Algorithms for Bioinformatics
|
|
|
- Marvin Gibbs
- 5 years ago
- Views:
Transcription
1 Adapted from slides by Alexandru Tomescu, Leena Salmela, Veli Mäkinen, Esa Pitkänen Algorithms for Bioinformatics Lecture 3: Greedy Algorithms and Genomic Rearrangements
2 Background We now have genomes of several species available It is possible to compare genomes of two or more different species = Comparative genomics Basic observation: Closely related species (such as human and mouse) can be almost identical in terms of genome contents but the order of genomic segments can be very different between species 2 / 51
3 Synteny blocks and segments Synteny describes how genomic segments are located on the same chromosome or close to each other Genes, markers (any sequence) Shared synteny between two species: genes are located close to each other in both of the species Synteny block (or syntenic block) A set of genes or markers that co-occur together in two species Synteny segment (or syntenic segment) Syntenic block where the order of genes or markers is preserved 3 / 51
4 Synteny blocks and segments Chromosome i, species B Synteny segment Synteny block Chromosome j, species C Homologs of the same gene 4 / 51
5 Chromosomes Linear chromosomes Eukaryotes (mostly) Circular chromosomes Prokaryotes (mostly) Mitochondria Chromosomes are double stranded: genes can be found on both strands (orientations) gene 2 gene 1 gene 3 5 / 51
6 Example: Human vs mouse genome Human and mouse genomes share thousands of homologous genes but they are Arranged in different order Located in different chromosomes Examples: Human chromosome 6 contains elements from six different mouse chromosomes Analysis of X chromosome indicates that rearrangements have happened primarily within chromosome 6 / 51
7 Jones & Pevzner, / 51
8 Representing genomic rearrangements When comparing genomes, we can find homologous sequences in both using sequence comparison algorithms (next lecture). This gives us a map between sequences in both genomes. 8 / 51
9 Representing genomic rearrangements We assign numbers 1,..., n to the found homologous sequences By convention, we number the sequences in the first genome by their order of appearance in the chromosomes If the homolog of i is in reverse orientation, it receives number i (signed data) For example consider human vs mouse gene numbering on the right List order corresponds to physical order on chromosomes! Human Mouse 1 (gnat2) 12 (inpp1) 2 (nras) 13 (cd28) 3 (ngfb) 14 (fn1) 4 (gba) 15 (pax3) 5 (pklr) -9 (il10) 6 (at3) -8 (pdc) 7 (lamc1) -7 (lamc1) 8 (pdc) -6 (at3) 9 (il10) / 51
10 Permutations The basic data structure in the study of genome rearrangements is permutation A permutation of a sequence of n numbers is a reordering of the sequence For example, is a permutation of / 51
11 Genome rearrangement problem Given two genomes (set of markers), how many duplications, inversions and translocations do we need to transform the first genome to the second? Minimum number of operations? What operations? Which order? 11 / 51
12 Genome rearrangement problem # duplications? # inversions? # translocations? / 51
13 Genome rearrangement problem π 1 π 2 π 3 π 4 π 5 Permutation of 1,..., Keep in mind that the two genomes have been evolved from a common ancestor genome! 13 / 51
14 Genome rearrangements using reversals (inversions) only Let s consider a simpler problem where we just study reversals with unsigned data A reversal p(i, j) reverses the order of the segment π i π i+1... π j 1 π j (indexing starts from 1) For example, given permutation and reversal ρ(2, 4) we get permutation Note that we do not care about the exact positions on the genome. 14 / 51
15 Sorting by Reversals problem Goal: Find the shortest series of reversals that tranforms a given permutation to the identity permutation Input: Permutation π of the numbers 1,..., n Output: A series of reversals ρ 1,..., ρ t that transforms π into (1, 2,..., n). Objective: Minimize t. The smallest possible value of t is called the reversal distance and is denoted by d(π). Reversal distance for a pair of chromosomes: Find synteny blocks in both Number synteny blocks in the first chromosome to identity Set π to corresponding matching of second chromosome s blocks against the first 15 / 51
16 A simple reversal sort Our first approach to solve the sorting by reversals problem resembles selection sort: Examine each position i of the permutation from left to right At each position, if πi i, do a reversal such that π i = i This is a greedy approach: we try to choose the option that looks best at the current step. It finds a solution that is valid but often not optimal. 16 / 51
17 Simple reversal sort: example = = = = Reversal series: ρ(1, 2), ρ(2, 3), ρ(3, 4), ρ(4, 5) Is d( ) then 4? 17 / 51
18 Simple reversal sort: example = = = = Reversal series: ρ(1, 2), ρ(2, 3), ρ(3, 4), ρ(4, 5) Is d( ) then 4? = = d( ) = 2 17 / 51
19 How good is simple reversal sort? Not so good actually It has to do at most n 1 reversals with permutation of length n In our previous example, the algorithm returned a solution that is as large as (n 1)/2 times the optimal result d(π) = 2 For example, if we extend the example for n = 1001, the result can be as bad as 500 d(π) 18 / 51
20 Approximation algorithms and approximation ratios Simple reversal sort is an approximation algorithm. It only produces an approximate solution. A(π): approximate solution returned by algorithm A OPT (π): optimal solution The approximation ratio of (minimization) algorithm A is the maximum approximation ratio over all inputs of size n: max π =n A(π) OPT (π) The approximation ratio for simple reversal sort is thus at least (n 1)/2 The approximation ratio tells how much off the solution given by the algorithm can in worst case be from the optimal solution 19 / 51
21 Approximation ratios for maximization problems Previous slide gave the approximation ratio for a minimization problem like reversal distance. For a maximization problem (e.g. motif finding, maximizing score) the approximation ratio of an algorithm is defined as the minimum approximation ratio over all inputs of size n: min π =n A(π) OPT (π) 20 / 51
22 Computing reversals with breakpoints Let s investigate a better way to sort by reversals First some concepts related to permutation π 1 π 2... π n 1 π n Breakpoint: two elements π i and π i+1 are a breakpoint if they are not consecutive numbers Adjacency: if πi and π i+1 are consecutive they are an adjacency 21 / 51
23 Breakpoints and adjacencies The permutation contains three breakpoints: begin-5, 31, 26 six adjacencies: 54, 43, 12, 67, 78, 8-end Breakpoints 22 / 51
24 Breakpoints Each breakpoint in permutation needs to be removed to get to the identity permutation (= our target) Identity permutation does not contain any breakpoints First and last positions special cases Note that each reversal can remove at most two breakpoints Denote the number of breakpoints by b(π) b(π) = 3 23 / 51
25 Breakpoint reversal sort Idea: Try to remove as many breakpoints as possible (max 2) in every step 1: while b(π) > 0 do 2: Choose reversal ρ that removes most breakpoints 3: Perform reversal ρ to π 4: Output π 5: return 24 / 51
26 Breakpoint removal: example b(π) = b(π) = b(π) = b(π) = b(π) = 0 25 / 51
27 Breakpoint removal The previous algorithm needs refinement to be correct Consider the following permutation There is no reversal that decreases the number of breakpoints! 26 / 51
28 Breakpoint removal Reversal can always decrease breakpoint count if permutation contains decreasing strips Strip: maximal segment without breakpoints Increasing strip Decreasing strip (including segments of length 1, except 1 and n if they are located at their correct locations) 27 / 51
29 Improved breakpoint reversal sort 1: while b(π) > 0 do 2: if π has a decreasing strip then 3: Apply reversal ρ such that it removes most BPs 4: else 5: Reverse an increasing strip 6: Output π 28 / 51
