Bo Wang 1,2, Yang-Jun Wen 2, Wen-Long Ren 2, Yuan-Li Ni
An ImageJ based measurement setup for automated phenotyping of plants
 An ImageJ based measurement setup for automated phenotyping of plants J. Kokorian a,c, G. Polder b, J.J.B. Keurentjes a, D. Vreugdenhil a,c, M. Olortegui Guzman a a Laboratory of Plant Physiology, Wageningen
An ImageJ based measurement setup for automated phenotyping of plants J. Kokorian a,c, G. Polder b, J.J.B. Keurentjes a, D. Vreugdenhil a,c, M. Olortegui Guzman a a Laboratory of Plant Physiology, Wageningen
Scott Wolfe Department of Horticulture and Crop Science The Ohio State University, OARDC Wooster, Ohio
 Scott Wolfe Department of Horticulture and Crop Science The Ohio State University, OARDC Wooster, Ohio wolfe.529@osu.edu Purpose Show how to download, install, and run MapMaker 3.0b Show how to properly
Scott Wolfe Department of Horticulture and Crop Science The Ohio State University, OARDC Wooster, Ohio wolfe.529@osu.edu Purpose Show how to download, install, and run MapMaker 3.0b Show how to properly
Gene coancestry in pedigrees and populations
 Gene coancestry in pedigrees and populations Thompson, Elizabeth University of Washington, Department of Statistics Box 354322 Seattle, WA 98115-4322, USA E-mail: eathomp@uw.edu Glazner, Chris University
Gene coancestry in pedigrees and populations Thompson, Elizabeth University of Washington, Department of Statistics Box 354322 Seattle, WA 98115-4322, USA E-mail: eathomp@uw.edu Glazner, Chris University
GEDmatch Home Page The upper left corner of your home page has Information about you and links to lots of helpful information. Check them out!
 USING GEDMATCH Created March 2015 GEDmatch is a free, non-profit site that accepts raw autosomal data files from Ancestry, FTDNA, and 23andme. As such, it provides a large autosomal database that spans
USING GEDMATCH Created March 2015 GEDmatch is a free, non-profit site that accepts raw autosomal data files from Ancestry, FTDNA, and 23andme. As such, it provides a large autosomal database that spans
LeCroy UWBSpekChek WiMedia Compliance Test Suite User Guide. Introduction
 LeCroy UWBSpekChek WiMedia Compliance Test Suite User Guide Version 3.10 March, 2008 Introduction LeCroy UWBSpekChek Application The UWBSpekChek application operates in conjunction with the UWBTracer/Trainer
LeCroy UWBSpekChek WiMedia Compliance Test Suite User Guide Version 3.10 March, 2008 Introduction LeCroy UWBSpekChek Application The UWBSpekChek application operates in conjunction with the UWBTracer/Trainer
Mixing Business Cards in a Box
 Mixing Business Cards in a Box I. Abstract... 2 II. Introduction... 2 III. Experiment... 2 1. Materials... 2 2. Mixing Procedure... 3 3. Data collection... 3 IV. Theory... 4 V. Statistics of the Data...
Mixing Business Cards in a Box I. Abstract... 2 II. Introduction... 2 III. Experiment... 2 1. Materials... 2 2. Mixing Procedure... 3 3. Data collection... 3 IV. Theory... 4 V. Statistics of the Data...
EKA Laboratory Muon Lifetime Experiment Instructions. October 2006
 EKA Laboratory Muon Lifetime Experiment Instructions October 2006 0 Lab setup and singles rate. When high-energy cosmic rays encounter the earth's atmosphere, they decay into a shower of elementary particles.
EKA Laboratory Muon Lifetime Experiment Instructions October 2006 0 Lab setup and singles rate. When high-energy cosmic rays encounter the earth's atmosphere, they decay into a shower of elementary particles.
CONGEN. Inbreeding vocabulary
 CONGEN Inbreeding vocabulary Inbreeding Mating between relatives. Inbreeding depression Reduction in fitness due to inbreeding. Identical by descent Alleles that are identical by descent are direct descendents
CONGEN Inbreeding vocabulary Inbreeding Mating between relatives. Inbreeding depression Reduction in fitness due to inbreeding. Identical by descent Alleles that are identical by descent are direct descendents
Package EILA. February 19, Index 6. The CEU-CHD-YRI admixed simulation data
 Type Package Title Efficient Inference of Local Ancestry Version 0.1-2 Date 2013-09-09 Package EILA February 19, 2015 Author James J. Yang, Jia Li, Anne Buu, and L. Keoki Williams Maintainer James J. Yang
Type Package Title Efficient Inference of Local Ancestry Version 0.1-2 Date 2013-09-09 Package EILA February 19, 2015 Author James J. Yang, Jia Li, Anne Buu, and L. Keoki Williams Maintainer James J. Yang
Selection of Significant Features Using Monte Carlo Feature Selection
 Selection of Significant Features Using Monte Carlo Feature Selection Susanne Bornelöv and Jan Komorowski Abstract Feature selection methods identify subsets of features in large datasets. Such methods
Selection of Significant Features Using Monte Carlo Feature Selection Susanne Bornelöv and Jan Komorowski Abstract Feature selection methods identify subsets of features in large datasets. Such methods
Introduction to ibbig
 Introduction to ibbig Aedin Culhane, Daniel Gusenleitner April 4, 2013 1 ibbig Iterative Binary Bi-clustering of Gene sets (ibbig) is a bi-clustering algorithm optimized for discovery of overlapping biclusters
