Rapid Non linear Image Scanning Microscopy, Supplementary Notes
|
|
|
- Dylan Flowers
- 5 years ago
- Views:
Transcription
1 Rapid Non linear Image Scanning Microscopy, Supplementary Notes Calculation of theoretical PSFs We calculated the electrical field distribution using the wave optical theory developed by Wolf 1, and Richards and Wolf 2, for imaging through optical systems with high numerical aperture. From the obtained field we calculated the intensity,, and the squared intensity,,. To describe the PSF we use the FWHM diameters of,, along and perpendicular to the z axis along which the light propagates. We approximated the incident laser beam at the back focal aperture of the objective by a Gaussian beam and calculated the extensions of the PSFs for back focal beam radii ranging from 2 mm to 4 mm. The values of the other parameters used were: numerical aperture NA 1.3; refractive index of immersion medium equal to 1.41; excitation wavelength 900 nm. Figure S1: (a) 3D representation of the 2PE PSF as an isosurface corresponding to ½ of its maximum value. (b) axial (+) and lateral (o) diameters (FWHM) of the 2PE PSF and their dependence on the beam diameter in the back focal aperture of the objective. 1. Wolf, E. Electromagnetic diffration in optical systems 1. An integral representation of the image field. Proceedings of the Royal Society, (1959). 2. Richards, B. & Wolf, E. Electromagnetic diffraction in optical systems 2: Structure of the image field in an aplanatic system. Proceedings of the Royal Society, (1959).
2 Rendered images of 3D SHG signals Figure S2 shows different views of a 3D stack of the SHG signal of a collagen I hydrogel as described in the online methods. Figure S2: 3D representations of conventional microscopy (a,c) and de convolved SHG ISM (b,d). (a) and (b) show pseudo colored z stacks; (c) and (d) show maximum intensity volumetric renderings. Image size: 63.5 µm x 29.5 µm.
3 Beam path in the ISM microscope The use of a galvo scanner with a resonant and a conventional scan axis has some implications to the beam path that are not obvious at first sight. The two mirrors create a lateral shift between the optical axis of the objective and the optical axis of the entrance port of the scanner. When feeding the emitted light back into the scanner, this shift needs to be compensated. The second implication is that the directions of the movement of the beam has to be matched when feeding the beam back to the scanner. This is illustrated in figure S3. There, the movement, generated by the first scan mirror (SM1) is upwards, as indicated by the green arrow on top of the beam. For the emitted beam (shown in blue) this movement is identical and if it was simply reflected upwards, the movement does not match the motion of SM1. Hence, in order to obtain a doubling of the scan angle, one needs to rotate and invert the beam in combination with the necessary lateral shift. In our setup this is achieved by a combination of three mirrors (M1 to M3). Figure S3: Schematic view of the beam path in the ISM microscope. For simplicity, the scan and tube lenses are not shown. The excitation light is shown in red, light
4 emitted by the sample is shown in blue. Sections where excitation and fluorescence overlap are shown in green. The arrows indicate the beam deflection due to movement of the resonant scan mirror (green) and the second scan mirror (red). The excitation beam enters the setup and is reflected by the dichroic mirror (DCM3) onto the optical axis of the scanner. The beam is scanned via the resonant scan mirror (SM1) and the second scan mirror (SM2). It propagates through the dichroic mirror (DCM1) and is reflected by the dichroic mirror (DCM2) into the microscope objective. The emitted light is collected by the objective, passes DCM2 and is reflected by a mirror to the arrangement of three mirrors (M1 M3). These mirrors align the beam with the axes of the scanner. Then, the beam passes DCM3 and is re scanned via SM1 and SM2. Finally, the light is coupled out by DCM1 and imaged onto a camera (not shown).
5 Image data processing Processing of the recorded imaging data is done according to the following steps: 1. Background subtraction. A background dataset consisting of 100 frames is recorded using the same settings as for the image recording, but without excitation. The frames are averaged and smoothed with a Gaussian filter. The resultant representation is then subtracted from the image data. Negative results are clipped to zero. It is also possible to simply subtract an average value taken from unexposed areas of the camera sensor if the background data is not available. 2. Removal of pixel artifacts. To remove artifacts from hot or cold pixels we proceed as follows. For each pixel we calculate the expected value E by linear interpolation for its surrounding pixels. We also compute the local variance based on the interpolation for the surrounding pixels. If the pixel value exceeds the range 3 we consider this as an artifact and set this pixel to the value of E. If one uses ImageJ one can use the Remove outliers function using a threshold based on the variance of the histogram of the image data. 3. Final processing. After an optional Fourier reweighting, the data is filtered using a 2D Gaussian filter with a radius of 1 pixel (about 46 nm) and, usually, 2x2 pixels are binned. Finally, the 16 bit data is downscaled to 8 bit dynamic range.
6 Fourier reweighting 1. The first aim is to compute the Fourier transform F(k) of the image data. To minimize problems by boundary effects, we pad the original data using accordingly flipped patches from each respective border. The aim is to obtain an image of the size N x N, where log ceil max, 1, and a, b denote the size of the image. 2. Additionally, we let the intensity of the synthesized image decay toward the borders using a linear ramp that falls from 1 to 0 in the range R = N/2 to R = N. 3. Next, we do the Fourier transform of the conditioned data. We separate the modulus and the phase angle of the FT data, since we only want to re weigh the modulus. The modulus can take values different from 0 only for max where max is the maximal length of a k vector that is transmitted by the objective: max 4 NA/ em. This, for all max F(k) has to be The ideal MTF curve of a microscope would be fi 2/π acos 1 with ma. Since the imaging process in ISM delivers a MTF that is basically raw fi one could use a weight function 1 fi to recover fi from raw. Since the fi has low amplitudes for 1 this would mean high values of and thus amplifying noise. Therefore one has to use a regularization avoiding this. A simple approach is 1 fi where takes reasonably small values. Typically 0.02 works well, but this depends on the SNR. 5. After applying the corrections on F(k) one can back transform the data into real space. 6. A MatLab script showing this procedure on sample data is shown below. 7. Since the ISM raw images already contain the expanded frequency support, one can also achieve the resolution enhancement using so called deconvolution algorithms, e.g. Richardson Lucy deconvolution. Classical deconvolution is not able to gain resolution improvements but sharpens blurring by trying to de convolve image data with the PSF. This essentially is the same operation that one implicitly does when reweighting the amplitudes in the MTF. Therefore, these algorithms can be applied in ISM, too.