30 Is improved BP removal enough? The algorithm works pretty well: A reversal removes at most two breakpoints = Optimal solution cannot be better than b(π)/2 Improved BP removal performs at most 2 b(π) reversals = The result is at most four times worse than the optimal = The approximation ratio of improved BP removal is at most 4. Is this good? We considered only reversals What about translocations? 29 / 51
31 Translocations via reversals Translocation of 2,3,4 ρ(2, 8) ρ(2, 4) ρ(5, 8) 30 / 51
32 Genome rearrangements with reversals With unsigned data, the problem of sorting by reversal is NP-complete An algorithm has been developed that achieves approximation (Berman et al. ESA 2002) However, the reversal distance in signed data can be computed quickly! It takes linear time w.r.t. the length of the permutation (Bader, Moret, Yan 2001) We will not cover that algorithm here but give some insight into central concepts leading to it 31 / 51
33 Cycle decomposition Let s represent permutation π = with the following graph where edges correspond to adjacencies (identity, permutation π) 32 / 51
34 Cycle decomposition Cycle decomposition: a set of cycles that have edges with alternating colors do not share edges with other cycles (=cycles are edge disjoint) every edge belongs to some cycle / 51
35 Estimating reversal distance by cycle decomposition Let c(π) be the maximum number of cycles in a cycle decomposition of π The following formula allows estimation of d(π) d(π) n + 1 c(π), where n is the permutation length d(π) = 2 34 / 51
36 Cycle decompositions Cycle decomposition is NP-complete for unsigned permutations However, with signed data cycle decomposition becomes a trivial task (the cycles are vertex disjoint) 35 / 51
37 Cycle decomposition with signed data Consider the following permutation that includes orientation of the markers We modify this representation to include both endpoints of each marker: 0 1a 1b 5b 5a 3b 3a 2b 2a 4a 4b 6 36 / 51
38 Graph representation of π and identity permutation 0 1a 1b 5b 5a 3b 3a 2b 2a 4a 4b 6 d(π) n + 1 c(π) = = 3 37 / 51
39 Reversal step 1 (ad hoc greedy algorithm) 0 1a 1b 5b 5a 3b 3a 2b 2a 4a 4b Step 1 0 1a 1b 2a 2b 3a 3b 5a 5b 4a 4b / 51
40 Reversal steps 2,3,4 0 1a 1b 2a 2b 3a 3b 5a 5b 4a 4b Step 2 0 1a 1b 2a 2b 3a 3b 4b 4a 5b 5a Step 3,4 0 1a 1b 2a 2b 3a 3b 4a 4b 5a 5b d(π) 4 39 / 51
41 Reversal distance with signed data However, the exact reversal distance in signed data can be computed quickly! It takes linear time w.r.t. the length of permutation (Bader, Moret, Yan 2001) The algorithm is quite involved 40 / 51
42 Multiple chromosomes In unichromosomal genomes, inversion (reversal) is the most common operation In multichromosomal genomes, inversions, translocations, fissions and fusions are most common 41 / 51
43 Fusions and fissions Fusion: merging of two chromosomes Fission: chromosome is split into two chromosomes Both events can be represented with a translocation 42 / 51
44 Fusion Fusion 43 / 51
45 Fission Fission 44 / 51
46 Algorithms for general genomic distance problem Hannenhalli, Pevzner: Transforming Men into Mice (polynomial algorithm for genomic distance problem), 36th Annual IEEE Symposium on Foundations of Computer Science, / 51
47 Human and mouse revisited Human and mouse are separated by about million years of evolutionary history Only a few hundred rearrangements have happened after speciation from the common ancestor Pevzner and Tesler identified in 2003 for 281 synteny blocks a rearrangement from mouse to human with 149 inversions 93 translocations 9 fissions 46 / 51
48 Discussion Genome rearrangement events are very rare compared to e.g. point mutations We can study rearrangements events further back in the evolutionary history Rearrangements are easier to detect in comparison to many other genomic events We cannot detect homologs 100% correctly so the input permutation can contain errors 47 / 51
49 Outline Biological background Permutations and genomic rearrangements Sorting by reversals Simple reversal sort Breakpoints Cycle decomposition Multiple chromosomes Study group assignments 48 / 51
50 Study Group 1: (random allocation at lecture) Read pages from Sung: Algorithms in Bioinformatics: A Practical Introduction, CRC Press approximation for sorting an unsigned permutation Copies distributed at the lecture. In the study group Go through the reasoning in the proof of Lemma 9.2. Simulate the 2-approximation algorithm on the permutation How many reversals does the 2-approximation algorithm need? Is this optimal? 49 / 51
51 Study Group 2: (if you did not get material at the lecture) Read pages 136 and 137 from Jones & Pevzner Greedy approach to motif finding At study group, solve Problem 5.18 Desing an input for the GreedyMotifSearch algorithm that causes the algorithm to output an incorrect result 50 / 51
52 Study Group 3: (random allocation at lecture) Read pages 15, 16, (sect. 2.3) from Vazirani: Approximation algorithms, Springer 2001 Shortest superstring and its greedy approximation through set cover Copies distributed at the lecture. At the study group: present the reduction to set cover with some example go through the proof of Lemma / 51
How good is simple reversal sort? Cycle decompositions. Cycle decompositions. Estimating reversal distance by cycle decomposition
 How good is simple reversal sort? p Not so good actually p It has to do at most n-1 reversals with permutation of length n p The algorithm can return a distance that is as large as (n 1)/2 times the correct
How good is simple reversal sort? p Not so good actually p It has to do at most n-1 reversals with permutation of length n p The algorithm can return a distance that is as large as (n 1)/2 times the correct
Transforming Cabbage into Turnip Genome Rearrangements Sorting By Reversals Greedy Algorithm for Sorting by Reversals Pancake Flipping Problem
 Transforming Cabbage into Turnip Genome Rearrangements Sorting By Reversals Greedy Algorithm for Sorting by Reversals Pancake Flipping Problem Approximation Algorithms Breakpoints: a Different Face of
Transforming Cabbage into Turnip Genome Rearrangements Sorting By Reversals Greedy Algorithm for Sorting by Reversals Pancake Flipping Problem Approximation Algorithms Breakpoints: a Different Face of
GENOMIC REARRANGEMENT ALGORITHMS
 GENOMIC REARRANGEMENT ALGORITHMS KAREN LOSTRITTO Abstract. In this paper, I discuss genomic rearrangement. Specifically, I describe the formal representation of these genomic rearrangements as well as
GENOMIC REARRANGEMENT ALGORITHMS KAREN LOSTRITTO Abstract. In this paper, I discuss genomic rearrangement. Specifically, I describe the formal representation of these genomic rearrangements as well as
Greedy Algorithms and Genome Rearrangements
 Greedy Algorithms and Genome Rearrangements 1. Transforming Cabbage into Turnip 2. Genome Rearrangements 3. Sorting By Reversals 4. Pancake Flipping Problem 5. Greedy Algorithm for Sorting by Reversals
Greedy Algorithms and Genome Rearrangements 1. Transforming Cabbage into Turnip 2. Genome Rearrangements 3. Sorting By Reversals 4. Pancake Flipping Problem 5. Greedy Algorithm for Sorting by Reversals
Greedy Algorithms and Genome Rearrangements
 Greedy Algorithms and Genome Rearrangements Outline 1. Transforming Cabbage into Turnip 2. Genome Rearrangements 3. Sorting By Reversals 4. Pancake Flipping Problem 5. Greedy Algorithm for Sorting by Reversals
Greedy Algorithms and Genome Rearrangements Outline 1. Transforming Cabbage into Turnip 2. Genome Rearrangements 3. Sorting By Reversals 4. Pancake Flipping Problem 5. Greedy Algorithm for Sorting by Reversals
baobabluna: the solution space of sorting by reversals Documentation Marília D. V. Braga
 baobabluna: the solution space of sorting by reversals Documentation Marília D. V. Braga March 15, 2009 II Acknowledgments This work was funded by the European Union Programme Alβan (scholarship no. E05D053131BR),
baobabluna: the solution space of sorting by reversals Documentation Marília D. V. Braga March 15, 2009 II Acknowledgments This work was funded by the European Union Programme Alβan (scholarship no. E05D053131BR),