Introduction to ibbig Aedin Culhane, Daniel Gusenleitner April 4, 2013 1 ibbig Iterative Binary Bi-clustering of Gene sets (ibbig) is a bi-clustering algorithm optimized for discovery of overlapping biclusters
USER GUIDE. NEED HELP? Call us on +44 (0)
 USER GUIDE NEED HELP? Call us on +44 (0) 121 250 3642 TABLE OF CONTENTS Document Control and Authority...3 User Guide...4 Create SPN Project...5 Open SPN Project...6 Save SPN Project...6 Evidence Page...7
USER GUIDE NEED HELP? Call us on +44 (0) 121 250 3642 TABLE OF CONTENTS Document Control and Authority...3 User Guide...4 Create SPN Project...5 Open SPN Project...6 Save SPN Project...6 Evidence Page...7
Objective: Why? 4/6/2014. Outlines:
 Objective: Develop mathematical models that quantify/model resemblance between relatives for phenotypes of a quantitative trait : - based on pedigree - based on markers Outlines: Causal model for covariances
Objective: Develop mathematical models that quantify/model resemblance between relatives for phenotypes of a quantitative trait : - based on pedigree - based on markers Outlines: Causal model for covariances
DFTSimuLab BJStrike Blackjack simulator and strategy analyzer Reference Manual Version 4.0
 DFTSimuLab BJStrike Blackjack simulator and strategy analyzer Reference Manual Version 4.0 Copyright 2010 DFTSimuLab. All rights reserved. Copyright 2009 DFTSimuLab. All rights reserved. Contents 1. Introduction
DFTSimuLab BJStrike Blackjack simulator and strategy analyzer Reference Manual Version 4.0 Copyright 2010 DFTSimuLab. All rights reserved. Copyright 2009 DFTSimuLab. All rights reserved. Contents 1. Introduction
AN-006 APPLICATION NOTE GOLDEN SAMPLE IDENTIFICATION USING CLIO AND SCILAB INTRODUCTION. by Daniele Ponteggia -
 AUDIOMATICA AN-006 APPLICATION NOTE INTRODUCTION GOLDEN SAMPLE IDENTIFICATION USING CLIO AND SCILAB by Daniele Ponteggia - dp@audiomatica.com The efficiency and quality of a manufacturing process can be
AUDIOMATICA AN-006 APPLICATION NOTE INTRODUCTION GOLDEN SAMPLE IDENTIFICATION USING CLIO AND SCILAB by Daniele Ponteggia - dp@audiomatica.com The efficiency and quality of a manufacturing process can be
Developing Conclusions About Different Modes of Inheritance
 Pedigree Analysis Introduction A pedigree is a diagram of family relationships that uses symbols to represent people and lines to represent genetic relationships. These diagrams make it easier to visualize
Pedigree Analysis Introduction A pedigree is a diagram of family relationships that uses symbols to represent people and lines to represent genetic relationships. These diagrams make it easier to visualize
Management and Analysis of Camera Trap Data: Alternative Approaches (Response to Harris et al. 2010)
 Emerging Technologies E m e r g i n g T e c h n o l o g i e s Management and Analysis of Camera Trap Data: Alternative Approaches (Response to Harris et al. 2010) Siva R. Sundaresan, Department of Conservation
Emerging Technologies E m e r g i n g T e c h n o l o g i e s Management and Analysis of Camera Trap Data: Alternative Approaches (Response to Harris et al. 2010) Siva R. Sundaresan, Department of Conservation
New Developments for Mixture Modeling using Mplus
 New Developments for Mixture Modeling using Mplus Bengt Muthén & Tihomir Asparouhov Mplus www.statmodel.com Presentation at the 2013 AERA Meeting in San Francisco April 28, 2013 Bengt Muthén & Tihomir
New Developments for Mixture Modeling using Mplus Bengt Muthén & Tihomir Asparouhov Mplus www.statmodel.com Presentation at the 2013 AERA Meeting in San Francisco April 28, 2013 Bengt Muthén & Tihomir
OzE Field Modules. OzE School. Quick reference pages OzE Main Opening Screen OzE Process Data OzE Order Entry OzE Preview School Promotion Checklist
 1 OzE Field Modules OzE School Quick reference pages OzE Main Opening Screen OzE Process Data OzE Order Entry OzE Preview School Promotion Checklist OzESchool System Features Field unit for preparing all
1 OzE Field Modules OzE School Quick reference pages OzE Main Opening Screen OzE Process Data OzE Order Entry OzE Preview School Promotion Checklist OzESchool System Features Field unit for preparing all
PERMUTATION TESTS FOR COMPLEX DATA
 PERMUTATION TESTS FOR COMPLEX DATA Theory, Applications and Software Fortunato Pesarin Luigi Salmaso University of Padua, Italy TECHNISCHE INFORMATIONSBiBUOTHEK UNIVERSITATSBIBLIOTHEK HANNOVER V WILEY
PERMUTATION TESTS FOR COMPLEX DATA Theory, Applications and Software Fortunato Pesarin Luigi Salmaso University of Padua, Italy TECHNISCHE INFORMATIONSBiBUOTHEK UNIVERSITATSBIBLIOTHEK HANNOVER V WILEY
Project summary. Key findings, Winter: Key findings, Spring:
 Summary report: Assessing Rusty Blackbird habitat suitability on wintering grounds and during spring migration using a large citizen-science dataset Brian S. Evans Smithsonian Migratory Bird Center October
Summary report: Assessing Rusty Blackbird habitat suitability on wintering grounds and during spring migration using a large citizen-science dataset Brian S. Evans Smithsonian Migratory Bird Center October
Kinship/relatedness. David Balding Professor of Statistical Genetics University of Melbourne, and University College London.