7 An exemplary MatLab script showing the steps of this procedure: % read data from file name = 'actin.tif'; data = double(imread(name,'tiff')); data = max(data,0); % clip negative values [nx,ny] = size(data); N = 2^ceil(log2(max(nx,ny))+1); % size of image for Fourier transform % stich patches of the original image to form a periodic 'super-image' im = [data fliplr(data) data fliplr(data) data]; im = [flipud(im); im; flipud(im); im; flipud(im); im; flipud(im)]; % cut out the center part of the 'super-image' of size NxX [sx,sy] = size(im); im = im(floor((sx-n)/2)+(1:n),floor((sy-n)/2)+(1:n)); % let the signal decay to zero towards the edges [X,Y] = meshgrid(1:n,1:n); X = (X-N/2-1); Y = (Y-N/2-1); R = sqrt(x.^2+y.^2); W = 2-8*R/N; W = min(w,1); W = max(w,0); im = W.*im; % Do the Fourier transform fim = fftshift(fft2(im)); fa = abs(fim); % modulus fp = angle(fim); % Phase angle % calculate k-vectors of the Fourier-transformed image fs = 1/0.091; % per µm k = fs/2/(n/2).*r; km = 5.5; ind = double(k<km); % Step function to clip everything outside km to zero % 'Ideal' MFT OTF = ind.*2/pi.*(acos(k./km)-k./km.*sqrt(1-(k./km).^2)); % Re-weighting function ep = 0.015; w = ind./(otf + ep.*k./km); % Back-transform the re-weighted FT tim = abs(ifft2(ifftshift(w.*fa.*exp(1i.*fp)))); % cut out the original field of the data tim = tim((n-nx)/2+(1:nx),(n-ny)/2+(1:ny)); % display original data vs. re-weighted data imagesc([data; tim]) colormap(gray(256)) caxis([0 350]) axis image axis off
8 Description of Additional Files and Videos SupplementaryVideo 1.mov : This is a rendering of a collagen I gel as described in the paper. The data was recorded done using a conventional SHG confocal microscope. SupplementaryVideo 2.mov : This is a rendering of the same sample as in SHG.mp4. In this case the data was recorded using the ISM confocal microscope. SupplementaryVideo 3.mov : This video shows fluorescent beads (d = 100 nm) diffusing freely in water. The recording frame rate is 20 fps. The acquisition speed is sufficient to track individual beads for several frames until they diffuse out of focus. SupplementaryVideo 4.mov, SupplementaryVideo 5.mov, SupplementaryVideo 6.mov, and SupplementaryVideo 7.mov are rendered video sequences time series of 3D stacks of living Drosophila embryos as described in the Online Methods. The frame interval is 60 s or 120 s in the case of SupplementaryVideo 7.mov. Spacing of the z planes is 3 µm. The sections are taken around the center height of the embryo.
Nature Methods: doi: /nmeth Supplementary Figure 1. Schematic of 2P-ISIM AO optical setup.
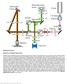 Supplementary Figure 1 Schematic of 2P-ISIM AO optical setup. Excitation from a femtosecond laser is passed through intensity control and shuttering optics (1/2 λ wave plate, polarizing beam splitting
Supplementary Figure 1 Schematic of 2P-ISIM AO optical setup. Excitation from a femtosecond laser is passed through intensity control and shuttering optics (1/2 λ wave plate, polarizing beam splitting
Point Spread Function. Confocal Laser Scanning Microscopy. Confocal Aperture. Optical aberrations. Alternative Scanning Microscopy
 Bi177 Lecture 5 Adding the Third Dimension Wide-field Imaging Point Spread Function Deconvolution Confocal Laser Scanning Microscopy Confocal Aperture Optical aberrations Alternative Scanning Microscopy
Bi177 Lecture 5 Adding the Third Dimension Wide-field Imaging Point Spread Function Deconvolution Confocal Laser Scanning Microscopy Confocal Aperture Optical aberrations Alternative Scanning Microscopy
Nature Neuroscience: doi: /nn Supplementary Figure 1. Optimized Bessel foci for in vivo volume imaging.
 Supplementary Figure 1 Optimized Bessel foci for in vivo volume imaging. (a) Images taken by scanning Bessel foci of various NAs, lateral and axial FWHMs: (Left panels) in vivo volume images of YFP + neurites
Supplementary Figure 1 Optimized Bessel foci for in vivo volume imaging. (a) Images taken by scanning Bessel foci of various NAs, lateral and axial FWHMs: (Left panels) in vivo volume images of YFP + neurites
3D light microscopy techniques
 3D light microscopy techniques The image of a point is a 3D feature In-focus image Out-of-focus image The image of a point is not a point Point Spread Function (PSF) 1D imaging 2D imaging 3D imaging Resolution
3D light microscopy techniques The image of a point is a 3D feature In-focus image Out-of-focus image The image of a point is not a point Point Spread Function (PSF) 1D imaging 2D imaging 3D imaging Resolution
Why and How? Daniel Gitler Dept. of Physiology Ben-Gurion University of the Negev. Microscopy course, Michmoret Dec 2005
 Why and How? Daniel Gitler Dept. of Physiology Ben-Gurion University of the Negev Why use confocal microscopy? Principles of the laser scanning confocal microscope. Image resolution. Manipulating the
Why and How? Daniel Gitler Dept. of Physiology Ben-Gurion University of the Negev Why use confocal microscopy? Principles of the laser scanning confocal microscope. Image resolution. Manipulating the
Kit for building your own THz Time-Domain Spectrometer
 Kit for building your own THz Time-Domain Spectrometer 16/06/2016 1 Table of contents 0. Parts for the THz Kit... 3 1. Delay line... 4 2. Pulse generator and lock-in detector... 5 3. THz antennas... 6
Kit for building your own THz Time-Domain Spectrometer 16/06/2016 1 Table of contents 0. Parts for the THz Kit... 3 1. Delay line... 4 2. Pulse generator and lock-in detector... 5 3. THz antennas... 6
Multicolor 4D Fluorescence Microscopy using Ultrathin Bessel Light sheets
 SUPPLEMENTARY MATERIAL Multicolor 4D Fluorescence Microscopy using Ultrathin Bessel Light sheets Teng Zhao, Sze Cheung Lau, Ying Wang, Yumian Su, Hao Wang, Aifang Cheng, Karl Herrup, Nancy Y. Ip, Shengwang
SUPPLEMENTARY MATERIAL Multicolor 4D Fluorescence Microscopy using Ultrathin Bessel Light sheets Teng Zhao, Sze Cheung Lau, Ying Wang, Yumian Su, Hao Wang, Aifang Cheng, Karl Herrup, Nancy Y. Ip, Shengwang
The DCS-120 Confocal Scanning FLIM System
 he DCS-120 Confocal Scanning FLIM System he bh DCS-120 confocal scanning FLIM system converts a conventional microscope into a high-performance fluorescence lifetime imaging system. he system is based
he DCS-120 Confocal Scanning FLIM System he bh DCS-120 confocal scanning FLIM system converts a conventional microscope into a high-performance fluorescence lifetime imaging system. he system is based
Instant super-resolution imaging in live cells and embryos via analog image processing
 Nature Methods Instant super-resolution imaging in live cells and embryos via analog image processing Andrew G. York, Panagiotis Chandris, Damian Dalle Nogare, Jeffrey Head, Peter Wawrzusin, Robert S.
Nature Methods Instant super-resolution imaging in live cells and embryos via analog image processing Andrew G. York, Panagiotis Chandris, Damian Dalle Nogare, Jeffrey Head, Peter Wawrzusin, Robert S.