A Genetic Approach with a Simple Fitness Function for Sorting Unsigned Permutations by Reversals
 A Genetic Approach with a Simple Fitness Function for Sorting Unsigned Permutations by Reversals José Luis Soncco Álvarez Department of Computer Science University of Brasilia Brasilia, D.F., Brazil Email:
A Genetic Approach with a Simple Fitness Function for Sorting Unsigned Permutations by Reversals José Luis Soncco Álvarez Department of Computer Science University of Brasilia Brasilia, D.F., Brazil Email:
Greedy Algorithms. Study Chapters /4/2014 COMP 555 Bioalgorithms (Fall 2014) 1
 Greedy Algorithms Study Chapters.1-.2 9//201 COMP Bioalgorithms (Fall 201) 1 Which version of Python? Use version 2.7 or 2.6 Python Information Where to run python? On your preferred platform Windows,
Greedy Algorithms Study Chapters.1-.2 9//201 COMP Bioalgorithms (Fall 201) 1 Which version of Python? Use version 2.7 or 2.6 Python Information Where to run python? On your preferred platform Windows,
SORTING BY REVERSALS. based on chapter 7 of Setubal, Meidanis: Introduction to Computational molecular biology
 SORTING BY REVERSALS based on chapter 7 of Setubal, Meidanis: Introduction to Computational molecular biology Motivation When comparing genomes across species insertions, deletions and substitutions of
SORTING BY REVERSALS based on chapter 7 of Setubal, Meidanis: Introduction to Computational molecular biology Motivation When comparing genomes across species insertions, deletions and substitutions of
A 2-Approximation Algorithm for Sorting by Prefix Reversals
 A 2-Approximation Algorithm for Sorting by Prefix Reversals c Springer-Verlag Johannes Fischer and Simon W. Ginzinger LFE Bioinformatik und Praktische Informatik Ludwig-Maximilians-Universität München
A 2-Approximation Algorithm for Sorting by Prefix Reversals c Springer-Verlag Johannes Fischer and Simon W. Ginzinger LFE Bioinformatik und Praktische Informatik Ludwig-Maximilians-Universität München
Genome Rearrangements - Continued
 Genome Rearrangements - Continued 1 A Greedy Algorithm for Sorting by Reversals Π = 1, 2, 3, 6, 4, 5 When sorting the permutation,, one notices that the first three elements are already in order. So it
Genome Rearrangements - Continued 1 A Greedy Algorithm for Sorting by Reversals Π = 1, 2, 3, 6, 4, 5 When sorting the permutation,, one notices that the first three elements are already in order. So it
A Approximation Algorithm for Sorting by Transpositions
 A 1.375-Approximation Algorithm for Sorting by Transpositions Isaac Elias 1 and Tzvika Hartman 2 1 Dept. of Numerical Analysis and Computer Science, Royal Institute of Technology, Stockholm, Sweden. isaac@nada.kth.se.
A 1.375-Approximation Algorithm for Sorting by Transpositions Isaac Elias 1 and Tzvika Hartman 2 1 Dept. of Numerical Analysis and Computer Science, Royal Institute of Technology, Stockholm, Sweden. isaac@nada.kth.se.
A Simpler and Faster 1.5-Approximation Algorithm for Sorting by Transpositions
 A Simpler and Faster 1.5-Approximation Algorithm for Sorting by Transpositions Tzvika Hartman Ron Shamir January 15, 2004 Abstract An important problem in genome rearrangements is sorting permutations
A Simpler and Faster 1.5-Approximation Algorithm for Sorting by Transpositions Tzvika Hartman Ron Shamir January 15, 2004 Abstract An important problem in genome rearrangements is sorting permutations
Exploiting the disjoint cycle decomposition in genome rearrangements
 Exploiting the disjoint cycle decomposition in genome rearrangements Jean-Paul Doignon Anthony Labarre 1 doignon@ulb.ac.be alabarre@ulb.ac.be Université Libre de Bruxelles June 7th, 2007 Ordinal and Symbolic
Exploiting the disjoint cycle decomposition in genome rearrangements Jean-Paul Doignon Anthony Labarre 1 doignon@ulb.ac.be alabarre@ulb.ac.be Université Libre de Bruxelles June 7th, 2007 Ordinal and Symbolic
Bounds for Cut-and-Paste Sorting of Permutations
 Bounds for Cut-and-Paste Sorting of Permutations Daniel Cranston Hal Sudborough Douglas B. West March 3, 2005 Abstract We consider the problem of determining the maximum number of moves required to sort
Bounds for Cut-and-Paste Sorting of Permutations Daniel Cranston Hal Sudborough Douglas B. West March 3, 2005 Abstract We consider the problem of determining the maximum number of moves required to sort
Parallel Algorithm to Enumerate Sorting Reversals for Signed Permutation
 Parallel Algorithm to Enumerate Sorting Reversals for Signed Permutation Amit Kumar Das and Amritanjali Dept. Of Computer Science and Engineering Birla Institute of Technology Mesra, Ranchi-835215,India
Parallel Algorithm to Enumerate Sorting Reversals for Signed Permutation Amit Kumar Das and Amritanjali Dept. Of Computer Science and Engineering Birla Institute of Technology Mesra, Ranchi-835215,India
Mathematical Representations of Ciliate Genome Decryption
 Mathematical Representations of Ciliate Genome Decryption Gustavus Adolphus College February 28, 2013 Ciliates Ciliates Single-celled Ciliates Single-celled Characterized by cilia Ciliates Single-celled
Mathematical Representations of Ciliate Genome Decryption Gustavus Adolphus College February 28, 2013 Ciliates Ciliates Single-celled Ciliates Single-celled Characterized by cilia Ciliates Single-celled
A New Tight Upper Bound on the Transposition Distance
 A New Tight Upper Bound on the Transposition Distance Anthony Labarre Université Libre de Bruxelles, Département de Mathématique, CP 16, Service de Géométrie, Combinatoire et Théorie des Groupes, Boulevard
A New Tight Upper Bound on the Transposition Distance Anthony Labarre Université Libre de Bruxelles, Département de Mathématique, CP 16, Service de Géométrie, Combinatoire et Théorie des Groupes, Boulevard
Perfect sorting by reversals is not always difficult 1
 Perfect sorting by reversals is not always difficult 1 S. Bérard and A. Bergeron and C. Chauve and C. Paul Juin 2005 Rapport de Recherche LIRMM RR-05042 161, rue Ada - F. 34394 Montpellier cedex 5 - Tél.
Perfect sorting by reversals is not always difficult 1 S. Bérard and A. Bergeron and C. Chauve and C. Paul Juin 2005 Rapport de Recherche LIRMM RR-05042 161, rue Ada - F. 34394 Montpellier cedex 5 - Tél.
Enumeration of Two Particular Sets of Minimal Permutations
 3 47 6 3 Journal of Integer Sequences, Vol. 8 (05), Article 5.0. Enumeration of Two Particular Sets of Minimal Permutations Stefano Bilotta, Elisabetta Grazzini, and Elisa Pergola Dipartimento di Matematica
3 47 6 3 Journal of Integer Sequences, Vol. 8 (05), Article 5.0. Enumeration of Two Particular Sets of Minimal Permutations Stefano Bilotta, Elisabetta Grazzini, and Elisa Pergola Dipartimento di Matematica
Sampling solution traces for the problem of sorting permutations by signed reversals
 Baudet et al. Algorithms for Molecular Biology 2012, 7:18 RESEARCH Open Access Sampling solution traces for the problem of sorting permutations by signed reversals Christian Baudet 1,2*,ZanoniDias 3 and
Baudet et al. Algorithms for Molecular Biology 2012, 7:18 RESEARCH Open Access Sampling solution traces for the problem of sorting permutations by signed reversals Christian Baudet 1,2*,ZanoniDias 3 and
On Hultman Numbers. 1 Introduction
 47 6 Journal of Integer Sequences, Vol 0 (007, Article 076 On Hultman Numbers Jean-Paul Doignon and Anthony Labarre Université Libre de Bruxelles Département de Mathématique, cp 6 Bd du Triomphe B-050
47 6 Journal of Integer Sequences, Vol 0 (007, Article 076 On Hultman Numbers Jean-Paul Doignon and Anthony Labarre Université Libre de Bruxelles Département de Mathématique, cp 6 Bd du Triomphe B-050
Design and Analysis of Algorithms Prof. Madhavan Mukund Chennai Mathematical Institute. Module 6 Lecture - 37 Divide and Conquer: Counting Inversions
 Design and Analysis of Algorithms Prof. Madhavan Mukund Chennai Mathematical Institute Module 6 Lecture - 37 Divide and Conquer: Counting Inversions Let us go back and look at Divide and Conquer again.