 Kinship/relatedness David Balding Professor of Statistical Genetics University of Melbourne, and University College London 2 Feb 2016 1 Ways to measure relatedness 2 Pedigree-based kinship coefficients
Kinship/relatedness David Balding Professor of Statistical Genetics University of Melbourne, and University College London 2 Feb 2016 1 Ways to measure relatedness 2 Pedigree-based kinship coefficients
ANSYS v14.5. Manager Installation Guide CAE Associates
 ANSYS v14.5 Remote Solve Manager Installation Guide 2013 CAE Associates What is the Remote Solve Manager? The Remote Solve Manager (RSM) is a job queuing system designed specifically for use with the ANSYS
ANSYS v14.5 Remote Solve Manager Installation Guide 2013 CAE Associates What is the Remote Solve Manager? The Remote Solve Manager (RSM) is a job queuing system designed specifically for use with the ANSYS
Introduction to ibbig
 Introduction to ibbig Aedin Culhane, Daniel Gusenleitner June 13, 2018 1 ibbig Iterative Binary Bi-clustering of Gene sets (ibbig) is a bi-clustering algorithm optimized for discovery of overlapping biclusters
Introduction to ibbig Aedin Culhane, Daniel Gusenleitner June 13, 2018 1 ibbig Iterative Binary Bi-clustering of Gene sets (ibbig) is a bi-clustering algorithm optimized for discovery of overlapping biclusters
Olympiad Combinatorics. Pranav A. Sriram
 Olympiad Combinatorics Pranav A. Sriram August 2014 Chapter 2: Algorithms - Part II 1 Copyright notices All USAMO and USA Team Selection Test problems in this chapter are copyrighted by the Mathematical
Olympiad Combinatorics Pranav A. Sriram August 2014 Chapter 2: Algorithms - Part II 1 Copyright notices All USAMO and USA Team Selection Test problems in this chapter are copyrighted by the Mathematical
Population Structure. Population Structure
 Nonrandom Mating HWE assumes that mating is random in the population Most natural populations deviate in some way from random mating There are various ways in which a species might deviate from random
Nonrandom Mating HWE assumes that mating is random in the population Most natural populations deviate in some way from random mating There are various ways in which a species might deviate from random
Principles of the Global Positioning System Lecture 20" Processing Software" Primary research programs"
 12.540 Principles of the Global Positioning System Lecture 20" Prof. Thomas Herring" Room 54-820A; 253-5941" tah@mit.edu" http://geoweb.mit.edu/~tah/12.540 " Processing Software" Examine basic features
12.540 Principles of the Global Positioning System Lecture 20" Prof. Thomas Herring" Room 54-820A; 253-5941" tah@mit.edu" http://geoweb.mit.edu/~tah/12.540 " Processing Software" Examine basic features
Methods for Assessor Screening
 Report ITU-R BS.2300-0 (04/2014) Methods for Assessor Screening BS Series Broadcasting service (sound) ii Rep. ITU-R BS.2300-0 Foreword The role of the Radiocommunication Sector is to ensure the rational,
Report ITU-R BS.2300-0 (04/2014) Methods for Assessor Screening BS Series Broadcasting service (sound) ii Rep. ITU-R BS.2300-0 Foreword The role of the Radiocommunication Sector is to ensure the rational,
Using Pedigrees to interpret Mode of Inheritance
 Using Pedigrees to interpret Mode of Inheritance Objectives Use a pedigree to interpret the mode of inheritance the given trait is with 90% accuracy. 11.2 Pedigrees (It s in your genes) Pedigree Charts
Using Pedigrees to interpret Mode of Inheritance Objectives Use a pedigree to interpret the mode of inheritance the given trait is with 90% accuracy. 11.2 Pedigrees (It s in your genes) Pedigree Charts
Analysing Illumina bead-based data using beadarray
 Analysing Illumina bead-based data using beadarray Mark Dunning 6th August 2007 The Bead Each silica bead is 3 microns in diameter 700,000 copies of same probe sequence are covalently attached to each
Analysing Illumina bead-based data using beadarray Mark Dunning 6th August 2007 The Bead Each silica bead is 3 microns in diameter 700,000 copies of same probe sequence are covalently attached to each
Laboratory 1: Uncertainty Analysis
 University of Alabama Department of Physics and Astronomy PH101 / LeClair May 26, 2014 Laboratory 1: Uncertainty Analysis Hypothesis: A statistical analysis including both mean and standard deviation can
University of Alabama Department of Physics and Astronomy PH101 / LeClair May 26, 2014 Laboratory 1: Uncertainty Analysis Hypothesis: A statistical analysis including both mean and standard deviation can
Lecture 1: Introduction to pedigree analysis
 Lecture 1: Introduction to pedigree analysis Magnus Dehli Vigeland NORBIS course, 8 th 12 th of January 2018, Oslo Outline Part I: Brief introductions Pedigrees symbols and terminology Some common relationships
Lecture 1: Introduction to pedigree analysis Magnus Dehli Vigeland NORBIS course, 8 th 12 th of January 2018, Oslo Outline Part I: Brief introductions Pedigrees symbols and terminology Some common relationships
cobindr package vignette