Characteristics of point-focus Simultaneous Spatial and temporal Focusing (SSTF) as a two-photon excited fluorescence microscopy
 Characteristics of point-focus Simultaneous Spatial and temporal Focusing (SSTF) as a two-photon excited fluorescence microscopy Qiyuan Song (M2) and Aoi Nakamura (B4) Abstracts: We theoretically and experimentally
Characteristics of point-focus Simultaneous Spatial and temporal Focusing (SSTF) as a two-photon excited fluorescence microscopy Qiyuan Song (M2) and Aoi Nakamura (B4) Abstracts: We theoretically and experimentally
BASICS OF CONFOCAL IMAGING (PART I)
 BASICS OF CONFOCAL IMAGING (PART I) INTERNAL COURSE 2012 LIGHT MICROSCOPY Lateral resolution Transmission Fluorescence d min 1.22 NA obj NA cond 0 0 rairy 0.61 NAobj Ernst Abbe Lord Rayleigh Depth of field
BASICS OF CONFOCAL IMAGING (PART I) INTERNAL COURSE 2012 LIGHT MICROSCOPY Lateral resolution Transmission Fluorescence d min 1.22 NA obj NA cond 0 0 rairy 0.61 NAobj Ernst Abbe Lord Rayleigh Depth of field
Supplementary Figure S1. Schematic representation of different functionalities that could be
 Supplementary Figure S1. Schematic representation of different functionalities that could be obtained using the fiber-bundle approach This schematic representation shows some example of the possible functions
Supplementary Figure S1. Schematic representation of different functionalities that could be obtained using the fiber-bundle approach This schematic representation shows some example of the possible functions
Leica TCS SP8 Quick Start Guide
 Leica TCS SP8 Quick Start Guide Leica TCS SP8 System Overview Start-Up Procedure 1. Turn on the CTR Control Box, Fluorescent Light for the microscope stand. 2. Turn on the Scanner Power (1) on the front
Leica TCS SP8 Quick Start Guide Leica TCS SP8 System Overview Start-Up Procedure 1. Turn on the CTR Control Box, Fluorescent Light for the microscope stand. 2. Turn on the Scanner Power (1) on the front
Supplemental Figure 1: Histogram of 63x Objective Lens z axis Calculated Resolutions. Results from the MetroloJ z axis fits for 5 beads from each
 Supplemental Figure 1: Histogram of 63x Objective Lens z axis Calculated Resolutions. Results from the MetroloJ z axis fits for 5 beads from each lens with a 1 Airy unit pinhole setting. Many water lenses
Supplemental Figure 1: Histogram of 63x Objective Lens z axis Calculated Resolutions. Results from the MetroloJ z axis fits for 5 beads from each lens with a 1 Airy unit pinhole setting. Many water lenses
Confocal Imaging Through Scattering Media with a Volume Holographic Filter
 Confocal Imaging Through Scattering Media with a Volume Holographic Filter Michal Balberg +, George Barbastathis*, Sergio Fantini % and David J. Brady University of Illinois at Urbana-Champaign, Urbana,
Confocal Imaging Through Scattering Media with a Volume Holographic Filter Michal Balberg +, George Barbastathis*, Sergio Fantini % and David J. Brady University of Illinois at Urbana-Champaign, Urbana,
Resolution. [from the New Merriam-Webster Dictionary, 1989 ed.]:
![Resolution. [from the New Merriam-Webster Dictionary, 1989 ed.]: Resolution. [from the New Merriam-Webster Dictionary, 1989 ed.]:](/thumbs/71/66141426.jpg) Resolution [from the New Merriam-Webster Dictionary, 1989 ed.]: resolve v : 1 to break up into constituent parts: ANALYZE; 2 to find an answer to : SOLVE; 3 DETERMINE, DECIDE; 4 to make or pass a formal
Resolution [from the New Merriam-Webster Dictionary, 1989 ed.]: resolve v : 1 to break up into constituent parts: ANALYZE; 2 to find an answer to : SOLVE; 3 DETERMINE, DECIDE; 4 to make or pass a formal
Nikon. King s College London. Imaging Centre. N-SIM guide NIKON IMAGING KING S COLLEGE LONDON
 N-SIM guide NIKON IMAGING CENTRE @ KING S COLLEGE LONDON Starting-up / Shut-down The NSIM hardware is calibrated after system warm-up occurs. It is recommended that you turn-on the system for at least
N-SIM guide NIKON IMAGING CENTRE @ KING S COLLEGE LONDON Starting-up / Shut-down The NSIM hardware is calibrated after system warm-up occurs. It is recommended that you turn-on the system for at least
Flatness of Dichroic Beamsplitters Affects Focus and Image Quality
 Flatness of Dichroic Beamsplitters Affects Focus and Image Quality Flatness of Dichroic Beamsplitters Affects Focus and Image Quality 1. Introduction Even though fluorescence microscopy has become a routine
Flatness of Dichroic Beamsplitters Affects Focus and Image Quality Flatness of Dichroic Beamsplitters Affects Focus and Image Quality 1. Introduction Even though fluorescence microscopy has become a routine
Be aware that there is no universal notation for the various quantities.
 Fourier Optics v2.4 Ray tracing is limited in its ability to describe optics because it ignores the wave properties of light. Diffraction is needed to explain image spatial resolution and contrast and
Fourier Optics v2.4 Ray tracing is limited in its ability to describe optics because it ignores the wave properties of light. Diffraction is needed to explain image spatial resolution and contrast and
Leica TCS SP8 Quick Start Guide
 Leica TCS SP8 Quick Start Guide Leica TCS SP8 System Overview Start-Up Procedure 1. Turn on the CTR Control Box, EL6000 fluorescent light source for the microscope stand. 2. Turn on the Scanner Power
Leica TCS SP8 Quick Start Guide Leica TCS SP8 System Overview Start-Up Procedure 1. Turn on the CTR Control Box, EL6000 fluorescent light source for the microscope stand. 2. Turn on the Scanner Power
3. are adherent cells (ie. cells in suspension are too far away from the coverslip)
 Before you begin, make sure your sample... 1. is seeded on #1.5 coverglass (thickness = 0.17) 2. is an aqueous solution (ie. fixed samples mounted on a slide will not work - not enough difference in refractive
Before you begin, make sure your sample... 1. is seeded on #1.5 coverglass (thickness = 0.17) 2. is an aqueous solution (ie. fixed samples mounted on a slide will not work - not enough difference in refractive
Information & Instructions
 KEY FEATURES 1. USB 3.0 For the Fastest Transfer Rates Up to 10X faster than regular USB 2.0 connections (also USB 2.0 compatible) 2. High Resolution 4.2 MegaPixels resolution gives accurate profile measurements
KEY FEATURES 1. USB 3.0 For the Fastest Transfer Rates Up to 10X faster than regular USB 2.0 connections (also USB 2.0 compatible) 2. High Resolution 4.2 MegaPixels resolution gives accurate profile measurements
a) How big will that physical image of the cells be your camera sensor?
 1. Consider a regular wide-field microscope set up with a 60x, NA = 1.4 objective and a monochromatic digital camera with 8 um pixels, properly positioned in the primary image plane. This microscope is
1. Consider a regular wide-field microscope set up with a 60x, NA = 1.4 objective and a monochromatic digital camera with 8 um pixels, properly positioned in the primary image plane. This microscope is
Training Guide for Leica SP8 Confocal/Multiphoton Microscope
 Training Guide for Leica SP8 Confocal/Multiphoton Microscope LAS AF v3.3 Optical Imaging & Vital Microscopy Core Baylor College of Medicine (2017) Power ON Routine 1 2 Turn ON power switch for epifluorescence
Training Guide for Leica SP8 Confocal/Multiphoton Microscope LAS AF v3.3 Optical Imaging & Vital Microscopy Core Baylor College of Medicine (2017) Power ON Routine 1 2 Turn ON power switch for epifluorescence
Enhancement of the lateral resolution and the image quality in a line-scanning tomographic optical microscope
 Summary of the PhD thesis Enhancement of the lateral resolution and the image quality in a line-scanning tomographic optical microscope Author: Dudás, László Supervisors: Prof. Dr. Szabó, Gábor and Dr.
Summary of the PhD thesis Enhancement of the lateral resolution and the image quality in a line-scanning tomographic optical microscope Author: Dudás, László Supervisors: Prof. Dr. Szabó, Gábor and Dr.