Design and Analysis of Algorithms Prof. Madhavan Mukund Chennai Mathematical Institute Module 6 Lecture - 37 Divide and Conquer: Counting Inversions Let us go back and look at Divide and Conquer again.
Algorithms for Finding Gene Clusters
 Algorithms for Finding Gene Clusters Steffen Heber 1 and Jens Stoye 2 1 Department of Computer Science & Engineering University of California, San Diego sheber@ucsd.edu 2 Max Planck Institute for Molecular
Algorithms for Finding Gene Clusters Steffen Heber 1 and Jens Stoye 2 1 Department of Computer Science & Engineering University of California, San Diego sheber@ucsd.edu 2 Max Planck Institute for Molecular
Efficient bounds for oriented chromosome inversion distance
 Efficient bounds for oriented chromosome inversion distance John Kececioglu* David Sanko~ Abstract We study the problem of comparing two circular chromosomes that have evolved by chromosome inversion,
Efficient bounds for oriented chromosome inversion distance John Kececioglu* David Sanko~ Abstract We study the problem of comparing two circular chromosomes that have evolved by chromosome inversion,
Factorization of permutation
 Department of Mathematics College of William and Mary Based on the paper: Zejun Huang,, Sharon H. Li, Nung-Sing Sze, Amidakuji/Ghost Leg Drawing Amidakuji/Ghost Leg Drawing It is a scheme for assigning
Department of Mathematics College of William and Mary Based on the paper: Zejun Huang,, Sharon H. Li, Nung-Sing Sze, Amidakuji/Ghost Leg Drawing Amidakuji/Ghost Leg Drawing It is a scheme for assigning
A group-theoretic approach to inversion distance
 A group-theoretic approach to inversion distance Andrew R Francis Centre for Research in Mathematics University of Western Sydney Australia Algebraic Statistics 2014 at IIT. Andrew R Francis (CRM @ UWS)
A group-theoretic approach to inversion distance Andrew R Francis Centre for Research in Mathematics University of Western Sydney Australia Algebraic Statistics 2014 at IIT. Andrew R Francis (CRM @ UWS)
Dyck paths, standard Young tableaux, and pattern avoiding permutations
 PU. M. A. Vol. 21 (2010), No.2, pp. 265 284 Dyck paths, standard Young tableaux, and pattern avoiding permutations Hilmar Haukur Gudmundsson The Mathematics Institute Reykjavik University Iceland e-mail:
PU. M. A. Vol. 21 (2010), No.2, pp. 265 284 Dyck paths, standard Young tableaux, and pattern avoiding permutations Hilmar Haukur Gudmundsson The Mathematics Institute Reykjavik University Iceland e-mail:
Edit Distances and Factorisations of Even Permutations
 Edit Distances and Factorisations of Even Permutations Anthony Labarre Université libre de Bruxelles (ULB), Département de Mathématique, CP 16 Service de Géométrie, Combinatoire et Théorie des Groupes
Edit Distances and Factorisations of Even Permutations Anthony Labarre Université libre de Bruxelles (ULB), Département de Mathématique, CP 16 Service de Géométrie, Combinatoire et Théorie des Groupes
arxiv: v2 [math.co] 16 Dec 2014
![arxiv: v2 [math.co] 16 Dec 2014 arxiv: v2 [math.co] 16 Dec 2014](/thumbs/86/93511762.jpg) SORTING PERMUTATIONS: GAMES, GENOMES, AND CYCLES K.L.M. ADAMYK, E. HOLMES, G.R. MAYFIELD, D.J. MORITZ, M. SCHEEPERS, B.E. TENNER, AND H.C. WAUCK arxiv:1410.2353v2 [math.co] 16 Dec 2014 Abstract. It has
SORTING PERMUTATIONS: GAMES, GENOMES, AND CYCLES K.L.M. ADAMYK, E. HOLMES, G.R. MAYFIELD, D.J. MORITZ, M. SCHEEPERS, B.E. TENNER, AND H.C. WAUCK arxiv:1410.2353v2 [math.co] 16 Dec 2014 Abstract. It has
Some algorithmic and combinatorial problems on permutation classes
 Some algorithmic and combinatorial problems on permutation classes The point of view of decomposition trees PhD Defense, 2009 December the 4th Outline 1 Objects studied : Permutations, Patterns and Classes
Some algorithmic and combinatorial problems on permutation classes The point of view of decomposition trees PhD Defense, 2009 December the 4th Outline 1 Objects studied : Permutations, Patterns and Classes
380 IEEE/ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS, VOL. 3, NO. 4, OCTOBER-DECEMBER 2006
 380 IEEE/ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS, VOL. 3, NO. 4, OCTOBER-DECEMBER 006 New Bounds and Tractable Instances for the Transposition Distance Anthony Labarre Abstract The
380 IEEE/ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS, VOL. 3, NO. 4, OCTOBER-DECEMBER 006 New Bounds and Tractable Instances for the Transposition Distance Anthony Labarre Abstract The
Medians of permutations and gene orders
 Medians of permutations and gene orders Sylvie Hamel Université de Montréal Work in collaboration with: Guillaume Blin Maxime Crochemore Stéphane Vialette from Université Marne-la-Vallée Medians of permutations
Medians of permutations and gene orders Sylvie Hamel Université de Montréal Work in collaboration with: Guillaume Blin Maxime Crochemore Stéphane Vialette from Université Marne-la-Vallée Medians of permutations
A Memory-Efficient Method for Fast Computation of Short 15-Puzzle Solutions
 A Memory-Efficient Method for Fast Computation of Short 15-Puzzle Solutions Ian Parberry Technical Report LARC-2014-02 Laboratory for Recreational Computing Department of Computer Science & Engineering
A Memory-Efficient Method for Fast Computation of Short 15-Puzzle Solutions Ian Parberry Technical Report LARC-2014-02 Laboratory for Recreational Computing Department of Computer Science & Engineering
arxiv: v1 [cs.cc] 21 Jun 2017
![arxiv: v1 [cs.cc] 21 Jun 2017 arxiv: v1 [cs.cc] 21 Jun 2017](/thumbs/75/72234152.jpg) Solving the Rubik s Cube Optimally is NP-complete Erik D. Demaine Sarah Eisenstat Mikhail Rudoy arxiv:1706.06708v1 [cs.cc] 21 Jun 2017 Abstract In this paper, we prove that optimally solving an n n n Rubik
Solving the Rubik s Cube Optimally is NP-complete Erik D. Demaine Sarah Eisenstat Mikhail Rudoy arxiv:1706.06708v1 [cs.cc] 21 Jun 2017 Abstract In this paper, we prove that optimally solving an n n n Rubik
MATH 433 Applied Algebra Lecture 12: Sign of a permutation (continued). Abstract groups.
 MATH 433 Applied Algebra Lecture 12: Sign of a permutation (continued). Abstract groups. Permutations Let X be a finite set. A permutation of X is a bijection from X to itself. The set of all permutations
MATH 433 Applied Algebra Lecture 12: Sign of a permutation (continued). Abstract groups. Permutations Let X be a finite set. A permutation of X is a bijection from X to itself. The set of all permutations
Sorting by Block Moves
 UNF Digital Commons UNF Theses and Dissertations Student Scholarship 2015 Sorting by Block Moves Jici Huang University of North Florida Suggested Citation Huang, Jici, "Sorting by Block Moves" (2015).
UNF Digital Commons UNF Theses and Dissertations Student Scholarship 2015 Sorting by Block Moves Jici Huang University of North Florida Suggested Citation Huang, Jici, "Sorting by Block Moves" (2015).