 cobindr package vignette October 30, 2018 Many transcription factors (TFs) regulate gene expression by binding to specific DNA motifs near genes. Often the regulation of gene expression is not only controlled
cobindr package vignette October 30, 2018 Many transcription factors (TFs) regulate gene expression by binding to specific DNA motifs near genes. Often the regulation of gene expression is not only controlled
The permax Package. May 26, 2004
 The permax Package May 26, 2004 Version 1.2.1 Author Robert J. Gray Maintainer Robert Gentleman The permax library consists of 7 functions, intended
The permax Package May 26, 2004 Version 1.2.1 Author Robert J. Gray Maintainer Robert Gentleman The permax library consists of 7 functions, intended
Design of Parallel Algorithms. Communication Algorithms
 + Design of Parallel Algorithms Communication Algorithms + Topic Overview n One-to-All Broadcast and All-to-One Reduction n All-to-All Broadcast and Reduction n All-Reduce and Prefix-Sum Operations n Scatter
+ Design of Parallel Algorithms Communication Algorithms + Topic Overview n One-to-All Broadcast and All-to-One Reduction n All-to-All Broadcast and Reduction n All-Reduce and Prefix-Sum Operations n Scatter
Unsupervised Classification
 Unsupervised Classification Using SAGA Tutorial ID: IGET_RS_007 This tutorial has been developed by BVIEER as part of the IGET web portal intended to provide easy access to geospatial education. This tutorial
Unsupervised Classification Using SAGA Tutorial ID: IGET_RS_007 This tutorial has been developed by BVIEER as part of the IGET web portal intended to provide easy access to geospatial education. This tutorial
Manual for Familias 3
 Manual for Familias 3 Daniel Kling 1 (daniellkling@gmailcom) Petter F Mostad 2 (mostad@chalmersse) ThoreEgeland 1,3 (thoreegeland@nmbuno) 1 Oslo University Hospital Department of Forensic Services Oslo,
Manual for Familias 3 Daniel Kling 1 (daniellkling@gmailcom) Petter F Mostad 2 (mostad@chalmersse) ThoreEgeland 1,3 (thoreegeland@nmbuno) 1 Oslo University Hospital Department of Forensic Services Oslo,
An Introduction to Data Visualization with RStudio November 29, 2018
 Good afternoon everyone my name is Bailey Maryfield and I am one of JRSA's research analysts. For those of you less familiar with JRSA, that stands for the Justice Research and Statistics Association.
Good afternoon everyone my name is Bailey Maryfield and I am one of JRSA's research analysts. For those of you less familiar with JRSA, that stands for the Justice Research and Statistics Association.
Real-time Forecast Combinations for the Oil Price
 Crawford School of Public Policy CAMA Centre for Applied Macroeconomic Analysis Real-time Forecast Combinations for the Oil Price CAMA Working Paper 38/2018 August 2018 Anthony Garratt University of Warwick
Crawford School of Public Policy CAMA Centre for Applied Macroeconomic Analysis Real-time Forecast Combinations for the Oil Price CAMA Working Paper 38/2018 August 2018 Anthony Garratt University of Warwick
Exercise 4 Exploring Population Change without Selection
 Exercise 4 Exploring Population Change without Selection This experiment began with nine Avidian ancestors of identical fitness; the mutation rate is zero percent. Since descendants can never differ in
Exercise 4 Exploring Population Change without Selection This experiment began with nine Avidian ancestors of identical fitness; the mutation rate is zero percent. Since descendants can never differ in
Comparing Across Categories Part of a Series of Tutorials on using Google Sheets to work with data for making charts in Venngage
 Comparing Across Categories Part of a Series of Tutorials on using Google Sheets to work with data for making charts in Venngage These materials are based upon work supported by the National Science Foundation
Comparing Across Categories Part of a Series of Tutorials on using Google Sheets to work with data for making charts in Venngage These materials are based upon work supported by the National Science Foundation
FLDIGI Users Manual: WEFAX
 w1hkj.com 10-13 minutes This modem is able to receive and transmit HF-Fax images, traditionally used for weather reports. More technical information is available on the wikipedia article Radiofax. Two
w1hkj.com 10-13 minutes This modem is able to receive and transmit HF-Fax images, traditionally used for weather reports. More technical information is available on the wikipedia article Radiofax. Two
arxiv: v1 [astro-ph.im] 7 Dec 2010
![arxiv: v1 [astro-ph.im] 7 Dec 2010 arxiv: v1 [astro-ph.im] 7 Dec 2010](/thumbs/88/114537853.jpg) arxiv:1012.1583v1 [astro-ph.im] 7 Dec 2010 University of Amsterdam (UvA), Amsterdam, The Netherlands E-mail: a.alexov@uva.nl Jason W. T. Hessels Netherlands Institute for Radio Astronomy (ASTRON), Dwingeloo,
arxiv:1012.1583v1 [astro-ph.im] 7 Dec 2010 University of Amsterdam (UvA), Amsterdam, The Netherlands E-mail: a.alexov@uva.nl Jason W. T. Hessels Netherlands Institute for Radio Astronomy (ASTRON), Dwingeloo,