Examination, TEN1, in courses SK2500/SK2501, Physics of Biomedical Microscopy,
 KTH Applied Physics Examination, TEN1, in courses SK2500/SK2501, Physics of Biomedical Microscopy, 2009-06-05, 8-13, FB51 Allowed aids: Compendium Imaging Physics (handed out) Compendium Light Microscopy
KTH Applied Physics Examination, TEN1, in courses SK2500/SK2501, Physics of Biomedical Microscopy, 2009-06-05, 8-13, FB51 Allowed aids: Compendium Imaging Physics (handed out) Compendium Light Microscopy
Supplementary Materials
 Supplementary Materials In the supplementary materials of this paper we discuss some practical consideration for alignment of optical components to help unexperienced users to achieve a high performance
Supplementary Materials In the supplementary materials of this paper we discuss some practical consideration for alignment of optical components to help unexperienced users to achieve a high performance
12.4 Alignment and Manufacturing Tolerances for Segmented Telescopes
 330 Chapter 12 12.4 Alignment and Manufacturing Tolerances for Segmented Telescopes Similar to the JWST, the next-generation large-aperture space telescope for optical and UV astronomy has a segmented
330 Chapter 12 12.4 Alignment and Manufacturing Tolerances for Segmented Telescopes Similar to the JWST, the next-generation large-aperture space telescope for optical and UV astronomy has a segmented
Determination of the Focal Width with the Focal Width Script
 Tutorial Determination of the Focal Width with the Focal Width Script Summary This tutorial shows step-by-step, how to determine the Point Spread Function (PSF) of a microscopic system using the Focal
Tutorial Determination of the Focal Width with the Focal Width Script Summary This tutorial shows step-by-step, how to determine the Point Spread Function (PSF) of a microscopic system using the Focal
More fancy SPIM, Even fancier SPIM
 More fancy SPIM, Even fancier SPIM Last class Light sheet microscopy Fancy SPIM (ispim, dspim, etc ) This class Multi camera SPIM SIM SPIM Bessels d x,y = λ em 2 NA d z = 2 NA λ ex + n(1 cosθ λ em 1 IsoView
More fancy SPIM, Even fancier SPIM Last class Light sheet microscopy Fancy SPIM (ispim, dspim, etc ) This class Multi camera SPIM SIM SPIM Bessels d x,y = λ em 2 NA d z = 2 NA λ ex + n(1 cosθ λ em 1 IsoView
plasmonic nanoblock pair
 Nanostructured potential of optical trapping using a plasmonic nanoblock pair Yoshito Tanaka, Shogo Kaneda and Keiji Sasaki* Research Institute for Electronic Science, Hokkaido University, Sapporo 1-2,
Nanostructured potential of optical trapping using a plasmonic nanoblock pair Yoshito Tanaka, Shogo Kaneda and Keiji Sasaki* Research Institute for Electronic Science, Hokkaido University, Sapporo 1-2,
Supplementary Information. Stochastic Optical Reconstruction Microscopy Imaging of Microtubule Arrays in Intact Arabidopsis thaliana Seedling Roots
 Supplementary Information Stochastic Optical Reconstruction Microscopy Imaging of Microtubule Arrays in Intact Arabidopsis thaliana Seedling Roots Bin Dong 1,, Xiaochen Yang 2,, Shaobin Zhu 1, Diane C.
Supplementary Information Stochastic Optical Reconstruction Microscopy Imaging of Microtubule Arrays in Intact Arabidopsis thaliana Seedling Roots Bin Dong 1,, Xiaochen Yang 2,, Shaobin Zhu 1, Diane C.
Nikon Instruments Europe
 Nikon Instruments Europe Recommendations for N-SIM sample preparation and image reconstruction Dear customer, We hope you find the following guidelines useful in order to get the best performance out of
Nikon Instruments Europe Recommendations for N-SIM sample preparation and image reconstruction Dear customer, We hope you find the following guidelines useful in order to get the best performance out of
Fast, high-contrast imaging of animal development with scanned light sheet based structured-illumination microscopy
 nature methods Fast, high-contrast imaging of animal development with scanned light sheet based structured-illumination microscopy Philipp J Keller, Annette D Schmidt, Anthony Santella, Khaled Khairy,
nature methods Fast, high-contrast imaging of animal development with scanned light sheet based structured-illumination microscopy Philipp J Keller, Annette D Schmidt, Anthony Santella, Khaled Khairy,
3D light microscopy techniques
 3D light microscopy techniques The image of a point is a 3D feature In-focus image Out-of-focus image The image of a point is not a point Point Spread Function (PSF) 1D imaging 1 1 2! NA = 0.5! NA 2D imaging
3D light microscopy techniques The image of a point is a 3D feature In-focus image Out-of-focus image The image of a point is not a point Point Spread Function (PSF) 1D imaging 1 1 2! NA = 0.5! NA 2D imaging
High resolution extended depth of field microscopy using wavefront coding
 High resolution extended depth of field microscopy using wavefront coding Matthew R. Arnison *, Peter Török #, Colin J. R. Sheppard *, W. T. Cathey +, Edward R. Dowski, Jr. +, Carol J. Cogswell *+ * Physical
High resolution extended depth of field microscopy using wavefront coding Matthew R. Arnison *, Peter Török #, Colin J. R. Sheppard *, W. T. Cathey +, Edward R. Dowski, Jr. +, Carol J. Cogswell *+ * Physical
Fiber Optic Communications
 Fiber Optic Communications ( Chapter 2: Optics Review ) presented by Prof. Kwang-Chun Ho 1 Section 2.4: Numerical Aperture Consider an optical receiver: where the diameter of photodetector surface area
Fiber Optic Communications ( Chapter 2: Optics Review ) presented by Prof. Kwang-Chun Ho 1 Section 2.4: Numerical Aperture Consider an optical receiver: where the diameter of photodetector surface area
1 Co Localization and Working flow with the lsm700
 1 Co Localization and Working flow with the lsm700 Samples -1 slide = mousse intestine, Dapi / Ki 67 with Cy3/ BrDU with alexa 488. -1 slide = mousse intestine, Dapi / Ki 67 with Cy3/ no BrDU (but with
1 Co Localization and Working flow with the lsm700 Samples -1 slide = mousse intestine, Dapi / Ki 67 with Cy3/ BrDU with alexa 488. -1 slide = mousse intestine, Dapi / Ki 67 with Cy3/ no BrDU (but with
Presented by Jerry Hubbell Lake of the Woods Observatory (MPC I24) President, Rappahannock Astronomy Club
 Presented by Jerry Hubbell Lake of the Woods Observatory (MPC I24) President, Rappahannock Astronomy Club ENGINEERING A FIBER-FED FED SPECTROMETER FOR ASTRONOMICAL USE Objectives Discuss the engineering
Presented by Jerry Hubbell Lake of the Woods Observatory (MPC I24) President, Rappahannock Astronomy Club ENGINEERING A FIBER-FED FED SPECTROMETER FOR ASTRONOMICAL USE Objectives Discuss the engineering
Practical work no. 3: Confocal Live Cell Microscopy
 Practical work no. 3: Confocal Live Cell Microscopy Course Instructor: Mikko Liljeström (MIU) 1 Background Confocal microscopy: The main idea behind confocality is that it suppresses the signal outside
Practical work no. 3: Confocal Live Cell Microscopy Course Instructor: Mikko Liljeström (MIU) 1 Background Confocal microscopy: The main idea behind confocality is that it suppresses the signal outside
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Supplemental correlative nanomanipulation-fluorescence traces probing nascent RNA and fluorescent Mfd during TCR initiation. Supplemental correlative nanomanipulation-fluorescence
Supplementary Figure 1 Supplemental correlative nanomanipulation-fluorescence traces probing nascent RNA and fluorescent Mfd during TCR initiation. Supplemental correlative nanomanipulation-fluorescence
Εισαγωγική στην Οπτική Απεικόνιση
 Εισαγωγική στην Οπτική Απεικόνιση Δημήτριος Τζεράνης, Ph.D. Εμβιομηχανική και Βιοϊατρική Τεχνολογία Τμήμα Μηχανολόγων Μηχανικών Ε.Μ.Π. Χειμερινό Εξάμηνο 2015 Light: A type of EM Radiation EM radiation:
Εισαγωγική στην Οπτική Απεικόνιση Δημήτριος Τζεράνης, Ph.D. Εμβιομηχανική και Βιοϊατρική Τεχνολογία Τμήμα Μηχανολόγων Μηχανικών Ε.Μ.Π. Χειμερινό Εξάμηνο 2015 Light: A type of EM Radiation EM radiation:
Optical coherence tomography
 Optical coherence tomography Peter E. Andersen Optics and Plasma Research Department Risø National Laboratory E-mail peter.andersen@risoe.dk Outline Part I: Introduction to optical coherence tomography
Optical coherence tomography Peter E. Andersen Optics and Plasma Research Department Risø National Laboratory E-mail peter.andersen@risoe.dk Outline Part I: Introduction to optical coherence tomography
Practical Flatness Tech Note
 Practical Flatness Tech Note Understanding Laser Dichroic Performance BrightLine laser dichroic beamsplitters set a new standard for super-resolution microscopy with λ/10 flatness per inch, P-V. We ll
Practical Flatness Tech Note Understanding Laser Dichroic Performance BrightLine laser dichroic beamsplitters set a new standard for super-resolution microscopy with λ/10 flatness per inch, P-V. We ll
Supplementary Information for. Surface Waves. Angelo Angelini, Elsie Barakat, Peter Munzert, Luca Boarino, Natascia De Leo,
 Supplementary Information for Focusing and Extraction of Light mediated by Bloch Surface Waves Angelo Angelini, Elsie Barakat, Peter Munzert, Luca Boarino, Natascia De Leo, Emanuele Enrico, Fabrizio Giorgis,
Supplementary Information for Focusing and Extraction of Light mediated by Bloch Surface Waves Angelo Angelini, Elsie Barakat, Peter Munzert, Luca Boarino, Natascia De Leo, Emanuele Enrico, Fabrizio Giorgis,
Development of a High-speed Super-resolution Confocal Scanner
 Development of a High-speed Super-resolution Confocal Scanner Takuya Azuma *1 Takayuki Kei *1 Super-resolution microscopy techniques that overcome the spatial resolution limit of conventional light microscopy
Development of a High-speed Super-resolution Confocal Scanner Takuya Azuma *1 Takayuki Kei *1 Super-resolution microscopy techniques that overcome the spatial resolution limit of conventional light microscopy
Shreyash Tandon M.S. III Year
 Shreyash Tandon M.S. III Year 20091015 Confocal microscopy is a powerful tool for generating high-resolution images and 3-D reconstructions of a specimen by using point illumination and a spatial pinhole
Shreyash Tandon M.S. III Year 20091015 Confocal microscopy is a powerful tool for generating high-resolution images and 3-D reconstructions of a specimen by using point illumination and a spatial pinhole
Adaptive optimisation of illumination beam profiles in fluorescence microscopy
 Adaptive optimisation of illumination beam profiles in fluorescence microscopy T. J. Mitchell a, C. D. Saunter a, W. O Nions a, J. M. Girkin a, G. D. Love a a Centre for Advanced nstrumentation & Biophysical
Adaptive optimisation of illumination beam profiles in fluorescence microscopy T. J. Mitchell a, C. D. Saunter a, W. O Nions a, J. M. Girkin a, G. D. Love a a Centre for Advanced nstrumentation & Biophysical
11Beamage-3. CMOS Beam Profiling Cameras
 11Beamage-3 CMOS Beam Profiling Cameras Key Features USB 3.0 FOR THE FASTEST TRANSFER RATES Up to 10X faster than regular USB 2.0 connections (also USB 2.0 compatible) HIGH RESOLUTION 2.2 MPixels resolution
11Beamage-3 CMOS Beam Profiling Cameras Key Features USB 3.0 FOR THE FASTEST TRANSFER RATES Up to 10X faster than regular USB 2.0 connections (also USB 2.0 compatible) HIGH RESOLUTION 2.2 MPixels resolution
SETTING UP OF A TOTAL INTERNAL REFLECTION FLUORESCENT MICROSCOPE (TIRFM) SYSTEM: A DETAILED OVERVIEW
 PK ISSN 0022-2941; CODEN JNSMAC Vol. 51, (2011) PP 31-45 SETTING UP OF A TOTAL INTERNAL REFLECTION FLUORESCENT MICROSCOPE (TIRFM) SYSTEM: A DETAILED OVERVIEW A. R. KHAN 1 *, S. AKHLAQ 1, M. N. B. ABID
PK ISSN 0022-2941; CODEN JNSMAC Vol. 51, (2011) PP 31-45 SETTING UP OF A TOTAL INTERNAL REFLECTION FLUORESCENT MICROSCOPE (TIRFM) SYSTEM: A DETAILED OVERVIEW A. R. KHAN 1 *, S. AKHLAQ 1, M. N. B. ABID
Supplementary information, Figure S1A-S1H The thickness and the uniformity of the light sheet at different DOFs. By
 Supplementary information, Figure S1A-S1H The thickness and the uniformity of the light sheet at different DOFs. By imaging FITC-containing solution, the thickness of the light sheet generated by the P3A-DSLM
Supplementary information, Figure S1A-S1H The thickness and the uniformity of the light sheet at different DOFs. By imaging FITC-containing solution, the thickness of the light sheet generated by the P3A-DSLM
TSBB09 Image Sensors 2018-HT2. Image Formation Part 1
 TSBB09 Image Sensors 2018-HT2 Image Formation Part 1 Basic physics Electromagnetic radiation consists of electromagnetic waves With energy That propagate through space The waves consist of transversal
TSBB09 Image Sensors 2018-HT2 Image Formation Part 1 Basic physics Electromagnetic radiation consists of electromagnetic waves With energy That propagate through space The waves consist of transversal
Dynamic beam shaping with programmable diffractive optics
 Dynamic beam shaping with programmable diffractive optics Bosanta R. Boruah Dept. of Physics, GU Page 1 Outline of the talk Introduction Holography Programmable diffractive optics Laser scanning confocal
Dynamic beam shaping with programmable diffractive optics Bosanta R. Boruah Dept. of Physics, GU Page 1 Outline of the talk Introduction Holography Programmable diffractive optics Laser scanning confocal
Optical Sectioning Deep Inside Live Embryos by Selective Plane Illumination Microscopy
 Supporting Online Material Optical Sectioning Deep Inside Live Embryos by Selective Plane Illumination Microscopy 1 Material and methods 1.1 Setup Jan Huisken, Jim Swoger, Filippo Del Bene, Joachim Wittbrodt,
Supporting Online Material Optical Sectioning Deep Inside Live Embryos by Selective Plane Illumination Microscopy 1 Material and methods 1.1 Setup Jan Huisken, Jim Swoger, Filippo Del Bene, Joachim Wittbrodt,
SUPPLEMENTARY INFORMATION
 Optically reconfigurable metasurfaces and photonic devices based on phase change materials S1: Schematic diagram of the experimental setup. A Ti-Sapphire femtosecond laser (Coherent Chameleon Vision S)
Optically reconfigurable metasurfaces and photonic devices based on phase change materials S1: Schematic diagram of the experimental setup. A Ti-Sapphire femtosecond laser (Coherent Chameleon Vision S)
Zeiss 880 Training Notes Zen 2.3
 Zeiss 880 Training Notes Zen 2.3 1 Turn on the HXP 120V Lamp 2 Turn on Main Power Switch Turn on the Systems PC Switch Turn on the Components Switch. 3 4 5 Turn on the PC and log into your account. Start
Zeiss 880 Training Notes Zen 2.3 1 Turn on the HXP 120V Lamp 2 Turn on Main Power Switch Turn on the Systems PC Switch Turn on the Components Switch. 3 4 5 Turn on the PC and log into your account. Start
ANSWER KEY Lab 2 (IGB): Bright Field and Fluorescence Optical Microscopy and Sectioning
 Phys598BP Spring 2016 University of Illinois at Urbana-Champaign ANSWER KEY Lab 2 (IGB): Bright Field and Fluorescence Optical Microscopy and Sectioning Location: IGB Core Microscopy Facility Microscope:
Phys598BP Spring 2016 University of Illinois at Urbana-Champaign ANSWER KEY Lab 2 (IGB): Bright Field and Fluorescence Optical Microscopy and Sectioning Location: IGB Core Microscopy Facility Microscope:
Instruction manual for T3DS software. Tool for THz Time-Domain Spectroscopy. Release 4.0
 Instruction manual for T3DS software Release 4.0 Table of contents 0. Setup... 3 1. Start-up... 5 2. Input parameters and delay line control... 6 3. Slow scan measurement... 8 4. Fast scan measurement...