Counting Permutations by Putting Balls into Boxes
 Counting Permutations by Putting Balls into Boxes Ira M. Gessel Brandeis University C&O@40 Conference June 19, 2007 I will tell you shamelessly what my bottom line is: It is placing balls into boxes. Gian-Carlo
Counting Permutations by Putting Balls into Boxes Ira M. Gessel Brandeis University C&O@40 Conference June 19, 2007 I will tell you shamelessly what my bottom line is: It is placing balls into boxes. Gian-Carlo
UNO Gets Easier for a Single Player
 UNO Gets Easier for a Single Player Palash Dey, Prachi Goyal, and Neeldhara Misra Indian Institute of Science, Bangalore {palash prachi.goyal neeldhara}@csa.iisc.ernet.in Abstract This work is a follow
UNO Gets Easier for a Single Player Palash Dey, Prachi Goyal, and Neeldhara Misra Indian Institute of Science, Bangalore {palash prachi.goyal neeldhara}@csa.iisc.ernet.in Abstract This work is a follow
More Great Ideas in Theoretical Computer Science. Lecture 1: Sorting Pancakes
 15-252 More Great Ideas in Theoretical Computer Science Lecture 1: Sorting Pancakes January 19th, 2018 Question If there are n pancakes in total (all in different sizes), what is the max number of flips
15-252 More Great Ideas in Theoretical Computer Science Lecture 1: Sorting Pancakes January 19th, 2018 Question If there are n pancakes in total (all in different sizes), what is the max number of flips
Successor Rules for Flipping Pancakes and Burnt Pancakes
 Successor Rules for Flipping Pancakes and Burnt Pancakes J. Sawada a, A. Williams b a School of Computer Science, University of Guelph, Canada. Research supported by NSERC. E-mail: jsawada@uoguelph.ca
Successor Rules for Flipping Pancakes and Burnt Pancakes J. Sawada a, A. Williams b a School of Computer Science, University of Guelph, Canada. Research supported by NSERC. E-mail: jsawada@uoguelph.ca
An evolution of a permutation
 An evolution of a permutation Huseyin Acan April 28, 204 Joint work with Boris Pittel Notation and Definitions S n is the set of permutations of {,..., n} Notation and Definitions S n is the set of permutations
An evolution of a permutation Huseyin Acan April 28, 204 Joint work with Boris Pittel Notation and Definitions S n is the set of permutations of {,..., n} Notation and Definitions S n is the set of permutations
Greedy Flipping of Pancakes and Burnt Pancakes
 Greedy Flipping of Pancakes and Burnt Pancakes Joe Sawada a, Aaron Williams b a School of Computer Science, University of Guelph, Canada. Research supported by NSERC. b Department of Mathematics and Statistics,
Greedy Flipping of Pancakes and Burnt Pancakes Joe Sawada a, Aaron Williams b a School of Computer Science, University of Guelph, Canada. Research supported by NSERC. b Department of Mathematics and Statistics,
Permutation graphs an introduction
 Permutation graphs an introduction Ioan Todinca LIFO - Université d Orléans Algorithms and permutations, february / Permutation graphs Optimisation algorithms use, as input, the intersection model (realizer)
Permutation graphs an introduction Ioan Todinca LIFO - Université d Orléans Algorithms and permutations, february / Permutation graphs Optimisation algorithms use, as input, the intersection model (realizer)
Algorithms for Genetics: Basics of Wright Fisher Model and Coalescent Theory
 Algorithms for Genetics: Basics of Wright Fisher Model and Coalescent Theory Vineet Bafna Harish Nagarajan and Nitin Udpa 1 Disclaimer Please note that a lot of the text and figures here are copied from
Algorithms for Genetics: Basics of Wright Fisher Model and Coalescent Theory Vineet Bafna Harish Nagarajan and Nitin Udpa 1 Disclaimer Please note that a lot of the text and figures here are copied from
Regulatory Motif Finding II
 Regulatory Motif Finding II Lectures 13 Nov 9, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20 Johnson Hall (JHN) 022 1 Outline Regulatory
Regulatory Motif Finding II Lectures 13 Nov 9, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20 Johnson Hall (JHN) 022 1 Outline Regulatory
Analysis of Power Assignment in Radio Networks with Two Power Levels
 Analysis of Power Assignment in Radio Networks with Two Power Levels Miguel Fiandor Gutierrez & Manuel Macías Córdoba Abstract. In this paper we analyze the Power Assignment in Radio Networks with Two
Analysis of Power Assignment in Radio Networks with Two Power Levels Miguel Fiandor Gutierrez & Manuel Macías Córdoba Abstract. In this paper we analyze the Power Assignment in Radio Networks with Two
MA/CSSE 473 Day 13. Student Questions. Permutation Generation. HW 6 due Monday, HW 7 next Thursday, Tuesday s exam. Permutation generation
 MA/CSSE 473 Day 13 Permutation Generation MA/CSSE 473 Day 13 HW 6 due Monday, HW 7 next Thursday, Student Questions Tuesday s exam Permutation generation 1 Exam 1 If you want additional practice problems
MA/CSSE 473 Day 13 Permutation Generation MA/CSSE 473 Day 13 HW 6 due Monday, HW 7 next Thursday, Student Questions Tuesday s exam Permutation generation 1 Exam 1 If you want additional practice problems
Heuristic Search with Pre-Computed Databases
 Heuristic Search with Pre-Computed Databases Tsan-sheng Hsu tshsu@iis.sinica.edu.tw http://www.iis.sinica.edu.tw/~tshsu 1 Abstract Use pre-computed partial results to improve the efficiency of heuristic
Heuristic Search with Pre-Computed Databases Tsan-sheng Hsu tshsu@iis.sinica.edu.tw http://www.iis.sinica.edu.tw/~tshsu 1 Abstract Use pre-computed partial results to improve the efficiency of heuristic
Quotients of the Malvenuto-Reutenauer algebra and permutation enumeration
 Quotients of the Malvenuto-Reutenauer algebra and permutation enumeration Ira M. Gessel Department of Mathematics Brandeis University Sapienza Università di Roma July 10, 2013 Exponential generating functions
Quotients of the Malvenuto-Reutenauer algebra and permutation enumeration Ira M. Gessel Department of Mathematics Brandeis University Sapienza Università di Roma July 10, 2013 Exponential generating functions
X = {1, 2,...,n} n 1f 2f 3f... nf
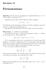 Section 11 Permutations Definition 11.1 Let X be a non-empty set. A bijective function f : X X will be called a permutation of X. Consider the case when X is the finite set with n elements: X {1, 2,...,n}.
Section 11 Permutations Definition 11.1 Let X be a non-empty set. A bijective function f : X X will be called a permutation of X. Consider the case when X is the finite set with n elements: X {1, 2,...,n}.
The Harassed Waitress Problem
 The Harassed Waitress Problem Harrah Essed Wei Therese Italian House of Pancakes Abstract. It is known that a stack of n pancakes can be rearranged in all n! ways by a sequence of n! 1 flips, and that
The Harassed Waitress Problem Harrah Essed Wei Therese Italian House of Pancakes Abstract. It is known that a stack of n pancakes can be rearranged in all n! ways by a sequence of n! 1 flips, and that
THE 15-PUZZLE (AND RUBIK S CUBE)
 THE 15-PUZZLE (AND RUBIK S CUBE) KEITH CONRAD 1. Introduction A permutation puzzle is a toy where the pieces can be moved around and the object is to reassemble the pieces into their beginning state We
THE 15-PUZZLE (AND RUBIK S CUBE) KEITH CONRAD 1. Introduction A permutation puzzle is a toy where the pieces can be moved around and the object is to reassemble the pieces into their beginning state We
((( ))) CS 19: Discrete Mathematics. Please feel free to ask questions! Getting into the mood. Pancakes With A Problem!