LASER server: ancestry tracing with genotypes or sequence reads
 LASER server: ancestry tracing with genotypes or sequence reads The LASER method Supplementary Data For each ancestry reference panel of N individuals, LASER applies principal components analysis (PCA)
LASER server: ancestry tracing with genotypes or sequence reads The LASER method Supplementary Data For each ancestry reference panel of N individuals, LASER applies principal components analysis (PCA)
NADFAS TRAINING DAY Presentation notes for attendees in Scotland
 NADFAS TRAINING DAY Presentation notes for attendees in Scotland CHURCH RECORDING HANDBOOK pp. 37-39, 42, 45-46 & PHOTO SUPPS 5, 12 & 13 Available in Members Section of NADFAS website under Church Recording
NADFAS TRAINING DAY Presentation notes for attendees in Scotland CHURCH RECORDING HANDBOOK pp. 37-39, 42, 45-46 & PHOTO SUPPS 5, 12 & 13 Available in Members Section of NADFAS website under Church Recording
Learning Dota 2 Team Compositions
 Learning Dota 2 Team Compositions Atish Agarwala atisha@stanford.edu Michael Pearce pearcemt@stanford.edu Abstract Dota 2 is a multiplayer online game in which two teams of five players control heroes
Learning Dota 2 Team Compositions Atish Agarwala atisha@stanford.edu Michael Pearce pearcemt@stanford.edu Abstract Dota 2 is a multiplayer online game in which two teams of five players control heroes
Steps toward reproducible research
 Steps toward reproducible research Karl Broman Biostatistics & Medical Informatics Univ. Wisconsin Madison kbroman.org github.com/kbroman @kwbroman Slides: bit.ly/jax2017-05 These are slides for a talk
Steps toward reproducible research Karl Broman Biostatistics & Medical Informatics Univ. Wisconsin Madison kbroman.org github.com/kbroman @kwbroman Slides: bit.ly/jax2017-05 These are slides for a talk
Greedy Algorithms. Study Chapters /4/2014 COMP 555 Bioalgorithms (Fall 2014) 1
 Greedy Algorithms Study Chapters.1-.2 9//201 COMP Bioalgorithms (Fall 201) 1 Which version of Python? Use version 2.7 or 2.6 Python Information Where to run python? On your preferred platform Windows,
Greedy Algorithms Study Chapters.1-.2 9//201 COMP Bioalgorithms (Fall 201) 1 Which version of Python? Use version 2.7 or 2.6 Python Information Where to run python? On your preferred platform Windows,
2. Inference for comparing two proportions
 Unit5: Inferenceforcategoricaldata 2. Inference for comparing two proportions Sta 101 - Spring 2016 Duke University, Department of Statistical Science Dr. Çetinkaya-Rundel Slides posted at http://bit.ly/sta101_s16
Unit5: Inferenceforcategoricaldata 2. Inference for comparing two proportions Sta 101 - Spring 2016 Duke University, Department of Statistical Science Dr. Çetinkaya-Rundel Slides posted at http://bit.ly/sta101_s16
The Microstation configuration must be set up with the correct CAD resource files as outlined in SPEC
 1/7 1.0 PURPOSE This specification describes the procedure for working with the Vale Drawing Checker Utility. This utility checks the drawing for adherence to many mandatory Vale drawing standards and
1/7 1.0 PURPOSE This specification describes the procedure for working with the Vale Drawing Checker Utility. This utility checks the drawing for adherence to many mandatory Vale drawing standards and
A Custom-made MATLAB Based Software to Manage Leakage Current Waveforms
 ETASR - Engineering, Technology & Applied Science Research Vol. 1, No.2, 2011, 36-42 36 A Custom-made MATLAB Based Software to Manage Leakage Current Waveforms Dionisios Pylarinos High Voltage Lab University
ETASR - Engineering, Technology & Applied Science Research Vol. 1, No.2, 2011, 36-42 36 A Custom-made MATLAB Based Software to Manage Leakage Current Waveforms Dionisios Pylarinos High Voltage Lab University
Copy number variations and quantitative trait loci in South African Brahman cattle
 Copy number variations and quantitative trait loci in South African Brahman cattle M.D. Wang 1, F.C. Muchadeyi 2, M. Makgahela 1,3, S. Mdyogolo 1 & A. Maiwashe 1,3 1 Agriculture Research Council-Animal
Copy number variations and quantitative trait loci in South African Brahman cattle M.D. Wang 1, F.C. Muchadeyi 2, M. Makgahela 1,3, S. Mdyogolo 1 & A. Maiwashe 1,3 1 Agriculture Research Council-Animal
)454 ' TELECOMMUNICATION STANDARDIZATION SECTOR OF ITU
 INTERNATIONAL TELECOMMUNICATION UNION )454 ' TELECOMMUNICATION STANDARDIZATION SECTOR OF ITU '%.%2!,!30%#43 /& $)')4!, 42!.3-)33)/. 3934%-3 4%2-).!, %15)0-%.43 4()2$ /2$%2 $)')4!, -5,4)0,%8 %15)0-%.4 /0%2!4).'!4
INTERNATIONAL TELECOMMUNICATION UNION )454 ' TELECOMMUNICATION STANDARDIZATION SECTOR OF ITU '%.%2!,!30%#43 /& $)')4!, 42!.3-)33)/. 3934%-3 4%2-).!, %15)0-%.43 4()2$ /2$%2 $)')4!, -5,4)0,%8 %15)0-%.4 /0%2!4).'!4
Prediction Method of Beef Marbling Standard Number Using Parameters Obtained from Image Analysis for Beef Ribeye
 Prediction Method of Beef Marbling Standard Number Using Parameters Obtained from Image Analysis for Beef Ribeye Keigo KUCHIDA, Shogo TSURUTA1, a, L. D. Van Vleck2, Mitsuyoshi SUZUKI and Shunzo MIYOSHI