Instruction manual for T3DS software Release 4.0 Table of contents 0. Setup... 3 1. Start-up... 5 2. Input parameters and delay line control... 6 3. Slow scan measurement... 8 4. Fast scan measurement...
Chapter 18 Optical Elements
 Chapter 18 Optical Elements GOALS When you have mastered the content of this chapter, you will be able to achieve the following goals: Definitions Define each of the following terms and use it in an operational
Chapter 18 Optical Elements GOALS When you have mastered the content of this chapter, you will be able to achieve the following goals: Definitions Define each of the following terms and use it in an operational
1 Set up the confocal light path for imaging a green dye (Alexa488-EGFP). For example, the
 1 Set up the confocal light path for imaging a green dye (Alexa488-EGFP). For example, the light path as shown here using the 488 nm LASER (Laser Unit 1) reflecting off of the 405/488 nm Dichroic mirror
1 Set up the confocal light path for imaging a green dye (Alexa488-EGFP). For example, the light path as shown here using the 488 nm LASER (Laser Unit 1) reflecting off of the 405/488 nm Dichroic mirror
An Optical Characteristic Testing System for the Infrared Fiber in a Transmission Bandwidth 9-11μm
 An Optical Characteristic Testing System for the Infrared Fiber in a Transmission Bandwidth 9-11μm Ma Yangwu *, Liang Di ** Center for Optical and Electromagnetic Research, State Key Lab of Modern Optical
An Optical Characteristic Testing System for the Infrared Fiber in a Transmission Bandwidth 9-11μm Ma Yangwu *, Liang Di ** Center for Optical and Electromagnetic Research, State Key Lab of Modern Optical
Heisenberg) relation applied to space and transverse wavevector
 2. Optical Microscopy 2.1 Principles A microscope is in principle nothing else than a simple lens system for magnifying small objects. The first lens, called the objective, has a short focal length (a
2. Optical Microscopy 2.1 Principles A microscope is in principle nothing else than a simple lens system for magnifying small objects. The first lens, called the objective, has a short focal length (a
Deep and fast live imaging with two-photon scanned light-sheet microscopy
 Nature Methods Deep and fast live imaging with two-photon scanned light-sheet microscopy Thai V Truong, Willy Supatto, Davis S Koos, John M Choi & Scott E Fraser Supplementary Figure 1 Supplementary Figure
Nature Methods Deep and fast live imaging with two-photon scanned light-sheet microscopy Thai V Truong, Willy Supatto, Davis S Koos, John M Choi & Scott E Fraser Supplementary Figure 1 Supplementary Figure
Basics of confocal imaging (part I)
 Basics of confocal imaging (part I) Swiss Institute of Technology (EPFL) Faculty of Life Sciences Head of BIOIMAGING AND OPTICS BIOP arne.seitz@epfl.ch Lateral resolution BioImaging &Optics Platform Light
Basics of confocal imaging (part I) Swiss Institute of Technology (EPFL) Faculty of Life Sciences Head of BIOIMAGING AND OPTICS BIOP arne.seitz@epfl.ch Lateral resolution BioImaging &Optics Platform Light
880 Quantum Electronics Optional Lab Construct A Pulsed Dye Laser
 880 Quantum Electronics Optional Lab Construct A Pulsed Dye Laser The goal of this lab is to give you experience aligning a laser and getting it to lase more-or-less from scratch. There is no write-up
880 Quantum Electronics Optional Lab Construct A Pulsed Dye Laser The goal of this lab is to give you experience aligning a laser and getting it to lase more-or-less from scratch. There is no write-up
Supporting Information
 Copyright WILEY-VCH Verlag GmbH & Co. KGaA, 69469 Weinheim, Germany, 2012. Supporting Information for Adv. Mater., DOI: 10.1002/adma.201203033 Solid Immersion Facilitates Fluorescence Microscopy with Nanometer
Copyright WILEY-VCH Verlag GmbH & Co. KGaA, 69469 Weinheim, Germany, 2012. Supporting Information for Adv. Mater., DOI: 10.1002/adma.201203033 Solid Immersion Facilitates Fluorescence Microscopy with Nanometer
The Zeiss AiryScan System, Confocal Four.
 The Zeiss AiryScan System, Confocal Four. Overview. The Zeiss AiryScan module is a segmented, radially stacked GaASP detector and collector system designed to subsample the airy disk of a point emission
The Zeiss AiryScan System, Confocal Four. Overview. The Zeiss AiryScan module is a segmented, radially stacked GaASP detector and collector system designed to subsample the airy disk of a point emission
Akinori Mitani and Geoff Weiner BGGN 266 Spring 2013 Non-linear optics final report. Introduction and Background
 Akinori Mitani and Geoff Weiner BGGN 266 Spring 2013 Non-linear optics final report Introduction and Background Two-photon microscopy is a type of fluorescence microscopy using two-photon excitation. It
Akinori Mitani and Geoff Weiner BGGN 266 Spring 2013 Non-linear optics final report Introduction and Background Two-photon microscopy is a type of fluorescence microscopy using two-photon excitation. It
Fundamentals of Light Microscopy II: Fluorescence, Deconvolution, Confocal, Multiphoton, Spectral microscopy. Integrated Microscopy Course
 Fundamentals of Light Microscopy II: Fluorescence, Deconvolution, Confocal, Multiphoton, Spectral microscopy Integrated Microscopy Course Review Lecture 1: Microscopy Basics Light train Kohler illumination*
Fundamentals of Light Microscopy II: Fluorescence, Deconvolution, Confocal, Multiphoton, Spectral microscopy Integrated Microscopy Course Review Lecture 1: Microscopy Basics Light train Kohler illumination*
Sizing of nano structures below the diffraction limit using laser scanning microscopy
 Sizing of nano structures below the diffraction limit using laser scanning microscopy JAN BERGSTRAND Master s Thesis Supervisor: Stefan Wennmalm Examiner: Jerker Widengren trita? Abstract The resolution
Sizing of nano structures below the diffraction limit using laser scanning microscopy JAN BERGSTRAND Master s Thesis Supervisor: Stefan Wennmalm Examiner: Jerker Widengren trita? Abstract The resolution
High-Resolution Bubble Printing of Quantum Dots
 SUPPORTING INFORMATION High-Resolution Bubble Printing of Quantum Dots Bharath Bangalore Rajeeva 1, Linhan Lin 1, Evan P. Perillo 2, Xiaolei Peng 1, William W. Yu 3, Andrew K. Dunn 2, Yuebing Zheng 1,*
SUPPORTING INFORMATION High-Resolution Bubble Printing of Quantum Dots Bharath Bangalore Rajeeva 1, Linhan Lin 1, Evan P. Perillo 2, Xiaolei Peng 1, William W. Yu 3, Andrew K. Dunn 2, Yuebing Zheng 1,*
ECEN 4606, UNDERGRADUATE OPTICS LAB
 ECEN 4606, UNDERGRADUATE OPTICS LAB Lab 2: Imaging 1 the Telescope Original Version: Prof. McLeod SUMMARY: In this lab you will become familiar with the use of one or more lenses to create images of distant
ECEN 4606, UNDERGRADUATE OPTICS LAB Lab 2: Imaging 1 the Telescope Original Version: Prof. McLeod SUMMARY: In this lab you will become familiar with the use of one or more lenses to create images of distant
AgilOptics mirrors increase coupling efficiency into a 4 µm diameter fiber by 750%.