 CS : Discrete Mathematics Professor Amit Chakrabarti Please feel free to ask questions! ((( ))) Teaching Assistants Chien-Chung Huang David Blinn http://www.cs cs.dartmouth.edu/~cs Getting into the mood
CS : Discrete Mathematics Professor Amit Chakrabarti Please feel free to ask questions! ((( ))) Teaching Assistants Chien-Chung Huang David Blinn http://www.cs cs.dartmouth.edu/~cs Getting into the mood
CS3334 Data Structures Lecture 4: Bubble Sort & Insertion Sort. Chee Wei Tan
 CS3334 Data Structures Lecture 4: Bubble Sort & Insertion Sort Chee Wei Tan Sorting Since Time Immemorial Plimpton 322 Tablet: Sorted Pythagorean Triples https://www.maa.org/sites/default/files/pdf/news/monthly105-120.pdf
CS3334 Data Structures Lecture 4: Bubble Sort & Insertion Sort Chee Wei Tan Sorting Since Time Immemorial Plimpton 322 Tablet: Sorted Pythagorean Triples https://www.maa.org/sites/default/files/pdf/news/monthly105-120.pdf
Permutation classes and infinite antichains
 Permutation classes and infinite antichains Robert Brignall Based on joint work with David Bevan and Nik Ruškuc Dartmouth College, 12th July 2018 Typical questions in PP For a permutation class C: What
Permutation classes and infinite antichains Robert Brignall Based on joint work with David Bevan and Nik Ruškuc Dartmouth College, 12th July 2018 Typical questions in PP For a permutation class C: What
Permutation Editing and Matching via Embeddings
 Permutation Editing and Matching via Embeddings Graham Cormode, S. Muthukrishnan, Cenk Sahinalp (grahamc@dcs.warwick.ac.uk) Permutation Editing and Matching Why study permutations? Distances between permutations
Permutation Editing and Matching via Embeddings Graham Cormode, S. Muthukrishnan, Cenk Sahinalp (grahamc@dcs.warwick.ac.uk) Permutation Editing and Matching Why study permutations? Distances between permutations
Creating a Dominion AI Using Genetic Algorithms
 Creating a Dominion AI Using Genetic Algorithms Abstract Mok Ming Foong Dominion is a deck-building card game. It allows for complex strategies, has an aspect of randomness in card drawing, and no obvious
Creating a Dominion AI Using Genetic Algorithms Abstract Mok Ming Foong Dominion is a deck-building card game. It allows for complex strategies, has an aspect of randomness in card drawing, and no obvious
LANDSCAPE SMOOTHING OF NUMERICAL PERMUTATION SPACES IN GENETIC ALGORITHMS
 LANDSCAPE SMOOTHING OF NUMERICAL PERMUTATION SPACES IN GENETIC ALGORITHMS ABSTRACT The recent popularity of genetic algorithms (GA s) and their application to a wide range of problems is a result of their
LANDSCAPE SMOOTHING OF NUMERICAL PERMUTATION SPACES IN GENETIC ALGORITHMS ABSTRACT The recent popularity of genetic algorithms (GA s) and their application to a wide range of problems is a result of their
Ancestral Recombination Graphs
 Ancestral Recombination Graphs Ancestral relationships among a sample of recombining sequences usually cannot be accurately described by just a single genealogy. Linked sites will have similar, but not
Ancestral Recombination Graphs Ancestral relationships among a sample of recombining sequences usually cannot be accurately described by just a single genealogy. Linked sites will have similar, but not
Fast Sorting and Pattern-Avoiding Permutations
 Fast Sorting and Pattern-Avoiding Permutations David Arthur Stanford University darthur@cs.stanford.edu Abstract We say a permutation π avoids a pattern σ if no length σ subsequence of π is ordered in
Fast Sorting and Pattern-Avoiding Permutations David Arthur Stanford University darthur@cs.stanford.edu Abstract We say a permutation π avoids a pattern σ if no length σ subsequence of π is ordered in
Computing common intervals of K permutations, with applications to modular decomposition of graphs
 Computing common intervals of K permutations, with applications to modular decomposition of graphs Anne Bergeron, Cedric Chauve LaCIM and Département d Informatique Université du Québec à Montréal Fabien
Computing common intervals of K permutations, with applications to modular decomposition of graphs Anne Bergeron, Cedric Chauve LaCIM and Département d Informatique Université du Québec à Montréal Fabien
Lecture 13 Register Allocation: Coalescing
 Lecture 13 Register llocation: Coalescing I. Motivation II. Coalescing Overview III. lgorithms: Simple & Safe lgorithm riggs lgorithm George s lgorithm Phillip. Gibbons 15-745: Register Coalescing 1 Review:
Lecture 13 Register llocation: Coalescing I. Motivation II. Coalescing Overview III. lgorithms: Simple & Safe lgorithm riggs lgorithm George s lgorithm Phillip. Gibbons 15-745: Register Coalescing 1 Review:
Pattern Matching for Permutations
 Pattern Matching for Permutations Stéphane Vialette 2 CNRS & LIGM, Université Paris-Est Marne-la-Vallée, France Permutation Pattern 2013, Paris Vialette (LIGM UPEMLV) Pattern Matching PP 2013 1 / 69 Outline
Pattern Matching for Permutations Stéphane Vialette 2 CNRS & LIGM, Université Paris-Est Marne-la-Vallée, France Permutation Pattern 2013, Paris Vialette (LIGM UPEMLV) Pattern Matching PP 2013 1 / 69 Outline
Math Fundamentals for Statistics (Math 52) Unit 2:Number Line and Ordering. By Scott Fallstrom and Brent Pickett The How and Whys Guys.
 Math Fundamentals for Statistics (Math 52) Unit 2:Number Line and Ordering By Scott Fallstrom and Brent Pickett The How and Whys Guys Unit 2 Page 1 2.1: Place Values We just looked at graphing ordered
Math Fundamentals for Statistics (Math 52) Unit 2:Number Line and Ordering By Scott Fallstrom and Brent Pickett The How and Whys Guys Unit 2 Page 1 2.1: Place Values We just looked at graphing ordered
The Problem. Tom Davis December 19, 2016
 The 1 2 3 4 Problem Tom Davis tomrdavis@earthlink.net http://www.geometer.org/mathcircles December 19, 2016 Abstract The first paragraph in the main part of this article poses a problem that can be approached
The 1 2 3 4 Problem Tom Davis tomrdavis@earthlink.net http://www.geometer.org/mathcircles December 19, 2016 Abstract The first paragraph in the main part of this article poses a problem that can be approached
Algorithmics of Directional Antennae: Strong Connectivity with Multiple Antennae
 Algorithmics of Directional Antennae: Strong Connectivity with Multiple Antennae Ioannis Caragiannis Stefan Dobrev Christos Kaklamanis Evangelos Kranakis Danny Krizanc Jaroslav Opatrny Oscar Morales Ponce
Algorithmics of Directional Antennae: Strong Connectivity with Multiple Antennae Ioannis Caragiannis Stefan Dobrev Christos Kaklamanis Evangelos Kranakis Danny Krizanc Jaroslav Opatrny Oscar Morales Ponce
MITOCW watch?v=-qcpo_dwjk4
 MITOCW watch?v=-qcpo_dwjk4 The following content is provided under a Creative Commons license. Your support will help MIT OpenCourseWare continue to offer high quality educational resources for free. To
MITOCW watch?v=-qcpo_dwjk4 The following content is provided under a Creative Commons license. Your support will help MIT OpenCourseWare continue to offer high quality educational resources for free. To
arxiv: v1 [math.co] 8 Aug 2018
![arxiv: v1 [math.co] 8 Aug 2018 arxiv: v1 [math.co] 8 Aug 2018](/thumbs/83/87604666.jpg) Permutation patterns in genome rearrangement problems Giulio Cerbai giuliocerbai14@gmailcom Luca Ferrari lucaferrari@unifiit Dipartimento di Matematica e Informatica U Dini, viale Morgagni 65, University
Permutation patterns in genome rearrangement problems Giulio Cerbai giuliocerbai14@gmailcom Luca Ferrari lucaferrari@unifiit Dipartimento di Matematica e Informatica U Dini, viale Morgagni 65, University
Pokémon Cards and the Shortest Common Superstring
 San Jose State University From the SelectedWorks of Mark Stamp 2004 Pokémon Cards and the Shortest Common Superstring Mark Stamp Austin E Stamp Available at: https://works.bepress.com/mark_stamp/90/ Graph
San Jose State University From the SelectedWorks of Mark Stamp 2004 Pokémon Cards and the Shortest Common Superstring Mark Stamp Austin E Stamp Available at: https://works.bepress.com/mark_stamp/90/ Graph
Scheduling. Radek Mařík. April 28, 2015 FEE CTU, K Radek Mařík Scheduling April 28, / 48
 Scheduling Radek Mařík FEE CTU, K13132 April 28, 2015 Radek Mařík (marikr@fel.cvut.cz) Scheduling April 28, 2015 1 / 48 Outline 1 Introduction to Scheduling Methodology Overview 2 Classification of Scheduling
Scheduling Radek Mařík FEE CTU, K13132 April 28, 2015 Radek Mařík (marikr@fel.cvut.cz) Scheduling April 28, 2015 1 / 48 Outline 1 Introduction to Scheduling Methodology Overview 2 Classification of Scheduling
Problem 2A Consider 101 natural numbers not exceeding 200. Prove that at least one of them is divisible by another one.