Prediction Method of Beef Marbling Standard Number Using Parameters Obtained from Image Analysis for Beef Ribeye Keigo KUCHIDA, Shogo TSURUTA1, a, L. D. Van Vleck2, Mitsuyoshi SUZUKI and Shunzo MIYOSHI
Stratigraphy Modeling Boreholes and Cross. Become familiar with boreholes and borehole cross sections in GMS
 v. 10.3 GMS 10.3 Tutorial Stratigraphy Modeling Boreholes and Cross Sections Become familiar with boreholes and borehole cross sections in GMS Objectives Learn how to import borehole data, construct a
v. 10.3 GMS 10.3 Tutorial Stratigraphy Modeling Boreholes and Cross Sections Become familiar with boreholes and borehole cross sections in GMS Objectives Learn how to import borehole data, construct a
Evolutions of communication
 Evolutions of communication Alex Bell, Andrew Pace, and Raul Santos May 12, 2009 Abstract In this paper a experiment is presented in which two simulated robots evolved a form of communication to allow
Evolutions of communication Alex Bell, Andrew Pace, and Raul Santos May 12, 2009 Abstract In this paper a experiment is presented in which two simulated robots evolved a form of communication to allow
Using the zoom adjustment, zoom on the gel Adjust the tray on the VGAU 3000 to see the image of the gel in the viewfinder
 Operation of Vakili 3000 Gel Analysis Unit Both qualitative and quantitative analysis of electrophoresis experiments can be accomplished by using the Vakili 3000 Gel Analysis Unit. There are three steps
Operation of Vakili 3000 Gel Analysis Unit Both qualitative and quantitative analysis of electrophoresis experiments can be accomplished by using the Vakili 3000 Gel Analysis Unit. There are three steps
I STATISTICAL TOOLS IN SIX SIGMA DMAIC PROCESS WITH MINITAB APPLICATIONS
 Six Sigma Quality Concepts & Cases- Volume I STATISTICAL TOOLS IN SIX SIGMA DMAIC PROCESS WITH MINITAB APPLICATIONS Chapter 7 Measurement System Analysis Gage Repeatability & Reproducibility (Gage R&R)
Six Sigma Quality Concepts & Cases- Volume I STATISTICAL TOOLS IN SIX SIGMA DMAIC PROCESS WITH MINITAB APPLICATIONS Chapter 7 Measurement System Analysis Gage Repeatability & Reproducibility (Gage R&R)
Repeated Measures Twoway Analysis of Variance
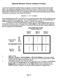 Repeated Measures Twoway Analysis of Variance A researcher was interested in whether frequency of exposure to a picture of an ugly or attractive person would influence one's liking for the photograph.
Repeated Measures Twoway Analysis of Variance A researcher was interested in whether frequency of exposure to a picture of an ugly or attractive person would influence one's liking for the photograph.
Data Processing: Visibility Calibration
 Data Processing: Visibility Calibration The delivered ALMA data consist of the amplitudes and phases for the combined signals from pairs of antennas. These are called visibility data. The goal of visibility
Data Processing: Visibility Calibration The delivered ALMA data consist of the amplitudes and phases for the combined signals from pairs of antennas. These are called visibility data. The goal of visibility
CMSC 372: Artificial Intelligence Lab#1: Designing Pac-Man Agents
 CMSC 372: Artificial Intelligence Lab#1: Designing Pac-Man Agents Figure 1: The Pac-Man World Introduction In this project, you will familiarize yourself with the Pac-Man World. Over the next few assignments
CMSC 372: Artificial Intelligence Lab#1: Designing Pac-Man Agents Figure 1: The Pac-Man World Introduction In this project, you will familiarize yourself with the Pac-Man World. Over the next few assignments
Stratigraphy Modeling Boreholes and Cross Sections
 GMS TUTORIALS Stratigraphy Modeling Boreholes and Cross Sections The Borehole module of GMS can be used to visualize boreholes created from drilling logs. Also three-dimensional cross sections between
GMS TUTORIALS Stratigraphy Modeling Boreholes and Cross Sections The Borehole module of GMS can be used to visualize boreholes created from drilling logs. Also three-dimensional cross sections between
UNIVERSITETET FOR MILJØ- OG BIOVITSKAP
 UNIVERSITETET FOR MILJØ- OG BIOVITSKAP 1 Photo: Ingunn Nævdal http://www.nsg.no/ind ex.cfm?id= 53192 MILK QUALITY BREEDING VALUE PREDICTION BASED ON FTIR SPECTRA Tormod ÅDNØY, Theo ME MEUWISSEN, Binyamin
UNIVERSITETET FOR MILJØ- OG BIOVITSKAP 1 Photo: Ingunn Nævdal http://www.nsg.no/ind ex.cfm?id= 53192 MILK QUALITY BREEDING VALUE PREDICTION BASED ON FTIR SPECTRA Tormod ÅDNØY, Theo ME MEUWISSEN, Binyamin
xdev Magazine Markup Guide
 xdev Magazine Markup Guide How to use Markdown to format articles July 12, 2013 v1.2 xdev Magazine Markup Guide 2 Contents Introduction.............................. 3 The Goals of Our Formatting System..............
xdev Magazine Markup Guide How to use Markdown to format articles July 12, 2013 v1.2 xdev Magazine Markup Guide 2 Contents Introduction.............................. 3 The Goals of Our Formatting System..............