 Application Note AN004: Fiber Coupling Improvement Introduction AgilOptics mirrors increase coupling efficiency into a 4 µm diameter fiber by 750%. Industrial lasers used for cutting, welding, drilling,
Application Note AN004: Fiber Coupling Improvement Introduction AgilOptics mirrors increase coupling efficiency into a 4 µm diameter fiber by 750%. Industrial lasers used for cutting, welding, drilling,
Lab Report 3: Speckle Interferometry LIN PEI-YING, BAIG JOVERIA
 Lab Report 3: Speckle Interferometry LIN PEI-YING, BAIG JOVERIA Abstract: Speckle interferometry (SI) has become a complete technique over the past couple of years and is widely used in many branches of
Lab Report 3: Speckle Interferometry LIN PEI-YING, BAIG JOVERIA Abstract: Speckle interferometry (SI) has become a complete technique over the past couple of years and is widely used in many branches of
BEAMAGE-3.0 KEY FEATURES BEAM DIAGNOSTICS AVAILABLE MODELS MAIN FUNCTIONS SEE ALSO ACCESSORIES. CMOS Beam Profiling Cameras
 BEAM DIAGNOSTICS BEAM DIAGNOSTICS SPECIAL PRODUCTS OEM DETECTORS THZ DETECTORS PHOTO DETECTORS HIGH POWER DETECTORS POWER DETECTORS ENERGY DETECTORS MONITORS CMOS Beam Profiling Cameras AVAILABLE MODELS
BEAM DIAGNOSTICS BEAM DIAGNOSTICS SPECIAL PRODUCTS OEM DETECTORS THZ DETECTORS PHOTO DETECTORS HIGH POWER DETECTORS POWER DETECTORS ENERGY DETECTORS MONITORS CMOS Beam Profiling Cameras AVAILABLE MODELS
Rates of excitation, emission, ISC
 Bi177 Lecture 4 Fluorescence Microscopy Phenomenon of Fluorescence Energy Diagram Rates of excitation, emission, ISC Practical Issues Lighting, Filters More on diffraction Point Spread Functions Thus Far,
Bi177 Lecture 4 Fluorescence Microscopy Phenomenon of Fluorescence Energy Diagram Rates of excitation, emission, ISC Practical Issues Lighting, Filters More on diffraction Point Spread Functions Thus Far,
D2.1 Operating 2D STED Microscope
 D2.1 Operating 2D STED Microscope Nature: Report Dissemination Level: Public Lead Beneficiary: UNIVDUN Author(s): Piotr Zdankowski Work Package: WP2 Task: ESR5 Version: 0.02 Last modified: 24/04/2017 Status:
D2.1 Operating 2D STED Microscope Nature: Report Dissemination Level: Public Lead Beneficiary: UNIVDUN Author(s): Piotr Zdankowski Work Package: WP2 Task: ESR5 Version: 0.02 Last modified: 24/04/2017 Status:
ECEN. Spectroscopy. Lab 8. copy. constituents HOMEWORK PR. Figure. 1. Layout of. of the
 ECEN 4606 Lab 8 Spectroscopy SUMMARY: ROBLEM 1: Pedrotti 3 12-10. In this lab, you will design, build and test an optical spectrum analyzer and use it for both absorption and emission spectroscopy. The
ECEN 4606 Lab 8 Spectroscopy SUMMARY: ROBLEM 1: Pedrotti 3 12-10. In this lab, you will design, build and test an optical spectrum analyzer and use it for both absorption and emission spectroscopy. The
Reflecting optical system to increase signal intensity. in confocal microscopy
 Reflecting optical system to increase signal intensity in confocal microscopy DongKyun Kang *, JungWoo Seo, DaeGab Gweon Nano Opto Mechatronics Laboratory, Dept. of Mechanical Engineering, Korea Advanced
Reflecting optical system to increase signal intensity in confocal microscopy DongKyun Kang *, JungWoo Seo, DaeGab Gweon Nano Opto Mechatronics Laboratory, Dept. of Mechanical Engineering, Korea Advanced
Zeiss LSM 780 Protocol
 Zeiss LSM 780 Protocol 1) System Startup F Please note the sign-up policy. You must inform the facility at least 24 hours beforehand if you can t come; otherwise, you will receive a charge for unused time.
Zeiss LSM 780 Protocol 1) System Startup F Please note the sign-up policy. You must inform the facility at least 24 hours beforehand if you can t come; otherwise, you will receive a charge for unused time.
Instructions for the Experiment
 Instructions for the Experiment Excitonic States in Atomically Thin Semiconductors 1. Introduction Alongside with electrical measurements, optical measurements are an indispensable tool for the study of
Instructions for the Experiment Excitonic States in Atomically Thin Semiconductors 1. Introduction Alongside with electrical measurements, optical measurements are an indispensable tool for the study of
Transferring wavefront measurements to ablation profiles. Michael Mrochen PhD Swiss Federal Institut of Technology, Zurich IROC Zurich
 Transferring wavefront measurements to ablation profiles Michael Mrochen PhD Swiss Federal Institut of Technology, Zurich IROC Zurich corneal ablation Calculation laser spot positions Centration Calculation
Transferring wavefront measurements to ablation profiles Michael Mrochen PhD Swiss Federal Institut of Technology, Zurich IROC Zurich corneal ablation Calculation laser spot positions Centration Calculation
Properties of Structured Light
 Properties of Structured Light Gaussian Beams Structured light sources using lasers as the illumination source are governed by theories of Gaussian beams. Unlike incoherent sources, coherent laser sources
Properties of Structured Light Gaussian Beams Structured light sources using lasers as the illumination source are governed by theories of Gaussian beams. Unlike incoherent sources, coherent laser sources
Single-photon excitation of morphology dependent resonance
 Single-photon excitation of morphology dependent resonance 3.1 Introduction The examination of morphology dependent resonance (MDR) has been of considerable importance to many fields in optical science.
Single-photon excitation of morphology dependent resonance 3.1 Introduction The examination of morphology dependent resonance (MDR) has been of considerable importance to many fields in optical science.