 1. Problems from 2007 contest Problem 1A Do there exist 10 natural numbers such that none one of them is divisible by another one, and the square of any one of them is divisible by any other of the original
1. Problems from 2007 contest Problem 1A Do there exist 10 natural numbers such that none one of them is divisible by another one, and the square of any one of them is divisible by any other of the original
Generating indecomposable permutations
 Discrete Mathematics 306 (2006) 508 518 www.elsevier.com/locate/disc Generating indecomposable permutations Andrew King Department of Computer Science, McGill University, Montreal, Que., Canada Received
Discrete Mathematics 306 (2006) 508 518 www.elsevier.com/locate/disc Generating indecomposable permutations Andrew King Department of Computer Science, McGill University, Montreal, Que., Canada Received
17. Symmetries. Thus, the example above corresponds to the matrix: We shall now look at how permutations relate to trees.
 7 Symmetries 7 Permutations A permutation of a set is a reordering of its elements Another way to look at it is as a function Φ that takes as its argument a set of natural numbers of the form {, 2,, n}
7 Symmetries 7 Permutations A permutation of a set is a reordering of its elements Another way to look at it is as a function Φ that takes as its argument a set of natural numbers of the form {, 2,, n}
Online Call Control in Cellular Networks Revisited
 Online Call Control in Cellular Networks Revisited Yong Zhang Francis Y.L. Chin Hing-Fung Ting Joseph Wun-Tat Chan Xin Han Ka-Cheong Lam Abstract Wireless Communication Networks based on Frequency Division
Online Call Control in Cellular Networks Revisited Yong Zhang Francis Y.L. Chin Hing-Fung Ting Joseph Wun-Tat Chan Xin Han Ka-Cheong Lam Abstract Wireless Communication Networks based on Frequency Division
A comparison of a genetic algorithm and a depth first search algorithm applied to Japanese nonograms
 A comparison of a genetic algorithm and a depth first search algorithm applied to Japanese nonograms Wouter Wiggers Faculty of EECMS, University of Twente w.a.wiggers@student.utwente.nl ABSTRACT In this
A comparison of a genetic algorithm and a depth first search algorithm applied to Japanese nonograms Wouter Wiggers Faculty of EECMS, University of Twente w.a.wiggers@student.utwente.nl ABSTRACT In this
Mitochondrial DNA (mtdna) JGSGO June 5, 2018
 Mitochondrial DNA (mtdna) JGSGO June 5, 2018 MtDNA - outline What is it? What do you do with it? How do you maximize its value? 2 3 mtdna a double-stranded, circular DNA that is stored in mitochondria
Mitochondrial DNA (mtdna) JGSGO June 5, 2018 MtDNA - outline What is it? What do you do with it? How do you maximize its value? 2 3 mtdna a double-stranded, circular DNA that is stored in mitochondria
Permutations with short monotone subsequences
 Permutations with short monotone subsequences Dan Romik Abstract We consider permutations of 1, 2,..., n 2 whose longest monotone subsequence is of length n and are therefore extremal for the Erdős-Szekeres
Permutations with short monotone subsequences Dan Romik Abstract We consider permutations of 1, 2,..., n 2 whose longest monotone subsequence is of length n and are therefore extremal for the Erdős-Szekeres
Convergence in competitive games
 Convergence in competitive games Vahab S. Mirrokni Computer Science and AI Lab. (CSAIL) and Math. Dept., MIT. This talk is based on joint works with A. Vetta and with A. Sidiropoulos, A. Vetta DIMACS Bounded
Convergence in competitive games Vahab S. Mirrokni Computer Science and AI Lab. (CSAIL) and Math. Dept., MIT. This talk is based on joint works with A. Vetta and with A. Sidiropoulos, A. Vetta DIMACS Bounded
THE ERDŐS-KO-RADO THEOREM FOR INTERSECTING FAMILIES OF PERMUTATIONS
 THE ERDŐS-KO-RADO THEOREM FOR INTERSECTING FAMILIES OF PERMUTATIONS A Thesis Submitted to the Faculty of Graduate Studies and Research In Partial Fulfillment of the Requirements for the Degree of Master
THE ERDŐS-KO-RADO THEOREM FOR INTERSECTING FAMILIES OF PERMUTATIONS A Thesis Submitted to the Faculty of Graduate Studies and Research In Partial Fulfillment of the Requirements for the Degree of Master
Inputs. Outputs. Outputs. Inputs. Outputs. Inputs
 Permutation Admissibility in Shue-Exchange Networks with Arbitrary Number of Stages Nabanita Das Bhargab B. Bhattacharya Rekha Menon Indian Statistical Institute Calcutta, India ndas@isical.ac.in Sergei
Permutation Admissibility in Shue-Exchange Networks with Arbitrary Number of Stages Nabanita Das Bhargab B. Bhattacharya Rekha Menon Indian Statistical Institute Calcutta, India ndas@isical.ac.in Sergei
A Hybrid Evolutionary Approach for Multi Robot Path Exploration Problem
 A Hybrid Evolutionary Approach for Multi Robot Path Exploration Problem K.. enthilkumar and K. K. Bharadwaj Abstract - Robot Path Exploration problem or Robot Motion planning problem is one of the famous
A Hybrid Evolutionary Approach for Multi Robot Path Exploration Problem K.. enthilkumar and K. K. Bharadwaj Abstract - Robot Path Exploration problem or Robot Motion planning problem is one of the famous
Three of these grids share a property that the other three do not. Can you find such a property? + mod
 PPMTC 22 Session 6: Mad Vet Puzzles Session 6: Mad Veterinarian Puzzles There is a collection of problems that have come to be known as "Mad Veterinarian Puzzles", for reasons which will soon become obvious.
PPMTC 22 Session 6: Mad Vet Puzzles Session 6: Mad Veterinarian Puzzles There is a collection of problems that have come to be known as "Mad Veterinarian Puzzles", for reasons which will soon become obvious.