CCD Image Processing of M15 Images Estimated time: 4 hours
 CCD Image Processing of M15 Images Estimated time: 4 hours For this part of the astronomy lab, you will use the astronomy software package IRAF (Image Reduction and Analysis Facility) to perform the basic
CCD Image Processing of M15 Images Estimated time: 4 hours For this part of the astronomy lab, you will use the astronomy software package IRAF (Image Reduction and Analysis Facility) to perform the basic
Diversity Image Inspector
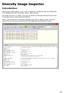 Diversity Image Inspector Introduction The Diversity Image Inspector scans a bulk of images for included barcodes and configurable EXIF metadata (e.g. GPS coordinates, author, date and time). The results
Diversity Image Inspector Introduction The Diversity Image Inspector scans a bulk of images for included barcodes and configurable EXIF metadata (e.g. GPS coordinates, author, date and time). The results
Advanced Data Analysis Pattern Recognition & Neural Networks Software for Acoustic Emission Applications. Topic: Waveforms in Noesis
 Advanced Data Analysis Pattern Recognition & Neural Networks Software for Acoustic Emission Applications Topic: Waveforms in Noesis 1 Noesis Waveforms Capabilities Noesis main features relating to Waveforms:
Advanced Data Analysis Pattern Recognition & Neural Networks Software for Acoustic Emission Applications Topic: Waveforms in Noesis 1 Noesis Waveforms Capabilities Noesis main features relating to Waveforms:
-f/d-b '') o, q&r{laniels, Advisor. 20rt. lmage Processing of Petrographic and SEM lmages. By James Gonsiewski. The Ohio State University
 lmage Processing of Petrographic and SEM lmages Senior Thesis Submitted in partial fulfillment of the requirements for the Bachelor of Science Degree At The Ohio State Universitv By By James Gonsiewski
lmage Processing of Petrographic and SEM lmages Senior Thesis Submitted in partial fulfillment of the requirements for the Bachelor of Science Degree At The Ohio State Universitv By By James Gonsiewski
Basic Use of XCMS -- Local. Xiuxia Du Department of Bioinformatics & Genomics University of North Carolina at Charlotte
 Basic Use of XCMS -- Local Xiuxia Du Department of Bioinformatics & Genomics University of North Carolina at Charlotte Preparation Required: install R Optional: install Rstudio, an IDE (Integrated Development
Basic Use of XCMS -- Local Xiuxia Du Department of Bioinformatics & Genomics University of North Carolina at Charlotte Preparation Required: install R Optional: install Rstudio, an IDE (Integrated Development
Package pedantics. R topics documented: April 18, Type Package
 Type Package Package pedantics April 18, 2018 Title Functions to Facilitate Power and Sensitivity Analyses for Genetic Studies of Natural Populations Version 1.7 Date 2018-04-18 Depends R (>= 2.4.0), MasterBayes,
Type Package Package pedantics April 18, 2018 Title Functions to Facilitate Power and Sensitivity Analyses for Genetic Studies of Natural Populations Version 1.7 Date 2018-04-18 Depends R (>= 2.4.0), MasterBayes,
Information system for a wheat breeding program
 Information system for a wheat breeding program L. LÁNG L Cs. KUTI Z. BEDŐ Agricultural Research Institute of the Hungarian Academy of Sciences Martonvásár The information system has to satisfy the demand
Information system for a wheat breeding program L. LÁNG L Cs. KUTI Z. BEDŐ Agricultural Research Institute of the Hungarian Academy of Sciences Martonvásár The information system has to satisfy the demand
Walter Steets Houston Genealogical Forum DNA Interest Group April 7, 2018
 Ancestry DNA and GEDmatch Walter Steets Houston Genealogical Forum DNA Interest Group April 7, 2018 Today s agenda Recent News about DNA Testing DNA Cautions: DNA Data Used for Forensic Purposes New Technology:
Ancestry DNA and GEDmatch Walter Steets Houston Genealogical Forum DNA Interest Group April 7, 2018 Today s agenda Recent News about DNA Testing DNA Cautions: DNA Data Used for Forensic Purposes New Technology:
Constructing Genetic Linkage Maps with MAPMAKER/EXP Version 3.0: A Tutorial and Reference Manual
 Whitehead Institute Constructing Genetic Linkage Maps with MAPMAKER/EXP Version 3.0: A Tutorial and Reference Manual Stephen E. Lincoln, Mark J. Daly, and Eric S. Lander A Whitehead Institute for Biomedical
Whitehead Institute Constructing Genetic Linkage Maps with MAPMAKER/EXP Version 3.0: A Tutorial and Reference Manual Stephen E. Lincoln, Mark J. Daly, and Eric S. Lander A Whitehead Institute for Biomedical
TIBCO FTL Part of the TIBCO Messaging Suite. Quick Start Guide
 TIBCO FTL 6.0.0 Part of the TIBCO Messaging Suite Quick Start Guide The TIBCO Messaging Suite TIBCO FTL is part of the TIBCO Messaging Suite. It includes not only TIBCO FTL, but also TIBCO eftl (providing
TIBCO FTL 6.0.0 Part of the TIBCO Messaging Suite Quick Start Guide The TIBCO Messaging Suite TIBCO FTL is part of the TIBCO Messaging Suite. It includes not only TIBCO FTL, but also TIBCO eftl (providing
SUPPLEMENT TO THE PAPER TESTING EQUALITY OF SPECTRAL DENSITIES USING RANDOMIZATION TECHNIQUES
 SUPPLEMENT TO THE PAPER TESTING EQUALITY OF SPECTRAL DENSITIES USING RANDOMIZATION TECHNIQUES CARSTEN JENTSCH AND MARKUS PAULY Abstract. In this supplementary material we provide additional supporting
SUPPLEMENT TO THE PAPER TESTING EQUALITY OF SPECTRAL DENSITIES USING RANDOMIZATION TECHNIQUES CARSTEN JENTSCH AND MARKUS PAULY Abstract. In this supplementary material we provide additional supporting
Biology 559R: Introduction to Phylogenetic Comparative Methods Topics for this week (Feb 3 & 5):