Imaging Retreat - UMASS Customized real-time confocal and 2-photon imaging
 Imaging Retreat - UMASS 2012 Customized real-time confocal and 2-photon imaging Mike Sanderson Department of Microbiology and Physiological Systems University of Massachusetts Medical School Thanks for
Imaging Retreat - UMASS 2012 Customized real-time confocal and 2-photon imaging Mike Sanderson Department of Microbiology and Physiological Systems University of Massachusetts Medical School Thanks for
Supporting Information
 Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2014 Two-Color RESOLFT Nanoscopy with Green and Red Fluorescent Photochromic Proteins** Flavie Lavoie-Cardinal, [a] Nickels
Supporting Information Copyright Wiley-VCH Verlag GmbH & Co. KGaA, 69451 Weinheim, 2014 Two-Color RESOLFT Nanoscopy with Green and Red Fluorescent Photochromic Proteins** Flavie Lavoie-Cardinal, [a] Nickels
ERS KEY FEATURES BEAM DIAGNOSTICS MAIN FUNCTIONS AVAILABLE MODEL. CMOS Beam Profiling Camera. 1 USB 3.0 for the Fastest Transfer Rates
 POWER DETECTORS ENERGY DETECTORS MONITORS SPECIAL PRODUCTS OEM DETECTORS THZ DETECTORS PHOTO DETECTORS HIGH POWER DETECTORS CAMERA PROFIL- CMOS Beam Profiling Camera KEY FEATURES ERS 1 USB 3.0 for the
POWER DETECTORS ENERGY DETECTORS MONITORS SPECIAL PRODUCTS OEM DETECTORS THZ DETECTORS PHOTO DETECTORS HIGH POWER DETECTORS CAMERA PROFIL- CMOS Beam Profiling Camera KEY FEATURES ERS 1 USB 3.0 for the
ASD and Speckle Interferometry. Dave Rowe, CTO, PlaneWave Instruments
 ASD and Speckle Interferometry Dave Rowe, CTO, PlaneWave Instruments Part 1: Modeling the Astronomical Image Static Dynamic Stochastic Start with Object, add Diffraction and Telescope Aberrations add Atmospheric
ASD and Speckle Interferometry Dave Rowe, CTO, PlaneWave Instruments Part 1: Modeling the Astronomical Image Static Dynamic Stochastic Start with Object, add Diffraction and Telescope Aberrations add Atmospheric
Fluorescence Imaging of Single Spins in Nitrogen-Vacancy centers using a Confocal Microscope. Advanced Lab Course University of Basel
 Fluorescence Imaging of Single Spins in Nitrogen-Vacancy centers using a Confocal Microscope Advanced Lab Course University of Basel October 6, 2015 Contents 1 Introduction 2 2 The Confocal Microscope
Fluorescence Imaging of Single Spins in Nitrogen-Vacancy centers using a Confocal Microscope Advanced Lab Course University of Basel October 6, 2015 Contents 1 Introduction 2 2 The Confocal Microscope
User manual for Olympus SD-OSR spinning disk confocal microscope
 User manual for Olympus SD-OSR spinning disk confocal microscope Ved Prakash, PhD. Research imaging specialist Imaging & histology core University of Texas, Dallas ved.prakash@utdallas.edu Once you open
User manual for Olympus SD-OSR spinning disk confocal microscope Ved Prakash, PhD. Research imaging specialist Imaging & histology core University of Texas, Dallas ved.prakash@utdallas.edu Once you open
Imaging Beyond the Basics: Optimizing Settings on the Leica SP8 Confocal
 Imaging Beyond the Basics: Optimizing Settings on the Leica SP8 Confocal Todays Goal: Introduce some additional functionalities of the Leica SP8 confocal HyD vs. PMT detectors Dye Assistant Scanning By
Imaging Beyond the Basics: Optimizing Settings on the Leica SP8 Confocal Todays Goal: Introduce some additional functionalities of the Leica SP8 confocal HyD vs. PMT detectors Dye Assistant Scanning By
Tutorial Zemax 9: Physical optical modelling I
 Tutorial Zemax 9: Physical optical modelling I 2012-11-04 9 Physical optical modelling I 1 9.1 Gaussian Beams... 1 9.2 Physical Beam Propagation... 3 9.3 Polarization... 7 9.4 Polarization II... 11 9 Physical
Tutorial Zemax 9: Physical optical modelling I 2012-11-04 9 Physical optical modelling I 1 9.1 Gaussian Beams... 1 9.2 Physical Beam Propagation... 3 9.3 Polarization... 7 9.4 Polarization II... 11 9 Physical
Bi/BE 227 Winter Assignment #3. Adding the third dimension: 3D Confocal Imaging
 Bi/BE 227 Winter 2016 Assignment #3 Adding the third dimension: 3D Confocal Imaging Schedule: Jan 20: Assignment Jan 20-Feb 8: Work on assignment Feb 10: Student PowerPoint presentations. Goals for this
Bi/BE 227 Winter 2016 Assignment #3 Adding the third dimension: 3D Confocal Imaging Schedule: Jan 20: Assignment Jan 20-Feb 8: Work on assignment Feb 10: Student PowerPoint presentations. Goals for this
Exercise questions for Machine vision
 Exercise questions for Machine vision This is a collection of exercise questions. These questions are all examination alike which means that similar questions may appear at the written exam. I ve divided
Exercise questions for Machine vision This is a collection of exercise questions. These questions are all examination alike which means that similar questions may appear at the written exam. I ve divided
Optomechanical enhancement of doubly resonant 2D optical nonlinearity
 Supporting information Optomechanical enhancement of doubly resonant 2D optical nonlinearity Fei Yi 3+, Mingliang Ren 3+, Jason C Reed 3, Hai Zhu 3, Jiechang Hou 3, Carl H. Naylor 4, Alan T. Charlie Johnson
Supporting information Optomechanical enhancement of doubly resonant 2D optical nonlinearity Fei Yi 3+, Mingliang Ren 3+, Jason C Reed 3, Hai Zhu 3, Jiechang Hou 3, Carl H. Naylor 4, Alan T. Charlie Johnson
Introduction to Light Microscopy. (Image: T. Wittman, Scripps)
 Introduction to Light Microscopy (Image: T. Wittman, Scripps) The Light Microscope Four centuries of history Vibrant current development One of the most widely used research tools A. Khodjakov et al. Major
Introduction to Light Microscopy (Image: T. Wittman, Scripps) The Light Microscope Four centuries of history Vibrant current development One of the most widely used research tools A. Khodjakov et al. Major
Spectral phase shaping for high resolution CARS spectroscopy around 3000 cm 1
 Spectral phase shaping for high resolution CARS spectroscopy around 3 cm A.C.W. van Rhijn, S. Postma, J.P. Korterik, J.L. Herek, and H.L. Offerhaus Mesa + Research Institute for Nanotechnology, University
Spectral phase shaping for high resolution CARS spectroscopy around 3 cm A.C.W. van Rhijn, S. Postma, J.P. Korterik, J.L. Herek, and H.L. Offerhaus Mesa + Research Institute for Nanotechnology, University
Rapid Adaptive Optical Recovery of Optimal Resolution over Large Volumes
 SUPPLEMENTARY MATERIAL Rapid Adaptive Optical Recovery of Optimal Resolution over Large Volumes Kai Wang, Dan Milkie, Ankur Saxena, Peter Engerer, Thomas Misgeld, Marianne E. Bronner, Jeff Mumm, and Eric
SUPPLEMENTARY MATERIAL Rapid Adaptive Optical Recovery of Optimal Resolution over Large Volumes Kai Wang, Dan Milkie, Ankur Saxena, Peter Engerer, Thomas Misgeld, Marianne E. Bronner, Jeff Mumm, and Eric
BEAMAGE KEY FEATURES AVAILABLE MODELS. CMOS Beam Profiling Cameras
 BEAM DIAGNOS TICS Beam Profiling Cameras KEY FEATURES SPECIAL PRODUCTS OEM DETECTORS THZ DETECTORS PHOTO DETECTORS HIGH POWER SOLUTIONS POWER DETECTORS ENERGY DETECTORS MONITORS AVAILABLE MODELS Beamage-3.0
BEAM DIAGNOS TICS Beam Profiling Cameras KEY FEATURES SPECIAL PRODUCTS OEM DETECTORS THZ DETECTORS PHOTO DETECTORS HIGH POWER SOLUTIONS POWER DETECTORS ENERGY DETECTORS MONITORS AVAILABLE MODELS Beamage-3.0
Coherent Laser Measurement and Control Beam Diagnostics
 Coherent Laser Measurement and Control M 2 Propagation Analyzer Measurement and display of CW laser divergence, M 2 (or k) and astigmatism sizes 0.2 mm to 25 mm Wavelengths from 220 nm to 15 µm Determination
Coherent Laser Measurement and Control M 2 Propagation Analyzer Measurement and display of CW laser divergence, M 2 (or k) and astigmatism sizes 0.2 mm to 25 mm Wavelengths from 220 nm to 15 µm Determination