Local Search: Hill Climbing. When A* doesn t work AIMA 4.1. Review: Hill climbing on a surface of states. Review: Local search and optimization
 Outline When A* doesn t work AIMA 4.1 Local Search: Hill Climbing Escaping Local Maxima: Simulated Annealing Genetic Algorithms A few slides adapted from CS 471, UBMC and Eric Eaton (in turn, adapted from
Outline When A* doesn t work AIMA 4.1 Local Search: Hill Climbing Escaping Local Maxima: Simulated Annealing Genetic Algorithms A few slides adapted from CS 471, UBMC and Eric Eaton (in turn, adapted from
Approximation Algorithms for Conflict-Free Vehicle Routing
 Approximation Algorithms for Conflict-Free Vehicle Routing Kaspar Schupbach and Rico Zenklusen Παπαηλίου Νικόλαος CFVRP Problem Undirected graph of stations and roads Vehicles(k): Source-Destination stations
Approximation Algorithms for Conflict-Free Vehicle Routing Kaspar Schupbach and Rico Zenklusen Παπαηλίου Νικόλαος CFVRP Problem Undirected graph of stations and roads Vehicles(k): Source-Destination stations
Olympiad Combinatorics. Pranav A. Sriram
 Olympiad Combinatorics Pranav A. Sriram August 2014 Chapter 2: Algorithms - Part II 1 Copyright notices All USAMO and USA Team Selection Test problems in this chapter are copyrighted by the Mathematical
Olympiad Combinatorics Pranav A. Sriram August 2014 Chapter 2: Algorithms - Part II 1 Copyright notices All USAMO and USA Team Selection Test problems in this chapter are copyrighted by the Mathematical
Determinants, Part 1
 Determinants, Part We shall start with some redundant definitions. Definition. Given a matrix A [ a] we say that determinant of A is det A a. Definition 2. Given a matrix a a a 2 A we say that determinant
Determinants, Part We shall start with some redundant definitions. Definition. Given a matrix A [ a] we say that determinant of A is det A a. Definition 2. Given a matrix a a a 2 A we say that determinant
CSE 417: Review. Larry Ruzzo
 CSE 417: Review Larry Ruzzo 1 Complexity, I Asymptotic Analysis Best/average/worst cases Upper/Lower Bounds Big O, Theta, Omega definitions; intuition Analysis methods loops recurrence relations common
CSE 417: Review Larry Ruzzo 1 Complexity, I Asymptotic Analysis Best/average/worst cases Upper/Lower Bounds Big O, Theta, Omega definitions; intuition Analysis methods loops recurrence relations common
NOTES ON SEPT 13-18, 2012
 NOTES ON SEPT 13-18, 01 MIKE ZABROCKI Last time I gave a name to S(n, k := number of set partitions of [n] into k parts. This only makes sense for n 1 and 1 k n. For other values we need to choose a convention
NOTES ON SEPT 13-18, 01 MIKE ZABROCKI Last time I gave a name to S(n, k := number of set partitions of [n] into k parts. This only makes sense for n 1 and 1 k n. For other values we need to choose a convention
Pattern Avoidance in Poset Permutations
 Pattern Avoidance in Poset Permutations Sam Hopkins and Morgan Weiler Massachusetts Institute of Technology and University of California, Berkeley Permutation Patterns, Paris; July 5th, 2013 1 Definitions
Pattern Avoidance in Poset Permutations Sam Hopkins and Morgan Weiler Massachusetts Institute of Technology and University of California, Berkeley Permutation Patterns, Paris; July 5th, 2013 1 Definitions
Variant Calling. Michael Schatz. Feb 20, 2018 Lecture 7: Applied Comparative Genomics
 Variant Calling Michael Schatz Feb 20, 2018 Lecture 7: Applied Comparative Genomics Mission Impossible 1. Setup VirtualBox 2. Initialize Tools 3. Download Reference Genome & Reads 4. Decode the secret
Variant Calling Michael Schatz Feb 20, 2018 Lecture 7: Applied Comparative Genomics Mission Impossible 1. Setup VirtualBox 2. Initialize Tools 3. Download Reference Genome & Reads 4. Decode the secret
Adaptive Hybrid Channel Assignment in Wireless Mobile Network via Genetic Algorithm
 Adaptive Hybrid Channel Assignment in Wireless Mobile Network via Genetic Algorithm Y.S. Chia Z.W. Siew A. Kiring S.S. Yang K.T.K. Teo Modelling, Simulation and Computing Laboratory School of Engineering
Adaptive Hybrid Channel Assignment in Wireless Mobile Network via Genetic Algorithm Y.S. Chia Z.W. Siew A. Kiring S.S. Yang K.T.K. Teo Modelling, Simulation and Computing Laboratory School of Engineering
Automedians sets of permutation: extended abstract
 Automedians sets of permutation: extended abstract Charles Desharnais and Sylvie Hamel DIRO - Université de Montréal, C. P. 6128 Succursale Centre-Ville, Montréal, Québec, Canada, H3C 3J7, {charles.desharnais,
Automedians sets of permutation: extended abstract Charles Desharnais and Sylvie Hamel DIRO - Université de Montréal, C. P. 6128 Succursale Centre-Ville, Montréal, Québec, Canada, H3C 3J7, {charles.desharnais,
Lecture 12: Divide and Conquer Algorithms. Divide and Conquer Algorithms
 Lecture 12: Divide and Conquer Algorithms Study Chapter 7.1 7.4 1 Divide and Conquer Algorithms Divide problem into sub-problems Conquer by solving sub-problems recursively. If the sub-problems are small
Lecture 12: Divide and Conquer Algorithms Study Chapter 7.1 7.4 1 Divide and Conquer Algorithms Divide problem into sub-problems Conquer by solving sub-problems recursively. If the sub-problems are small
Phylogeny and Molecular Evolution
 Phylogeny and Molecular Evolution Character Based Phylogeny Large Parsimony 1/50 Credit Ron Shamir s lecture notes Notes by Nir Friedman Dan Geiger, Shlomo Moran, Sagi Snir and Ron Shamir Durbin et al.
Phylogeny and Molecular Evolution Character Based Phylogeny Large Parsimony 1/50 Credit Ron Shamir s lecture notes Notes by Nir Friedman Dan Geiger, Shlomo Moran, Sagi Snir and Ron Shamir Durbin et al.
ON THE PERMUTATIONAL POWER OF TOKEN PASSING NETWORKS.
 ON THE PERMUTATIONAL POWER OF TOKEN PASSING NETWORKS. M. H. ALBERT, N. RUŠKUC, AND S. LINTON Abstract. A token passing network is a directed graph with one or more specified input vertices and one or more
ON THE PERMUTATIONAL POWER OF TOKEN PASSING NETWORKS. M. H. ALBERT, N. RUŠKUC, AND S. LINTON Abstract. A token passing network is a directed graph with one or more specified input vertices and one or more
What is a Sorting Function?
 Department of Computer Science University of Copenhagen Email: henglein@diku.dk WG 2.8 2008, Park City, June 15-22, 2008 Outline 1 Sorting algorithms Literature definitions What is a sorting criterion?
Department of Computer Science University of Copenhagen Email: henglein@diku.dk WG 2.8 2008, Park City, June 15-22, 2008 Outline 1 Sorting algorithms Literature definitions What is a sorting criterion?
Lecture Notes 3: Paging, K-Server and Metric Spaces
 Online Algorithms 16/11/11 Lecture Notes 3: Paging, K-Server and Metric Spaces Professor: Yossi Azar Scribe:Maor Dan 1 Introduction This lecture covers the Paging problem. We present a competitive online
Online Algorithms 16/11/11 Lecture Notes 3: Paging, K-Server and Metric Spaces Professor: Yossi Azar Scribe:Maor Dan 1 Introduction This lecture covers the Paging problem. We present a competitive online
Application of the GenomeZipper-Approach on Wheat. Mihaela Martis MIPS, Munich
 Applicati of the GenomeZipper-Approach Wheat Mihaela Martis MIPS, Munich Overview From barley to wheat Introducti to the GenomeZipper Pipeline Repeat masking Homology search Synteny detecti GenomeZipper
Applicati of the GenomeZipper-Approach Wheat Mihaela Martis MIPS, Munich Overview From barley to wheat Introducti to the GenomeZipper Pipeline Repeat masking Homology search Synteny detecti GenomeZipper
Coalescence. Outline History. History, Model, and Application. Coalescence. The Model. Application
 Coalescence History, Model, and Application Outline History Origins of theory/approach Trace the incorporation of other s ideas Coalescence Definition and descriptions The Model Assumptions and Uses Application
Coalescence History, Model, and Application Outline History Origins of theory/approach Trace the incorporation of other s ideas Coalescence Definition and descriptions The Model Assumptions and Uses Application