 Biology 559R: Introduction to Phylogenetic Comparative Methods Topics for this week (Feb 3 & 5): Chronogram estimation: Penalized Likelihood Approach BEAST Presentations of your projects 1 The Anatomy
Biology 559R: Introduction to Phylogenetic Comparative Methods Topics for this week (Feb 3 & 5): Chronogram estimation: Penalized Likelihood Approach BEAST Presentations of your projects 1 The Anatomy
Locating Molecules Using GSD Technology Project Folders: Organization of Experiment Files...1
 .....................................1 1 Project Folders: Organization of Experiment Files.................................1 2 Steps........................................................................2
.....................................1 1 Project Folders: Organization of Experiment Files.................................1 2 Steps........................................................................2
I STATISTICAL TOOLS IN SIX SIGMA DMAIC PROCESS WITH MINITAB APPLICATIONS
 Six Sigma Quality Concepts & Cases- Volume I STATISTICAL TOOLS IN SIX SIGMA DMAIC PROCESS WITH MINITAB APPLICATIONS Chapter 7 Measurement System Analysis Gage Repeatability & Reproducibility (Gage R&R)
Six Sigma Quality Concepts & Cases- Volume I STATISTICAL TOOLS IN SIX SIGMA DMAIC PROCESS WITH MINITAB APPLICATIONS Chapter 7 Measurement System Analysis Gage Repeatability & Reproducibility (Gage R&R)
Homework Assignment (20 points): MORPHOMETRICS (Bivariate and Multivariate Analyses)
 Fossils and Evolution Due: Tuesday, Jan. 31 Spring 2012 Homework Assignment (20 points): MORPHOMETRICS (Bivariate and Multivariate Analyses) Introduction Morphometrics is the use of measurements to assess
Fossils and Evolution Due: Tuesday, Jan. 31 Spring 2012 Homework Assignment (20 points): MORPHOMETRICS (Bivariate and Multivariate Analyses) Introduction Morphometrics is the use of measurements to assess
The method requires foreground and background sequence datasets. The users can use fasta files as input.
 1 Introduction he emergence of hip-seq technology for genome-wide profiling of transcription factor binding sites (FBS) has made it possible to categorize very precisely the FBS motifs. How to harness
1 Introduction he emergence of hip-seq technology for genome-wide profiling of transcription factor binding sites (FBS) has made it possible to categorize very precisely the FBS motifs. How to harness
Permutation inference for the General Linear Model
 Permutation inference for the General Linear Model Anderson M. Winkler fmrib Analysis Group 3.Sep.25 Winkler Permutation for the glm / 63 in jalapeno: winkler/bin/palm Winkler Permutation for the glm 2
Permutation inference for the General Linear Model Anderson M. Winkler fmrib Analysis Group 3.Sep.25 Winkler Permutation for the glm / 63 in jalapeno: winkler/bin/palm Winkler Permutation for the glm 2
Attaching & detaching
 The Attach tool has two functions. Attach holds your cuts in position so that images on the cutting mat will appear exactly as they show up on the design canvas. Attach can also fasten a write or score
The Attach tool has two functions. Attach holds your cuts in position so that images on the cutting mat will appear exactly as they show up on the design canvas. Attach can also fasten a write or score
Office 2016 Excel Basics 24 Video/Class Project #36 Excel Basics 24: Visualize Quantitative Data with Excel Charts. No Chart Junk!!!
 Office 2016 Excel Basics 24 Video/Class Project #36 Excel Basics 24: Visualize Quantitative Data with Excel Charts. No Chart Junk!!! Goal in video # 24: Learn about how to Visualize Quantitative Data with
Office 2016 Excel Basics 24 Video/Class Project #36 Excel Basics 24: Visualize Quantitative Data with Excel Charts. No Chart Junk!!! Goal in video # 24: Learn about how to Visualize Quantitative Data with
A general quadratic programming method for the optimisation of genetic contributions using interior point algorithm. R Pong-Wong & JA Woolliams
 A general quadratic programming method for the optimisation of genetic contributions using interior point algorithm R Pong-Wong & JA Woolliams Introduction Inbreeding is a risk and it needs to be controlled
A general quadratic programming method for the optimisation of genetic contributions using interior point algorithm R Pong-Wong & JA Woolliams Introduction Inbreeding is a risk and it needs to be controlled
Reading 14 : Counting
 CS/Math 240: Introduction to Discrete Mathematics Fall 2015 Instructors: Beck Hasti, Gautam Prakriya Reading 14 : Counting In this reading we discuss counting. Often, we are interested in the cardinality
CS/Math 240: Introduction to Discrete Mathematics Fall 2015 Instructors: Beck Hasti, Gautam Prakriya Reading 14 : Counting In this reading we discuss counting. Often, we are interested in the cardinality
Permutation and Randomization Tests 1
 Permutation and 1 STA442/2101 Fall 2012 1 See last slide for copyright information. 1 / 19 Overview 1 Permutation Tests 2 2 / 19 The lady and the tea From Fisher s The design of experiments, first published
Permutation and 1 STA442/2101 Fall 2012 1 See last slide for copyright information. 1 / 19 Overview 1 Permutation Tests 2 2 / 19 The lady and the tea From Fisher s The design of experiments, first published
Interleaved PC-OFDM to reduce the peak-to-average power ratio
 1 Interleaved PC-OFDM to reduce the peak-to-average power ratio A D S Jayalath and C Tellambura School of Computer Science and Software Engineering Monash University, Clayton, VIC, 3800 e-mail:jayalath@cssemonasheduau
1 Interleaved PC-OFDM to reduce the peak-to-average power ratio A D S Jayalath and C Tellambura School of Computer Science and Software Engineering Monash University, Clayton, VIC, 3800 e-mail:jayalath@cssemonasheduau
